Abstract
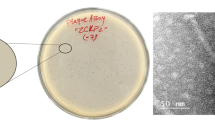
Klebsiella pneumoniae (family Enterobacteriaceae) is a gram-negative bacterium that has strong pathogenicity to humans and can cause sepsis, pneumonia, and urinary tract infection. In recent years, the unreasonable use of antibacterial drugs has led to an increase in drug-resistant strains of K. pneumoniae, a serious threat to public health. Bacteriophages, viruses that infect bacteria, are ubiquitous in the natural environment. They are considered to be the most promising substitute for antibiotics because of their high specificity, high efficiency, high safety, low cost, and short development cycle. In this study, a novel phage designated vB_KpnP_IME279 was successfully isolated from hospital sewage using a multidrug-resistant strain of K. pneumoniae as an indicator. A one-step growth curve showed that vB_KpnP_IME279 has a burst size of 140 plaque-forming units/cell and a latent period of 20 min at its optimal multiplicity of infection (MOI = 0.1). Phage vB_KpnP_IME279 survives in a wide pH range between 3 and 11 and is stable at temperatures ranging from 40 to 60 °C. Ten of the 20 strains of K. pneumoniae including the host bacteria were lysed by the phage vB_KpnP_IME279, and the multilocus sequence typing and wzi typing of the 10 strains were ST11, ST37, ST375, wzi209, wzi52, and wzi72, respectively. The genome of vB_KpnP_IME279 is 42,518 bp long with a G + C content of 59.3%. Electron microscopic observation showed that the phage belongs to the family Podoviridae. BLASTN alignment showed that the genome of the phage has low similarity with currently known phages. The evolutionary relationship between phage vB_KpnP_IME279 and other Podoviridae was analyzed using a phylogenetic tree based on sequences of phage major capsid protein and indicates that the phage vB_KpnP_IME279 belongs to the Podoviridae subfamily. These data enhance understanding of K. pneumoniae phages and will help in development of treatments for multidrug-resistant bacteria using phages.






Similar content being viewed by others
References
Altschul SF, Madden TL, Schaffer AA, Zhang J, Zhang Z, Miller W, Lipman DJ (1997) Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res 25:3389–3402
Bagley ST (1985) Habitat association of Klebsiella species. Infect Control 6:52–58
Benson SD, Bamford JK, Bamford DH, Burnett RM (1999) Viral evolution revealed by bacteriophage PRD1 and human adenovirus coat protein structures. Cell 98:825–833
Brisse S, Fevre C, Passet V, Issenhuth-Jeanjean S, Tournebize R, Diancourt L, Grimont P (2009) Virulent clones of Klebsiella pneumoniae: identification and evolutionary scenario based on genomic and phenotypic characterization. PLoS One 4:e4982
Brisse S, Passet V, Haugaard AB, Babosan A, Kassis-Chikhani N, Struve C, Decre D (2013) wzi gene sequencing, a rapid method for determination of capsular type for Klebsiella strains. J Clin Microbiol 51:4073–4078
Bruynoghe R, Maisin J (1921) Essais de thérapeutique au moyen du bacteriophage. CR Soc Biol 85:1120–1121
Chen Y, Sun E, Song J, Yang L, Wu B (2018) Complete genome sequence of a novel T7-like bacteriophage from a Pasteurella multocida capsular type a isolate. Curr Microbiol 75:574–579
Cisek AA, Dabrowska I, Gregorczyk KP, Wyzewski Z (2017) Phage therapy in bacterial infections treatment: one hundred years after the discovery of bacteriophages. Curr Microbiol 74:277–283
Craven D (2006) What is healthcare-associated pneumonia, and how should it be treated? Curr Opin Infect Dis 19:153–160
Eppler K, Wyckoff E, Goates J, Parr R, Casjens S (1991) Nucleotide sequence of the bacteriophage P22 genes required for DNA packaging. Virology 183:519–538
Gloster TM (2014) Advances in understanding glycosyltransferases from a structural perspective. Curr Opin Struct Biol 28:131–141
Guilliam TA, Keen BA, Brissett NC, Doherty AJ (2015) Primase-polymerases are a functionally diverse superfamily of replication and repair enzymes. Nucleic Acids Res 43:6651–6664
Gupta A (2002) Hospital-acquired infections in the neonatal intensive care unit-Klebsiella pneumoniae. Semin Perinatol 26:340–345
Hooi SH, Hooi ST (2005) Culture-proven bacterial keratitis in a Malaysian general hospital. Med J Malaysia 60:614–623
Kropinski AM, Waddell T, Meng J, Franklin K, Ackermann HW, Ahmed R, Mazzocco A, Yates J 3rd, Lingohr EJ, Johnson RP (2013) The host-range, genomics and proteomics of Escherichia coli O157:H7 bacteriophage rV5. Virol J 10:76
Lefkowitz EJ, Dempsey DM, Hendrickson RC, Orton RJ, Siddell SG, Smith DB (2018) Virus taxonomy: the database of the international committee on taxonomy of viruses (ICTV). Nucleic Acids Res 46:D708–d717
Lu S, Le S, Tan Y, Zhu J, Li M, Rao X, Zou L, Li S, Wang J, Jin X, Huang G, Zhang L, Zhao X, Hu F (2013) Genomic and proteomic analyses of the terminally redundant genome of the Pseudomonas aeruginosa phage PaP1: establishment of genus PaP1-like phages. PLoS One 8:e62933
Patil NA, Basu B, Deobagkar DD, Apte SK, Deobagkar DN (2017) Putative DNA modification methylase DR_C0020 of Deinococcus radiodurans is an atypical SAM dependent C-5 cytosine DNA methylase. Biochim Biophys Acta 1861:593–602
Pereira V, Lopes C, Castro A, Silva J, Gibbs P, Teixeira P (2009) Characterization for enterotoxin production, virulence factors, and antibiotic susceptibility of Staphylococcus aureus isolates from various foods in Portugal. Food Microbiol 26:278–282
Prevelige PE Jr, Cortines JR (2018) Phage assembly and the special role of the portal protein. Curr Opin Virol 31:66–73
Queenan AM, Bush K (2007) Carbapenemases: the versatile beta-lactamases. Clin Microbiol Rev 20:440–458
Ronald A (2003) The etiology of urinary tract infection: traditional and emerging pathogens. Disease-a-month 49:71–82
Shankar PR, Balasubramanium R (2014) Antimicrobial resistance: global report on surveillance 2014. Australas Med J (Online) 7:–237
Sun S, Rao VB, Rossmann MG (2010) Genome packaging in viruses. Curr Opin Struct Biol 20:114–120
Wang H, Jin S, Chen X, Gen X, He Y (2012) Target deletion of the AAA ATPase PpCDC48II in Physcomitrella patens results in freezing sensitivity after cold acclimation. Sci China Life Sci 55:150–157
Wang Y, Wang W, Lv Y, Zheng W, Mi Z, Pei G, An X, Xu X, Han C, Liu J, Zhou C, Tong Y (2014) Characterization and complete genome sequence analysis of novel bacteriophage IME-EFm1 infecting Enterococcus faecium. J Gen Virol 95:2565–2575
Wang R, Xing S, Zhao F, Li P, Mi Z, Shi T, Liu H, Tong Y (2018) Characterization and genome analysis of novel phage vB_EfaP_IME195 infecting Enterococcus faecalis. Virus Genes 54:804–811
Wang R, Cong Y, Mi Z, Fan H, Shi T, Liu H, Tong Y (2019) Characterization and complete genome sequence analysis of phage GP4, a novel lytic Bcep22-like podovirus. Arch Virol 164:2339–2343
Weber-Dabrowska B, Mulczyk M, Gorski A (2000) Bacteriophage therapy of bacterial infections: an update of our institute's experience. Arch Immunol Ther Exp 48:547–551
Wright A, Hawkins CH, Anggard EE, Harper DR (2009) A controlled clinical trial of a therapeutic bacteriophage preparation in chronic otitis due to antibiotic-resistant Pseudomonas aeruginosa; a preliminary report of efficacy. Clin Otolaryngol 34:349–357
Yazdi M, Bouzari M, Ghaemi EA (2019) Isolation and characterization of a potentially novel Siphoviridae phage (vB_SsapS-104) with lytic activity against Staphylococcus saprophyticus isolated from urinary tract infection. Folia Microbiol 64:283–294
Zhang X, Wang Y, Li S, An X, Pei G, Huang Y, Fan H, Mi Z, Zhang Z, Wang W, Chen Y, Tong Y (2015) A novel termini analysis theory using HTS data alone for the identification of Enterococcus phage EF4-like genome termini. BMC Genomics 16:414
Zhu J, Rao X, Tan Y, Xiong K, Hu Z, Chen Z, Jin X, Li S, Chen Y, Hu F (2010) Identification of lytic bacteriophage MmP1, assigned to a new member of T7-like phages infecting Morganella morganii. Genomics 96:167–172
Funding
This research was supported by a grant from the National Key Research and Development Program of China (2016YFC1202705, AWS16J020, and AWS15J006), the National Natural Science Foundation of China (81572045, 81672001, and 81621005), the Fundamental Research Funds for the Central Universities and Research Projects on Biomedical Transformation of China-Japan Friendship Hospital (No. PYBZ1820), the National Science and Technology Major Project (Grant No. 2018ZX10201001), and Project 31900489 supported by National Natural Science Foundation of China.
Author information
Authors and Affiliations
Contributions
Yigang Tong, Hui Liu, Taoxing Shi, and Zhiqiang Mi conceived and designed experiments and critically evaluated the manuscript. Rongrong Zhang, Feiyang Zhao, and Jiuru Wang contributed equally to the writing of this article. Rongrong Zhang was responsible for data and sequence analyses and wrote the manuscript. Feiyang Zhao extracted the phage nucleotide and conducted the sequencing experiments. Jiuru Wang isolated and identified the phage and conducted the biological characterization experiments. Guangqian Pei helps measuring phage DNA sequences. Hang Fan and Lilan Zhangxiang help analysis the bioinformation of the phage. All authors read and approved the final manuscript
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Informed consent
Informed consent was obtained from all individual participants included in the study.
Research involving human participants and/or animals
This article does not contain any studies with human participants or animals performed by any of the authors.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Zhang, R., Zhao, F., Wang, J. et al. Biological characteristics and genome analysis of a novel phage vB_KpnP_IME279 infecting Klebsiella pneumoniae. Folia Microbiol 65, 925–936 (2020). https://doi.org/10.1007/s12223-020-00775-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12223-020-00775-8