Abstract
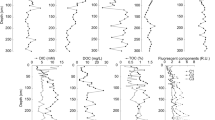
Nitrogen (N) is often a limiting nutrient in lacustrine systems, and bulk organic matter stable isotope ratios of N (δ15N) are widely used in lake sediment studies to interpret N source inputs and lake trophic status. Although records of lacustrine sedimentary δ15N can provide critical information relating to past environmental change, often productivity interpretations from δ15N and lacustrine fossil records yield conflicting interpretations. Furthermore, components of the internal N cycle have substantial isotopic fractionation factors, and likely wield an enormous influence on bulk lacustrine sedimentary δ15N values. Yet apart from cyanobacteria N-fixation, few studies link specific microbial, N-related activity to δ15N variability in lake sediment records. Here, we assess the relationship between lacustrine sedimentary δ15N and microbiome profiles analyzed from extracted sediment DNA using metagenomics. In a ~ 1600-year-long sediment record from a hypersaline lake located on Kiritimati, Republic of Kiribati (1.9° N, 157.4° W), both δ15N and the taxonomy annotations from five unique metagenomes vary with depth. Despite the relatively high abundance of Cyanobacteria, Bacteroidetes, and N-fixation genes in the uppermost sediment, we find the highest δ15N values of the sediment record there. These high values are likely due to denitrification, supported by a relatively high abundance of denitrification genes and taxa responsible for denitrification, such as those found in family Chromatiaceae within the Gamma-proteobacteria. In the deep sediment, N-related biochemical processes are likely suppressed considering the low energy, low nutrient subsurface environment. Low δ15N values observed in deeper sediments co-occur with genes for assimilatory nitrate reduction and ammonification. Thus, metagenomics provides greater clarity with respect to the specific, microbial processes that alter primary δ15N signatures in the subsurface sediment.






Similar content being viewed by others
Change history
27 September 2021
A Correction to this paper has been published: https://doi.org/10.1007/s10933-021-00223-8
References
Anderson MJ (2005) PERMANOVA Permutational multivariate analysis of variance. Austral Ecol. https://doi.org/10.1139/cjfas-58-3-626
Andrews S (2010) FastQC: a quality control tool for high throughput sequence data. http://www.bioinformatics.babraham.ac.uk/projects/fastqc
Bae HS, Morrison E, Chanton JP, Ogram A (2018) Methanogens are major contributors to nitrogen fixation in soils of the Florida Everglades. Appl Environ Microbiol. https://doi.org/10.1128/AEM.02222-17
Blaauw M, Christeny JA (2011) Flexible paleoclimate age-depth models using an autoregressive gamma process. Bayesian Anal. https://doi.org/10.1214/11-BA618
Bolger AM, Lohse M, Usadel B (2014) Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics. https://doi.org/10.1093/bioinformatics/btu170
Botrel M, Gregory-Eaves I, Maranger R (2014) Defining drivers of nitrogen stable isotopes (δ15N) of surface sediments in temperate lakes. J Paleolimnol 52:419–433. https://doi.org/10.1007/s10933-014-9802-6
Brahney J, Ballantyne AP, Turner BL, Spaulding SA, Otu M, Neff JC (2014) Separating the influences of diagenesis, productivity and anthropogenic nitrogen deposition on sedimentary δ15N variations. Org Geochem 75:140–150. https://doi.org/10.1016/j.orggeochem.2014.07.003
Brenner M, Whitmore TJ, Curtis JH, Hodell DA, Schelske CL (1999) Stable isotope (δ13C and δ15N) signatures of sedimented organic matter as indicators of historic lake trophic state. J Paleolimnol 22:205–221. https://doi.org/10.1023/A:1008078222806
Brodie CR, Heaton THE, Leng MJ, Kendrick CP, Casford JSL, Lloyd JM (2011) Evidence for bias in measured δ15N values of terrestrial and aquatic organic materials due to pre-analysis acid treatment methods. Rapid Commun Mass Spectrom. https://doi.org/10.1002/rcm.4970
Buchfink B, Xie C, Huson DH (2014) Fast and sensitive protein alignment using DIAMOND. Nat Methods 12:59
Casciotti KL (2009) Inverse kinetic isotope fractionation during bacterial nitrite oxidation. Geochim Cosmochim Acta. https://doi.org/10.1016/j.gca.2008.12.022
Castelle CJ, Hug LA, Wrighton KC, Thomas BC, Williams KH, Wu D, Tringe SG, Singer SW, Eisen JA, Banfield JF (2013) Extraordinary phylogenetic diversity and metabolic versatility in aquifer sediment. Nat Commun. https://doi.org/10.1038/ncomms3120
Chen I-MA, Chu K, Palaniappan K, Pillay M, Ratner A, Huang J, Huntemann M, Varghese N, White JR, Seshadri R, Smirnova T, Kirton E, Jungbluth SP, Woyke T, Eloe-Fadrosh EA, Ivanova NN, Kyrpides NC (2019) IMG/M vol 5.0: an integrated data management and comparative analysis system for microbial genomes and microbiomes. Nucleic Acids Res 47:D666–D677. https://doi.org/10.1093/nar/gky901
Clarke KR, Gorley R, Somerfield P, Warwick R (2014) Change in marine communities: an approach to statistical analysis and interpretation, 3rd edn. Primer-E Ltd, Plymouth
Conroy JL, Collins AF, Overpeck JT, Bush MB, Cole JE, Anderson DJ (2015) A 400-year isotopic record of seabird response to eastern tropical Pacific productivity. Geo Geogr Environ. https://doi.org/10.1002/geo2.11
Gälman V, Rydberg J, Bigler C (2009) Decadal diagenetic effects on δ13C and δ15N studied in varved lake sediment. Limnol Oceanogr. https://doi.org/10.4319/lo.2009.54.3.0917
Gasparin F, Roemmich D (2016) The strong freshwater anomaly during the onset of the 2015/2016 El Niño. Geophys Res Lett. https://doi.org/10.1002/2016GL069542
Gruber-Vodicka HR, Seah BK, Pruesse E (2019) phyloFlash — Rapid SSU rRNA profiling and targeted assembly from metagenomes. bioRxiv. Doi: https://doi.org/10.1101/521922
Hadas O, Altabet MA, Agnihotri R (2009) Seasonally varying nitrogen isotope biogeochemistry of particulate organic matter in Lake Kinneret. Limnol Oceanogr, Israel. https://doi.org/10.4319/lo.2009.54.1.0075
Higgins MB, Robinson RS, Husson JM, Carter SJ, Pearson A (2012) Dominant eukaryotic export production during ocean anoxic events reflects the importance of recycled NH 4+. Proc Natl Acad Sci U S A. https://doi.org/10.1073/pnas.1104313109
Higley MC, Conroy JL (2019) The hydrological response of surface water to recent climate variability: A remote sensing case study from the central tropical Pacific. Hydrol Process. https://doi.org/10.1002/hyp.13465
Higley MC, Conroy JL, Schmitt S (2018) Last Millennium Meridional Shifts in Hydroclimate in the Central Tropical Pacific. Paleoceanogr Paleoclimatology 33:354–366. https://doi.org/10.1002/2017PA003233
Hodell DA, Schelske CL (1998) Production, sedimentation, and isotopic composition of organic matter in Lake Ontario. Limnol Oceanogr. https://doi.org/10.4319/lo.1998.43.2.0200
Holtgrieve GW, Schindler DE, Hobbs WO et al (2011) A coherent signature of anthropogenic nitrogen deposition to remote watersheds of the Northern Hemisphere. Science 334:1545–1548. https://doi.org/10.1126/science.1212267
Huang TC, Lin RF, Chu MK, Chen HM (1999) Organization and expression of nitrogen-fixation genes in the aerobic nitrogen-fixing unicellular cyanobacterium Synechococcus sp. Strain RF-1. Microbiology. https://doi.org/10.1099/13500872-145-3-743
Huson DH, Beier S, Flade I, Górska A, El-Hadidi M, Mitra S, Ruscheweyh HJ, Tappu R (2016) MEGAN Community Edition: Interactive Exploration and Analysis of Large-Scale Microbiome Sequencing Data. PLoS Comput Biol. https://doi.org/10.1371/journal.pcbi.1004957
Kanehisa M, Sato Y, Kawashima M, Furumichi M, Tanabe M (2016) KEGG as a reference resource for gene and protein annotation. Nucleic Acids Res. https://doi.org/10.1093/nar/gkv1070
Kebschull JM, Zador AM (2015) Sources of PCR-induced distortions in high-throughput sequencing data sets. Nucleic Acids Res. https://doi.org/10.1093/nar/gkv717
Lazar CS, Baker BJ, Seitz K, Hyde AS, Dick GJ, Hinrichs KU, Teske AP (2016) Genomic evidence for distinct carbon substrate preferences and ecological niches of Bathyarchaeota in estuarine sediments. Environ Microbiol. https://doi.org/10.1111/1462-2920.13142
Leavitt PR, Brock CS, Ebel C, Patoine A (2006) Landscape-scale effects of urban nitrogen on a chain of freshwater lakes in central North America. Limnol Oceanogr. https://doi.org/10.4319/lo.2006.51.5.2262
Leininger S, Urich T, Schloter M, Schwark L, Qi J, Nicol GW, Prosser JI, Schuster SC, Schleper C (2006) Archaea predominate among ammonia-oxidizing prokaryotes in soils. Nature. https://doi.org/10.1038/nature04983
Levy-Booth DJ, Prescott CE, Grayston SJ (2014) Microbial functional genes involved in nitrogen fixation, nitrification and denitrification in forest ecosystems. Soil Biol Biochem 75:11
Li D, Liu CM, Luo R, Sadakane K, Lam TW (2015) MEGAHIT: An ultra-fast single-node solution for large and complex metagenomics assembly via succinct de Bruijn graph. Bioinformatics. https://doi.org/10.1093/bioinformatics/btv033
Love MI, Huber W, Anders S (2014) Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. https://doi.org/10.1186/s13059-014-0550-8
Maus I, Rumming M, Bergmann I, Heeg K, Pohl M, Nettmann E, Jaenicke S, Blom J, Pühler A, Schlüter A, Sczyrba A, Klocke M (2018) Characterization of Bathyarchaeota genomes assembled from metagenomes of biofilms residing in mesophilic and thermophilic biogas reactors. Biotechnol Biofuels. https://doi.org/10.1186/s13068-018-1162-4
McMurdie PJ, Holmes S (2013) Phyloseq: An R Package for Reproducible Interactive Analysis and Graphics of Microbiome Census Data. PLoS ONE. https://doi.org/10.1371/journal.pone.0061217
Meng J, Xu J, Qin D, He Y, Xiao X, Wang F (2014) Genetic and functional properties of uncultivated MCG archaea assessed by metagenome and gene expression analyses. ISME J. https://doi.org/10.1038/ismej.2013.174
Nishizawa M, Miyazaki J, Makabe A, Koba K, Takai K (2014) Physiological and isotopic characteristics of nitrogen fixation by hyperthermophilic methanogens: Key insights into nitrogen anabolism of the microbial communities in Archean hydrothermal systems. Geochim Cosmochim Acta. https://doi.org/10.1016/j.gca.2014.04.021
Oksanen J, Blanchet FG, Kindt R, Legendre P, Minchin PR, O’Hara RB (2013) Package vegan. R Packag ver
Overbeek R, Olson R, Pusch GD, Olsen GJ, Davis JJ, Disz T, Edwards RA, Gerdes S, Parrello B, Shukla M, Vonstein V, Wattam AR, Xia F, Stevens R (2014) The SEED and the Rapid Annotation of microbial genomes using Subsystems Technology (RAST). Nucleic Acids Res. https://doi.org/10.1093/nar/gkt1226
Patoine A, Graham MD, Leavitt PR (2006) Spatial variation of nitrogen fixation in lakes of the northern Great Plains. Limnol Oceanogr. https://doi.org/10.4319/lo.2006.51.4.1665
Pereira AD, Cabezas A, Etchebehere C, de Chernicharo CA, L, de Araújo JC, (2017) Microbial communities in anammox reactors: a review. Environ. Technol, Rev
Quast C, Pruesse E, Yilmaz P, Gerken J, Schweer T, Yarza P, Peplies J, Glöckner FO (2013) The SILVA ribosomal RNA gene database project: Improved data processing and web-based tools. Nucleic Acids Res. https://doi.org/10.1093/nar/gks1219
Rausch P, Rühlemann M, Hermes BM et al (2019) Comparative analysis of amplicon and metagenomic sequencing methods reveals key features in the evolution of animal metaorganisms. Microbiome. https://doi.org/10.1186/s40168-019-0743-1
Reimer PJ, Bard E, Bayliss A et al (2013) IntCal13 and Marine13 Radiocarbon Age Calibration Curves 0–50,000 Years cal BP. Radiocarbon. https://doi.org/10.2458/azu_js_rc.55.16947
Reyes C, Schneider D, Lipka M, Thürmer A, Böttcher ME, Friedrich MW (2017) Nitrogen Metabolism Genes from Temperate Marine Sediments. Mar Biotechnol. https://doi.org/10.1007/s10126-017-9741-0
Robinson RS, Kienast M, Luiza Albuquerque A et al (2012) A review of nitrogen isotopic alteration in marine sediments. Paleoceanography. https://doi.org/10.1029/2012PA002321
Rodysill JR, Russell JM, Bijaksana S, Brown ET, Safiuddin LO, Eggermont H (2012) A paleolimnological record of rainfall and drought from East Java, Indonesia during the last 1,400 years. J Paleolimnol. https://doi.org/10.1007/s10933-011-9564-3
Rodysill JR, Russell JM, Vuille M, Dee S, Lunghino B, Bijaksana S (2019) La Niña-driven flooding in the Indo-Pacific warm pool during the past millennium. Quat Sci Rev 225:106020. https://doi.org/10.1016/j.quascirev.2019.106020
Rognes T, Flouri T, Nichols B, Quince C, Mahé F (2016) VSEARCH: A versatile open source tool for metagenomics. PeerJ. https://doi.org/10.7717/peerj.2584
Saenger C, Miller M, Smittenberg RH, Sachs JP (2006) A physico-chemical survey of inland lakes and saline ponds: Christmas Island (Kiritimati) and Washington (Teraina) Islands. Republic of Kiribati. Saline Syst. https://doi.org/10.1186/1746-1448-2-8
Schmitt S, Conroy JL, Flynn TM, Sanford RA, Higley MC, Chen M, Fouke BW (2019) Salinity, microbe and carbonate mineral relationships in brackish and hypersaline lake sediments: a case study from the tropical Pacific coral atoll of Kiritimati. Depos Rec. https://doi.org/10.1002/dep2.71
Schneider D, Arp G, Reimer A, Reitner J, Daniel R (2013) Phylogenetic analysis of a microbialite-forming microbial mat from a hypersaline lake of the Kiritimati Atoll, Central Pacific. PLoS One. https://doi.org/10.1371/journal.pone.0066662
Sigman DM, Karsh KL, Casciotti KL (2009) Nitrogen Isotopes in the Ocean. In: Encyclopedia of Ocean Sciences
Spring S, Brinkmann N, Murrja M, Spröer C, Reitner J, Klenk HP (2015) High diversity of culturable prokaryotes in a lithifying hypersaline. Microbial Mat Geomicrobiol J. https://doi.org/10.1080/01490451.2014.913095
Talbot MR (2001) Nitrogen Isotopes in Palaeolimnology. In: Last WM, Smol JP (eds) Tracking Environmental Change Using Lake Sediments: Physical and Geochemical Methods. Springer, Netherlands, pp 401–439
Trichet J, Défarge C, Tribble J, Tribble G, Sansone F (2001) Christmas Island lagoonal lakes, models for the deposition of carbonate-evaporite-organic laminated sediments. Sediment Geol. https://doi.org/10.1016/S0037-0738(00)00177-9
Wong HL, White RA, Visscher PT, Charlesworth JC, Vázquez-Campos X, Burns BP (2018) Disentangling the drivers of functional complexity at the metagenomic level in Shark Bay microbial mat microbiomes. ISME J. https://doi.org/10.1038/s41396-018-0208-8
Wuthrich D (2006) Google Earth Pro. Geospatial Solut 16:30
Xu L, Zeng XC, Nie Y, Luo X, Zhou E, Zhou L, Pan Y, Li W (2014) Pontibacter diazotrophicus sp nov, a novel nitrogen-fixing bacterium of the family Cytophagaceae. PLoS One. https://doi.org/10.1371/journal.pone.0092294
Acknowledgements
This research was funded by ACS-PRF 57417-DNI2 and NSF-EAR 1602590 to JLC. We thank the UIUC Roy J. Carver Biotechnology Center, specifically the High-throughput Sequencing and Genotyping Unit, for DNA sample analysis. We thank A. Wyman, M. Higley, and C. Karamperidou for field assistance and B. Fouke and M. Christie for useful comments and advice.
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Chen, M., Conroy, J.L., Sanford, R.A. et al. Interpreting lacustrine bulk sediment δ15N values using metagenomics in a tropical hypersaline lake system. J Paleolimnol 65, 151–168 (2021). https://doi.org/10.1007/s10933-020-00157-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10933-020-00157-7