Abstract
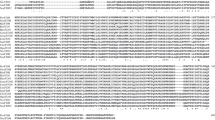
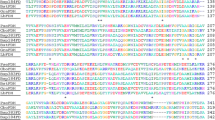
NAD-dependent formate dehydrogenase (FDH) enzymes are frequently used in industrial and scientific applications. FDH is a reversible enzyme that reduces the NAD molecule to NADH and produces CO2 by oxidation of the formate ion, whereas it causes CO2 reduction in the reverse reaction. Some transition metal elements – Fe3+, Mo6+ and W6 + – can be found in the FDH structure of anaerobic and archaeal microorganisms, and these enzymes require cations and other redox-active cofactors for their FDH activity. While NAD-dependent FDHs do not necessarily require any metal cations, the presence of various metal cations can still affect FDH activities. To study the effect of 11 different metal ions, NAD-dependent FDH enzymes from ten different microorganisms were tested: Ancylobacter aquaticus (AaFDH), Candida boidinii (CboFDH), Candida methylica (CmFDH), Ceriporiopsis subvermispora (CsFDH), Chaetomium thermophilum (CtFDH), Moraxella sp. (MsFDH), Myceliophthora thermophila (MtFDH), Paracoccus sp. (PsFDH), Saccharomyces cerevisiae (ScFDH) and Thiobacillus sp. (TsFDH). It was found that metal ions (mainly Cu2+ and Zn2+) could have quite strong inhibition effects on several enzymes in the forward reaction, whereas several cations (Li+, Mg2+, Mn2+, Fe3+ and W6+) could increase the forward reaction of two FDHs. The highest activity increase (1.97 fold) was caused by Fe3+ in AaFDH. The effect on the reverse reaction was minimal. The modelled structures of ten FDHs showed that the active site is formed by 15 highly conserved amino acid residues spatially settling around the formate binding site in a conserved way. However, the residue differences at some of the sites close to the substrate do not explain the activity differences. The active site space is very tight, excluding water molecules, as observed in earlier studies. Structural examination indicated that smaller metal ions might be spaced close to the active site to affect the reaction. Metal ion size showed partial correlation to the effect on inhibition or activation. Affinity of the substrate may also affect the sensitivity to the metal’s effect. In addition, amino acid differences on the protein surface may also be important for the metal ion effect.




Similar content being viewed by others
Abbreviations
- AaFDH:
-
Ancylobacter aquaticus FDH
- CboFDH:
-
Candida boidinii FDH
- CmFDH:
-
Candida methylica FDH
- CsFDH:
-
Ceriporiopsis subvermispora FDH
- CtFDH:
-
Chaetomium thermophilum FDH
- IPTG:
-
Isopropyl β-D-1-thiogalactopyranoside
- KDa:
-
Kilodalton
- mM:
-
Millimolar
- MsFDH:
-
Moraxella sp. FDH
- MtFDH:
-
Myceliopthora thermophila FDH
- NAD:
-
Nicotinamide adenine dinucleotide
- NADH:
-
Nicotinamide adenine dinucleotide hydrogen
- PsFDH:
-
Paracoccus sp. FDH
- SceFDH:
-
Saccharomyces cerevisiae FDH
- TsFDH:
-
Thiobacillus sp. FDH
- μM:
-
Micromolar
References
Zhushan F, Shuhua X (2020) The effects of heavy metals on human metabolism. Toxicol Mech Methods 30:3
Popov VO and Lamzin V. S (1994) NAD+-dependent formate dehydrogenase. Biochem J 301:625–643
Tishkov VI and Popov V. O (2006) Protein engineering of formate dehydrogenase. Biomol Eng 23:89–110
Altinkaynak C, Gulmez C, Atakisi O, Özdemir N (2020) Evaluation of organic-inorganic hybrid nanoflower’s enzymatic activity in the presence of different metal ions and organic solvents. Int J Biol Macromol 164:162–171
Ballinger P, Long FA (1960) Acid Ionization Constants of Alcohols. II. Acidities of Some Substituted Methanols and Related Compounds. J Am Chem Soc 82:795–798
Tishkov VI and Popov V. O (2004) Catalytic mechanism and application of formate dehydrogenase. Biochemistry 69:1252–1267
Choe H, Joo JC, Cho DH, Kim MH, Lee SH, Jung KD, Kim YH (2014) Efficient CO2-reducing activity of NAD-dependent formate dehydrogenase from Thiobacillus sp. KNK65MA for formate production from CO2 gas. PLoS One 25(7):e103111 9(
Jayathilake BS, Bhattacharya S, Vaidehi N, Narayanan SR (2019) Efficient and Selective Electrochemically Driven Enzyme-Catalyzed Reduction of Carbon Dioxide to Formate using Formate Dehydrogenase and an Artificial Cofactor. Acc Chem Res 52:676–685
Aslan AS, Valjakka J, Ruupunen J, Yildirim D, Turner NJ, Turunen O, Binay B (2017) Chaetomium thermophilum formate dehydrogenase has high activity in the reduction of hydrogen carbonate (HCO3–) to formate. Protein Eng Des Sel 30:47–55
Pietzke M, Meiser J, Vazquez A (2020) Formate metabolism in health and disease. Molecular Metabolism 33:23–37
Hovda KE, Urdal P, Jacobsen D (2005) Increased serum formate in the diagnosis of methanol poisoning. J Anal Toxicol 29:586–588
Hantson P, Haufroid V, Wallemacq P (2005) Formate kinetics in methanol poisoning. Hum Exp Toxicol. 24(2):55–9
Onyekwere N, Nwadiuto I, Maleghemi S, Maduka O, Numbere TW, Akpuh N, Kanu E, Katchy I, Okeafor I (2018) Methanol poisoning in South-South Nigeria: Reflections on the outbreak response. J Public Health Africa 9:49–53
Goldberg SL, Nanduri VB, Chu L, Johnston RM, Patel RN (2006) Enantioselective microbial reduction of 6-oxo-8-[4-[4-(2-pyrimidinyl)-1- piperazinyl]butyl]-8-azaspiro[4.5]decane-7,9-dione. Tetrahedron Asymmetry 16:2778–2783
Coleman JS, Gaydos CA, Witter F (2013) Trichomonas vaginalis vaginitis in obstetrics and gynecology practice: New concepts and controversies. Obstet Gynecol Surv 68:43–50
Janson JC (2012) Protein Purification: Principles, High Resolution Methods, and Applications. John Wiley and Sons
Altas N, Aslan AS, Karatas E, Chronopoulou E, Labrou NE, Binay B (2017) Heterologous production of extreme alkaline thermostable NAD+-dependent formate dehydrogenase with wide-range pH activity from Myceliophthora thermophila. Process Biochem 61:110–118
Pala U, Yelmazer B, Çorbacıoglu M, Ruupunen J, Valjakka J, Turunen O, Binay B (2018) Functional effects of active site mutations in NAD+-dependent formate dehydrogenases on transformation of hydrogen carbonate to formate. Protein Eng Des Sel 31:327–335
Lamzin VS, Dauter Z, Popov VO, Harutyunyan EH, Wilson KS (1994) High resolution structures of holo and apo formate dehydrogenase. JMolBiol 236:759–785
Guo Q, Gakhar L, Wickersham K, Francis K, Vardi-Kilshtain A, Major DT, Cheatum CM, Kohen A (2016) Structural and Kinetic Studies of Formate Dehydrogenase from. Candida boidiniiBiochemistry 55:2760–2771
Shabalin IG, Polyakov KM, Serov AE, Skirgello OE, Sadykhov EG, Dorovatovskiy PV, Tishkov VI, Popov VO (2010) NAD-dependent formate dehydrogenase from higher-plant Arabidopsis thaliana in complex with NAD and azide. PDB Databank
Filippova EV, Polyakov KM, Tikhonova TV, Sadykhov IG, Shabalin IG, Tishkov VI, Popov VO (2009) NAD-dependent formate dehydrogenase from bacterium Moraxella sp.C2 in complex with NAD and azide. Acta Crystallogr D Biol Crystallogr 65:1315–1325
Yoch DC, Chen YP, Hardin MG (1990) Formate dehydrogenase from the methane oxidizer Methylosinus trichosporium OB3b. J Bacteriol 172:4456–4463
Ding HT, Liu DF, Li ZL, Du YQ, Xu XH, Zhao YH (2011) Characterization of a thermally stable and organic solvent-adaptative NAD +-dependent formate dehydrogenase from Bacillus sp. F1. J Appl Microbiol 111:1075–1085
Robinson WE, Bassegoda A, Reisner E, Hirst J (2017) Oxidation-State-Dependent Binding Properties of the Active Site in a Mo-Containing Formate Dehydrogenase. J Am Chem Soc 139:9927–9936
Schlapbach L, Züttel A (2010) Hydrogen-storage materials for mobile applications. in Materials for Sustainable Energy: A Collection of Peer-Reviewed Research and Review Articles from Nature Publishing Group 265–270
Srikanth S, Alvarez-Gallego Y, Vanbroekhoven K, Pant D (2017) Enzymatic Electrosynthesis of Formic Acid through Carbon Dioxide Reduction in a Bioelectrochemical System: Effect of Immobilization and Carbonic Anhydrase Addition. ChemPhysChem 18:3174–3181
Liu A, Feng R, Liang B (2016) Microbial surface displaying formate dehydrogenase and its application in optical detection of formate. Enzyme Microb Technol 91:59–65
Yu S, Zhu L, Zhou C, An T, Zhang T, Jiang B, Mu W (2014) Promising properties of a formate dehydrogenase from a methanol- assimilating yeast Ogataea parapolymorpha DL-1 in His-tagged form. Appl Microbiol Biotechnol 98:1621–1630
Shannon RD (1976) Revised effective ionic radii and systematic studies of interatomic distances in halides and chalcogenides. Acta Crystallogr Sect A 32:751–767
Shabalin IG, Filippova EV, Polyako KM, Sadykhov EG, Safonova TN, Tikhonova TV, Tishkov VI, Popov VO (2009) Structures of the apo and holo forms of formate dehydrogenase from the bacterium Moraxella sp. C-1: towards understanding the mechanism of the closure of the interdomain cleft. Acta Crystallogr SectD 65:1315–1325
Madej T, Lanczycki CJ, Zhang D, Thiessen PA, Geer RC, Marchler-Bauer A, Bryant SH (2014) MMDB and VAST+: tracking structural similarities between macromolecular complexes. Nucleic Acids 42:297–303
Pagano P, Guo Q, Kohen A, and Christopher M (2016) Oscillatory Enzyme Dynamics Revealed by Two-Dimensional Infrared Spectroscopy. J Phys Chem Lett 7(13):2507–2511 7 )
Çakar MM, Mangas-Sanchez J, Birmingham WR, Turner NJ, Binay B (2018) Discovery of a new metal and NAD+-dependent formate dehydrogenase from Clostridium ljungdahlii. Prep Biochem Biotechnol 48:327–334
Karimäki J, Parkkinen T, Santa H, Pastinen O, Leisola M, Rouvinen J, Turunen O (2004) Engineering the substrate specificity of xylose isomerase. Protein Eng Des Sel 17:861–869
Santa H, Kammonen J, Karimäki J, Leisola M, Turunen O (2005) Stochastic boundary molecular dynamics simulation of L-ribose in the active site of Actinoplanes missouriensis xylose isomerase and its Val135Asn mutant with improved reaction rate. Biochim Biophys Acta - Proteins Proteomics 1749:65–73
Acknowledgements
Special thanks to Gebze Technical University Enzyme Test and Consultancy Center for providing infrastructure during the research.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no conflicts of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Bulut, H., Valjakka, J., Yuksel, B. et al. Effect of Metal Ions on the Activity of Ten NAD-Dependent Formate Dehydrogenases. Protein J 39, 519–530 (2020). https://doi.org/10.1007/s10930-020-09924-x
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10930-020-09924-x