Abstract
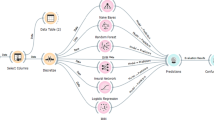
Proteins class and function prediction is one of the most significant task in computational bioinformatics. The information about the protein functions and class plays a vital role in understanding biological cells and has a great impact on human life in factors such as personalized medicine. The technical advancement in the areas of biological aspects and understanding of biological processes results in features and characteristics of important Proteins. Prediction of amino acid sequence involves prediction of amino sequence folding and its structures from the primary sequence obtained. In this work, Machine learning prediction algorithms have applied for protein class prediction. This method takes consideration of macromolecules of biological significances. Later the solution focuses on the understanding of different protein family, subsequently classify the protein family type sequence. This is achieved through machine learning algorithms Naive Bayes (NB) and Random forest (RF) algorithms with count vectorized feature and LSTM. These algorithms are used to classify the protein family on its protein sequence. Finally, result shows that LSTM predicts the protein class more accurately than the RF, and NB algorithm. LSTM achieves an accuracy of 96% whereas RF & NB with an accuracy of 91% and 86%.









Similar content being viewed by others
References
Pauling L, Corey RB, Branson HR (1951) The structure of proteins; two hydrogen-bonded helical configurations of the polypeptide chain. Proc Natl Acad Sci USA 37:205
Rehman HU, Azam N, Yao J, Benso A (2017) A three-way approach for protein function classification. PLoS ONE 12(2):0171702
Kabli F, Hamou RM, Amine A (2017) New classification system for protein sequences. In 2017 First International Conference on Embedded and Distributed Systems (EDiS), IEEE. Oran, Algeria, pp. 1–6
Bankapur, Sanjay, and Nagamma Patil (2018) Protein Secondary Structural Class Prediction Using Effective Feature Modeling and Machine Learning Techniques. In 2018 IEEE 18th International Conference on Bioinformatics and Bioengineering (BIBE). IEEE pp.18–21
Lima, Emerson Correia, Fábio Lima Custódio, Gregório Kappaun Rocha, and Laurent E. Dardenne (2018) Estimating Protein Structure Prediction Models Quality Using Convolutional Neural Networks. In 2018 International Joint Conference on Neural Networks (IJCNN), IEEE pp. 1–6
Fang, Chao, Yi Shang, and Dong Xu. (2017) A New Deep Neighbor Residual Network for Protein Secondary Structure Prediction. In 2017 IEEE 29th International Conference on Tools with Artificial Intelligence (ICTAI). IEEE pp. 66–71
Iqbal MJ, Faye I, Said AM, Samir BB (2014) Data mining of protein sequences with amino acid position-based feature encoding technique. In: Herawan T, Deris MM, Abawajy J (eds) Proceedings of the First International Conference on Advanced Data and Information Engineering. Springer, Singapore
Anfinsen C (1972) The formation and stabilization of protein structure. Biochem J 128:737
Dictionary (2019) Amino. https://www.dictionary.com/. Accessed 25 March 2019
Amino acid, [Online]. Available: https://en.wikipedia.org/. Accessed 22 May 2015
Robles V, Larrañaga P, Peña JM, Menasalvas E, Pérez MS, Herves V, Wasilewska A (2004) Bayesian network multi-classifiers for protein secondary structure prediction. Artif Intell Med 31:117
Breiman L (2001) Random forests. Mach Learn 45(1):5–32
Protein data bank. Availabe https://www.kaggle.com/shahir/protein-data-set#pdb_data_seq.csv
Hochreiter S, Schmidhuber J (1997) Long short-term memory. Neural Comput 9(8):1735–1780
Hawkins J, Boden M (2005) The Applicability of recurrent neural networks for biological sequence analysis. IEEE/ACM Trans Comput Biol Bioinform 2(3):243–253
Jain G, Sharma M, Agarwal B (2019) Optimizing semantic LSTM for spam detection. Int J Inf Technol 11:239–250
Chhachhiya D, Sharma A, Gupta M (2019) Designing optimal architecture of recurrent neural network (LSTM) with particle swarm optimization technique specifically for educational dataset. Int J Inf Technol 11(1):159–163
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Sekhar, S.R.M., Siddesh, G.M., Raj, M. et al. Protein class prediction based on Count Vectorizer and long short term memory. Int. j. inf. tecnol. 13, 341–348 (2021). https://doi.org/10.1007/s41870-020-00528-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s41870-020-00528-3