Abstract
Purpose
This contribution is the first example of a small molecule pharmaceutical product development ontology. This ontology allows the portfolio-wide visualization of the small molecule pharmaceutical development business as a network of decisions and activities. This ontology was built not only to digitalize, rapidly access, and gain insights from prior decision-making but also to deliver input data for dynamic resource allocation via optimal scheduling of research activities subject to resource, cost, and time constraints.
Methods
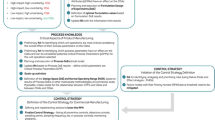
This ontology can be understood as a generic wiring diagram for small molecule product development which outlines the predecessors and successors of activities and decisions, along with needed or allowed overlap between activities. Each activity has multiple modes with direct dollars, resource consumption, and time taken to complete each mode.
Results
This generic wiring diagram was instanced for 17 portfolio assets by creating 32 specific scenarios. These scenarios were stored in an ontology as instances and served as foundational digitalized data for a host of future applications. These specific scenarios across the multiple assets served as input to the optimal resource allocation model.
Conclusion
In summary, this work describing pharmaceutical Chemistry Manufacturing and Controls (CMC) development activities and decisions is a foundational transformative project that enables full digitization of pharmaceutical CMC data, not only for optimal scheduling but also to generate heuristics on selection of research activities given development risks, a topic which has received virtually no mention in literature.








Similar content being viewed by others
References
CDISC. https://www.cdisc.org/standards.
Dali M, Stewart A, Behling RW, Raglione T, Stamato HJ, Tom JW. Optimizing knowledge creation at Bristol-Myers Squibb—a case study within pharmaceutical development. J Pharm Innov. 2015;10(1):1–12.
Gene Ontology. http://geneontology.org/docs/ontology-documentation/.
Remolona MFM, Conway MF, Balasubramanian S, Fan L, Feng Z, Gu T, et al. Hybrid ontology-learning materials engineering system for pharmaceutical products: multi-label entity recognition and concept detection. Comput Chem Eng. 2017;107:49–60.
Suresh P, Hsu S-H, Reklaitis GV, Venkatasubramanian VJI, Research EC. OntoMODEL: ontological mathematical modeling knowledge management in pharmaceutical product development, 2: applications. 2010;49(17):7768–81.
Venkatasubramanian V, Zhao C, Joglekar G, Jain A, Hailemariam L, Suresh P, et al. Ontological informatics infrastructure for pharmaceutical product development and manufacturing. 2006;30(10–12):1482–96.
Sankar P, Aghila G. Design and development of chemical ontologies for reaction representation. 2006;46(6):2355–68.
Takahashi L, Miyazato I, Takahashi K. Redesigning the Materials and Catalysts Database Construction Process Using Ontologies. 2018;58(9):1742–54.
Papadakis E, Anantpinijwatna A, Woodley J, Gani RJP. A reaction database for small molecule pharmaceutical processes integrated with process information. 2017;5(4):58.
Foundation A. https://www.allotrope.org/.
Hua Wang JD, Guntz Steve, Vaidyaraman Shankarraman, Viswanath Shekhar, Gounaris Chrysanthos E. Portfolio-wide optimization of pharmaceutical R&D activities using mathematical programming. (Under Review).
Pharmaceutical development, Q8 (2009).
Quality risk management, Q9 (2006).
Pharmaceutical quality system (2009).
Development and manufacture of drug substances, Q11 (2012).
Hartmann S, Briskorn D. A survey of variants and extensions of the resource-constrained project scheduling problem. J Eur J Oper Res. 2010;207(1):1–14. https://doi.org/10.1016/j.ejor.2009.11.005.
Ho D. Addressing COVID-19 drug development with artificial intelligence. 2020;2(5):2000070.
Benny P, Nair SK, Krishnakumar K, Dineshkumar B. Artificial intelligence in drug discovery and development.
RSC’s RXNO Ontology. https://bioportal.bioontology.org/ontologies/RXNO.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of Interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Viswanath, S., Guntz, S., Dieringer, J. et al. An Ontology to Describe Small Molecule Pharmaceutical Product Development and Methodology for Optimal Activity Scheduling. J Pharm Innov 17, 155–169 (2022). https://doi.org/10.1007/s12247-020-09505-6
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12247-020-09505-6