Abstract
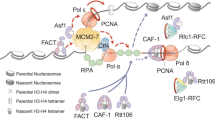
Replication-coupled (RC) nucleosome assembly is an essential process in eukaryotic cells to maintain chromatin structure during DNA replication. The deposition of newly-synthesized H3/H4 histones during DNA replication is facilitated by specialized histone chaperones. CAF-1 is an important histone chaperone complex and its main subunit, Cac1p, contains a PIP and WHD domain for interaction with PCNA and the DNA, respectively. While Cac1p subunit was extensively studied in different systems much less is known regarding the importance of the PIP and WHD domains in replication fork progression and genome stability. By exploiting a time-lapse microscopy system for monitoring DNA replication in individual live cells, we examined how mutations in these Cac1p domains affect replication fork progression and post-replication characteristics. Our experiments revealed that mutations in the Cac1p WHD domain, which abolished the CAF-1–DNA interaction, slows down replication fork progression. In contrast, mutations in Cac1p PIP domain, abolishing Cac1p–PCNA interaction, lead to extended late-S/Anaphase duration, elevated number of RPA foci and increased spontaneous mutation rate. Our research shows that Cac1p WHD and PIP domains have distinct roles in high replisome progression and maintaining genome stability during cell cycle progression.



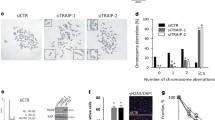
adapted from Fig. 2 and are presented for comparison with the cac1 PIP and WHD mutants. Significance was determined by Monte Carlo resampling and p values relative to WT are shown. The number of cells analyzed for each strain and all median values and statistical results are shown in Table S1



Similar content being viewed by others
References
Alabert C, Groth A (2012) Chromatin replication and epigenome maintenance. Nat Rev Mol Cell Biol 13:153–167. https://doi.org/10.1038/nrm3288
Alvino GM, Collingwood D, Murphy JM et al (2007) Replication in hydroxyurea: it’s a matter of time. Mol Cell Biol 27:6396–6406. https://doi.org/10.1128/MCB.00719-07
Amon A (1999) The spindle checkpoint. Curr Opin Genet Dev 9:69–75. https://doi.org/10.1016/S0959-437X(99)80010-0
Andrews AJ, Luger K (2011) Nucleosome structure(s) and stability: variations on a theme. Annu Rev Biophys 40:99–117. https://doi.org/10.1146/annurev-biophys-042910-155329
Boeri-Erba E, Ochsenbein F, Sitbon D et al (2017) Insights into the molecular architecture and histone H3–H4 deposition mechanism of yeast Chromatin assembly factor 1. Elife. https://doi.org/10.7554/elife.23474
Clemente-Ruiz M, González-Prieto R, Prado F (2011) Histone H3K56 acetylation, CAF1, and Rtt106 coordinate nucleosome assembly and stability of advancing replication forks. PLoS Genet. https://doi.org/10.1371/journal.pgen.1002376
Dahan D, Tsirkas I, Dovrat D et al (2018) Pif1 is essential for efficient replisome progression through lagging strand G-quadruplex DNA secondary structures. Nucleic Acids Res. https://doi.org/10.1093/nar/gky1065
Dovrat D, Dahan D, Sherman S et al (2018) A live-cell imaging approach for measuring DNA replication rates. Cell Rep 24:252–258. https://doi.org/10.1016/j.celrep.2018.06.018
Falbo KB, Shen X (2006) Chromatin remodeling in DNA replication. J Cell Biochem 97:684–689. https://doi.org/10.1002/jcb.20752
Franco AA, Lam WM, Burgers PM, Kaufman PD (2005) Histone deposition protein Asf1 maintains DNA replisome integrity and interacts with replication factor C. Genes Dev. https://doi.org/10.1101/gad.1305005
Fridman Y, Palgi N, Dovrat D et al (2010) Subtle alterations in PCNA-partner interactions severely impair DNA replication and repair. PLoS Biol 8:e1000507. https://doi.org/10.1371/journal.pbio.1000507
Game JC, Kaufman PD (1999) Role of Saccharomyces cerevisiae chromatin assembly factor-I in repair of ultraviolet radiation damage in vivo. Genetics 151:485–497
Gurard-Levin ZA, Quivy J-P, Almouzni G (2014) Histone chaperones: assisting histone traffic and nucleosome dynamics. Annu Rev Biochem 83:487–517. https://doi.org/10.1146/annurev-biochem-060713-035536
Hammond CM, Strømme CB, Huang H et al (2017) Histone chaperone networks shaping chromatin function. Nat Rev Mol Cell Biol 18:141–158. https://doi.org/10.1038/nrm.2016.159
Han J, Zhou H, Li Z et al (2007) Acetylation of Lysine 56 of Histone H3 catalyzed by RTT109 and regulated by ASF1 is required for replisome integrity. J Biol Chem 282:28587–28596. https://doi.org/10.1074/jbc.M702496200
Hedglin M, Benkovic SJ (2015) Regulation of Rad6/Rad18 activity during DNA damage tolerance. Annu Rev Biophys. https://doi.org/10.1146/annurev-biophys-060414-033841
Hendel A, Krijger PHL, Diamant N et al (2011) PCNA ubiquitination is important, but not essential for translesion DNA synthesis in mammalian cells. PLoS Genet. https://doi.org/10.1371/journal.pgen.1002262
Hoek M, Stillman B (2003) Chromatin assembly factor 1 is essential and couples chromatin assembly to DNA replication in vivo. Proc Natl Acad Sci 100:12183–12188. https://doi.org/10.1073/pnas.1635158100
Huang S, Zhou H, Katzmann D et al (2005) Rtt106p is a histone chaperone involved in heterochromatin-mediated silencing. Proc Natl Acad Sci U S A 102(38):13410–13415
Huang TH, Fowler F, Chen CC et al (2018) The histone chaperones ASF1 and CAF-1 promote MMS22L-TONSL-mediated Rad51 loading onto ssDNA during homologous recombination in human cells. Mol Cell 69:879–892.e5. https://doi.org/10.1016/j.molcel.2018.01.031
Ito H, Fukuda Y, Murata K, Kimura A (1983) Transformation of intact yeast cells treated with alkali cations. J Bacteriol 153:163–168
Kaufman PD, Kobayashi R, Kessler N, Stillman B (1995) The p150 and p60 subunits of chromatin assembly factor I: a molecular link between newly synthesized histories and DNA replication. Cell 81:1105–1114. https://doi.org/10.1016/S0092-8674(05)80015-7
Krawitz DC, Kama T, Kaufman PD (2002) Chromatin assembly factor I mutants defective for PCNA binding require Asf1/Hir proteins for silencing. Mol Cell Biol 22:614–625
Lea DE, Coulson CA (1949) The distribution of the numbers of mutants in bacterial populations. J Genet 49:264–285. https://doi.org/10.1007/BF02986080
Li Q, Zhou H, Wurtele H et al (2008) Acetylation of histone H3 lysine 56 regulates replication-coupled nucleosome assembly. Cell 134:244–255. https://doi.org/10.1016/j.cell.2008.06.018
Li Q, Burgess R, Zhang Z (2012) All roads lead to chromatin: Multiple pathways for histone deposition. Biochim Biophys Acta - Gene Regul Mech 1819:238–246. https://doi.org/10.1016/j.bbagrm.2011.06.013
Lisby M, Barlow JH, Burgess RC, Rothstein R (2004) Choreography of the DNA damage response: spatiotemporal relationships among checkpoint and repair proteins. Cell 118:699–713. https://doi.org/10.1016/j.cell.2004.08.015
Liu WH, Roemer SC, Zhou Y et al (2016) The Cac1 subunit of histone chaperone CAF-1 organizes CAF-1-H3/H4 architecture and tetramerizes histones. Elife. https://doi.org/10.7554/eLife.18023
Mattiroli F, D’Arcy S, Luger K (2015) The right place at the right time: chaperoning core histone variants. EMBO Rep 16:1454–1466. https://doi.org/10.15252/embr.201540840
Mattiroli F, Gu Y, Balsbaugh JL et al (2017a) The Cac2 subunit is essential for productive histone binding and nucleosome assembly in CAF-1. Sci Rep 7:1–11. https://doi.org/10.1038/srep46274
Mattiroli F, Gu Y, Yadav T et al (2017b) DNA-mediated association of two histone-bound complexes of yeast Chromatin Assembly Factor-1 (CAF-1) drives tetrasome assembly in the wake of DNA replication. Elife. https://doi.org/10.7554/eLife.22799
Mattiroli F, Gu Y, Yadav T et al (2017c) DNA-mediated association of two histone-bound complexes of yeast chromatin assembly factor-1 (CAF-1) drives tetrasome assembly in the wake of DNA replication. Elife 6:1–23. https://doi.org/10.7554/eLife.22799
Moldovan G-L, Pfander B, Jentsch S (2007) PCNA, the maestro of the replication fork. Cell 129:665–679. https://doi.org/10.1016/j.cell.2007.05.003
Monson EK, De Bruin D, Zakian VA (1997) The yeast Cac1 protein is required for the stable inheritance of transcriptionally repressed chromatin at telomeres. Proc Natl Acad Sci U S A 94:13081–13086. https://doi.org/10.1073/pnas.94.24.13081
Myung K, Pennaneach V, Kats ES, Kolodner RD (2003) Saccharomyces cerevisiae chromatin-assembly factors that act during DNA replication function in the maintenance of genome stability. Proc Natl Acad Sci 100:6640–6645. https://doi.org/10.1073/pnas.1232239100
Panne D, Sauer PV, Gu Y et al (2018) Mechanistic insights into histone deposition and nucleosome assembly by the chromatin assembly factor-1. Nucleic Acids Res 46:9907–9917. https://doi.org/10.1093/nar/gky823
Papamichos-Chronakis M, Peterson CL (2012) Chromatin and the genome integrity network. Nat Rev Genet 14:62–75. https://doi.org/10.1038/nrg3345
Pardo B, Crabbé L, Pasero P (2017) Signaling pathways of replication stress in yeast. FEMS Yeast Res. https://doi.org/10.1093/femsyr/fow101
Plate I, Hallwyl SCL, Shi I et al (2008) Interaction with RPA is necessary for Rad52 repair center formation and for its mediator activity. J Biol Chem 283:29077–29085. https://doi.org/10.1074/jbc.M804881200
Prado F, Maya D (2017) Regulation of replication fork advance and stability by nucleosome assembly. Genes (Basel). https://doi.org/10.3390/genes8020049
Sanchez Y, Bachant J, Wang H et al (1999) Control of the DNA damage checkpoint by Chk1 and Rad53 protein kinases through distinct mechanisms. Science 286:1166–1171. https://doi.org/10.1126/science.286.5442.1166
Selth L, Svejstrup JQ (2007) Vps75, a new yeast member of the NAP histone chaperone. J Biol Chem 282:12358–12362. https://doi.org/10.1074/jbc.C700012200
Serra-Cardona A, Zhang Z (2018) Replication-coupled nucleosome assembly in the passage of epigenetic information and cell identity. Trends Biochem Sci 43:136–148. https://doi.org/10.1016/j.tibs.2017.12.003
Sharp JA, Franco AA, Osley MA, Kaufman PD (2002) Chromatin assembly factor I and Hir proteins contribute to building functional kinetochores in S. cerevisiae. Genes Dev 16:85–100. https://doi.org/10.1101/gad.925302
Soreanu I, Hendler A, Dahan D et al (2018) Marker-free genetic manipulations in yeast using CRISPR/CAS9 system. Curr Genet 64:1129–1139. https://doi.org/10.1007/s00294-018-0831-y
Su D, Hu Q, Li Q et al (2012) Structural basis for recognition of H3K56-acetylated histone H3–H4 by the chaperone Rtt106. Nature 483:104–109. https://doi.org/10.1038/nature10861
Ulrich HD, Walden H (2010) Ubiquitin signalling in DNA replication and repair. Nat Rev Mol Cell Biol 117(11):479. https://doi.org/10.1038/nrm2921
Verreault A, Kaufman PD, Kobayashi R, Stillman B (1996) Nucleosome assembly by a complex of CAF-1 and acetylated histones H3/H4. Cell 87:95–104. https://doi.org/10.1016/S0092-8674(00)81326-4
Weill U, Yofe I, Sass E et al (2018) Genome-wide SWAp-Tag yeast libraries for proteome exploration. Nat Methods 15:617–622. https://doi.org/10.1038/s41592-018-0044-9
Weinert TA, Kiser GL, Hartwell LH (1994) Mitotic checkpoint genes in budding yeast and the dependence of mitosis on DNA replication and repair. Genes Dev 8:652–665. https://doi.org/10.1101/gad.8.6.652
Wellinger RE, Prado F, Aguilera A (2006) Replication fork progression is impaired by transcription in hyperrecombinant yeast cells lacking a functional THO complex. Mol Cell Biol 26:3327–3334. https://doi.org/10.1128/MCB.26.8.3327
Wong RP, García-Rodríguez N, Zilio N et al (2020) Processing of DNA polymerase-blocking lesions during genome replication is spatially and temporally segregated from replication forks. Mol Cell 77:3–16.e4. https://doi.org/10.1016/j.molcel.2019.09.015
Zeman MK, Cimprich KA (2014) Causes and consequences of replication stress. Nat Cell Biol 16:2–9. https://doi.org/10.1038/ncb2897
Zhang K, Gao Y, Li J et al (2016) A DNA binding winged helix domain in CAF-1 functions with PCNA to stabilize CAF-1 at replication forks. Nucleic Acids Res 44:5083–5094. https://doi.org/10.1093/nar/gkw106
Zheng Q (2017) rSalvador: an R package for the fluctuation experiment. G3 (Bethesda) 7:3849–3856. https://doi.org/10.1534/g3.117.300120
Zunder RM, Antczak AJ, Berger JM, Rine J (2011) Two surfaces on the histone chaperone Rtt106 mediate histone binding, replication, and silencing. Proc Natl Acad Sci 109:E144–E153. https://doi.org/10.1073/pnas.1119095109
Acknowledgements
This work was supported by the Israeli Science foundation (ISF) grant numbers 1340/17 and 2359/18, the German-Israeli Foundation (GIF) grant number 1-1425-415.13/2017, the Binational Science Foundation (BSF-NSF) grant number 2019617 and the European research training network (ITN, Horizon 2020) ES-cat grant number 722610.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by M. Kupiec.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Tsirkas, I., Dovrat, D., Lei, Y. et al. Cac1 WHD and PIP domains have distinct roles in replisome progression and genomic stability. Curr Genet 67, 129–139 (2021). https://doi.org/10.1007/s00294-020-01113-8
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00294-020-01113-8