Abstract
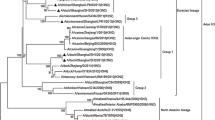
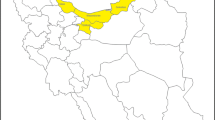
To date, avian influenza viruses (AIVs) have persisted in domestic poultry in wet markets in East Asian countries. We have performed ongoing virus surveillance in poultry populations in Vietnam since 2011, with the goal of controlling avian influenza. Throughout this study, 110 H3 AIVs were isolated from 2760 swab samples of poultry in markets and duck farms. H3 hemagglutinin (HA) genes of the isolates were phylogenetically classified into eight groups (I–VIII). Genetic diversity was also observed in the other seven gene segments. Groups I–IV also included AIVs from wild waterbirds. The epidemic strains in poultry switched from groups I–III and VI to groups I, IV, V, and VIII around 2013. H3 AIVs in groups I and V were maintained in poultry until at least 2016, which likely accompanied their dissemination from the northern to the southern regions of Vietnam. Groups VI–VIII AIVs were antigenically distinct from the other groups. Some H3 AIV isolates had similar N6 neuraminidase and matrix genes as H5 highly pathogenic avian influenza viruses (HPAIVs). These results reveal that genetically and antigenically different H3 AIVs have been co-circulating in poultry in Vietnam. Poultry is usually reared outside in this country and is at risk of infection with wild waterbird-originating AIVs. In poultry flocks, the intruded H3 AIVs must have experienced antigenic drift/shift and genetic reassortment, which could contribute to the emergence of H5 HPAIVs with novel gene constellations.




Similar content being viewed by others
References
Alexander DJ (1982) Ecological aspects of influenza A viruses in animals and their relationship to human influenza: a review. J R Soc Med 75:799–811
Asha K, Kumar B (2019) Emerging influenza D virus threat: what we know so far! J Clin Med. https://doi.org/10.3390/jcm8020192
Hause BM, Collin EA, Liu R, Huang B, Sheng Z, Lu W, Wang D, Nelson EA, Li F (2014) Characterization of a novel influenza virus in cattle and swine: proposal for a new genus in the orthomyxoviridae family. MBio 5:e00031
Fouchier RA, Munster V, Wallensten A, Bestebroer TM, Herfst S, Smith D, Rimmelzwaan GF, Olsen B, Osterhaus AD (2005) Characterization of a novel influenza A virus hemagglutinin subtype (H16) obtained from black-headed gulls. J Virol 79:2814–2822
Webster RG, Bean WJ, Gorman OT, Chambers TM, Kawaoka Y (1992) Evolution and ecology of influenza A viruses. Microbiol Rev 56:152–179
Wu Y, Wu Y, Tefsen B, Shi Y, Gao GF (2014) Bat-derived influenza-like viruses H17N10 and H18N11. Trends Microbiol 22:183–191
Kida H, Kawaoka Y, Naeve CW, Webster RG (1987) Antigenic and genetic conservation of H3 influenza virus in wild ducks. Virology 159:109–119
Cui H, Shi Y, Ruan T, Li X, Teng Q, Chen H, Yang J, Liu Q, Li Z (2016) Phylogenetic analysis and pathogenicity of H3 subtype avian influenza viruses isolated from live poultry markets in China. Sci Rep 6:27360
Bean WJ, Schell M, Katz J, Kawaoka Y, Naeve C, Gorman O, Webster RG (1992) Evolution of the H3 influenza virus hemagglutinin from human and nonhuman hosts. J Virol 66:1129–1138
Ito K, Igarashi M, Miyazaki Y, Murakami T, Iida S, Kida H, Takada A (2011) Gnarled-trunk evolutionary model of influenza A virus hemagglutinin. PLoS ONE 6:e25953
Laver WG, Air GM, Webster RG (1981) Mechanism of antigenic drift in influenza virus. Amino acid sequence changes in an antigenically active region of Hong Kong (H3N2) influenza virus hemagglutinin. J Mol Biol 145:339–361
Anthony SJ, St Leger JA, Pugliares K, Ip HS, Chan JM, Carpenter ZW, Navarrete-Macias I, Sanchez-Leon M, Saliki JT, Pedersen J, Karesh W, Daszak P, Rabadan R, Rowles T, Lipkin WI (2012) Emergence of fatal avian influenza in New England harbor seals. MBio 3:e00166
Crawford PC, Dubovi EJ, Castleman WL, Stephenson I, Gibbs EP, Chen L, Smith C, Hill RC, Ferro P, Pompey J, Bright RA, Medina MJ, Johnson CM, Olsen CW, Cox NJ, Klimov AI, Katz JM, Donis RO (2005) Transmission of equine influenza virus to dogs. Science 310:482–485
Guo Y, Wang M, Kawaoka Y, Gorman O, Ito T, Saito T, Webster RG (1992) Characterization of a new avian-like influenza A virus from horses in China. Virology 188:245–255
Olsen CW (2002) The emergence of novel swine influenza viruses in North America. Virus Res 85:199–210
Takemae N, Parchariyanon S, Damrongwatanapokin S, Uchida Y, Ruttanapumma R, Watanabe C, Yamaguchi S, Saito T (2008) Genetic diversity of swine influenza viruses isolated from pigs during 2000 to 2005 in Thailand. Influenza Other Respir Viruses 2:181–189
Campitelli L, Fabiani C, Puzelli S, Fioretti A, Foni E, De Marco A, Krauss S, Webster RG, Donatelli I (2002) H3N2 influenza viruses from domestic chickens in Italy: an increasing role for chickens in the ecology of influenza? J Gen Virol 83:413–420
Choi YK, Seo SH, Kim JA, Webby RJ, Webster RG (2005) Avian influenza viruses in Korean live poultry markets and their pathogenic potential. Virology 332:529–537
Liu M, He S, Walker D, Zhou N, Perez DR, Mo B, Li F, Huang X, Webster RG, Webby RJ (2003) The influenza virus gene pool in a poultry market in South central china. Virology 305:267–275
Pu J, Zhang GZ, Ma JH, Xia YJ, Liu QF, Jiang ZL, Wang Z, Brown EG, Tian FL, Liu JH (2009) Serologic evidence of prevalent avian H3 subtype influenza virus infection in chickens. Avian Dis 53:198–204
Choi JG, Kang HM, Kim MC, Paek MR, Kim HR, Kim BS, Kwon JH, Kim JH, Lee YJ (2012) Genetic relationship of H3 subtype avian influenza viruses isolated from domestic ducks and wild birds in Korea and their pathogenic potential in chickens and ducks. Vet Microbiol 155:147–157
Li X, Yang J, Liu B, Jia Y, Guo J, Gao X, Weng S, Yang M, Wang L, Wang LF, Cui J, Chen H, Zhu Q (2016) Co-circulation of H5N6, H3N2, H3N8, and emergence of novel reassortant H3N6 in a local community in Hunan Province in China. Sci Rep 6:25549
Brooks-Moizer F, Roberton SI, Edmunds K, Bell D (2009) Avian influenza H5N1 and the wild bird trade in Hanoi Vietnam. Ecol Soc. https://doi.org/10.5751/ES-02760-140128
Nguyen LT, Firestone SM, Stevenson MA, Young ND, Sims LD, Chu DH, Nguyen TN, Van Nguyen L, Le Thanh T, Van Nguyen H, Nguyen HN, Tien TN, Nguyen TD, Tran BN, Matsuno K, Okamatsu M, Kida H, Sakoda Y (2019) A systematic study towards evolutionary and epidemiological dynamics of currently predominant H5 highly pathogenic avian influenza viruses in Vietnam. Sci Rep 9:7723
Chu DH, Okamatsu M, Matsuno K, Hiono T, Ogasawara K, Nguyen LT, Van Nguyen L, Nguyen TN, Nguyen TT, Van Pham D, Nguyen DH, Nguyen TD, To TL, Van Nguyen H, Kida H, Sakoda Y (2016) Genetic and antigenic characterization of H5, H6 and H9 avian influenza viruses circulating in live bird markets with intervention in the center part of Vietnam. Vet Microbiol 192:194–203
Kim KI, Choi JG, Kang HM, To TL, Nguyen TD, Song BM, Hong MS, Choi KS, Kye SJ, Kim JY, Lee HS, Lee YJ (2013) Geographical distribution of low pathogenic avian influenza viruses of domestic poultry in Vietnam and their genetic relevance with Asian isolates. Poult Sci 92:2012–2023
Okamatsu M, Nishi T, Nomura N, Yamamoto N, Sakoda Y, Sakurai K, Chu HD, Thanh LP, Van Nguyen L, Van Hoang N, Tien TN, Yoshida R, Takada A, Kida H (2013) The genetic and antigenic diversity of avian influenza viruses isolated from domestic ducks, muscovy ducks, and chickens in northern and southern Vietnam, 2010–2012. Virus Genes 47:317–329
Takakuwa H, Yamashiro T, Le MQ, Phuong LS, Ozaki H, Tsunekuni R, Usui T, Ito H, Morimatsu M, Tomioka Y, Yamaguchi T, Ito T, Murase T, Ono E, Otsuki K (2012) Molecular epidemiology of avian influenza viruses circulating among healthy poultry flocks in farms in northern Vietnam. Prev Vet Med 103:192–200
Kida H, Yangawa R (1979) Isolation and characterization of influenza a viruses from wild free-flying ducks in Hokkaido, Japan. Zentralbl Bakteriol [Orig A] 244:135–143
Lee MS, Chang PC, Shien JH, Cheng MC, Shieh HK (2001) Identification and subtyping of avian influenza viruses by reverse transcription-PCR. J Virol Methods 97:13–22
Tsukamoto K, Ashizawa H, Nakanishi K, Kaji N, Suzuki K, Okamatsu M, Yamaguchi S, Mase M (2008) Subtyping of avian influenza viruses H1 to H15 on the basis of hemagglutinin genes by PCR assay and molecular determination of pathogenic potential. J Clin Microbiol 46:3048–3055
Tsukamoto K, Ashizawa T, Nakanishi K, Kaji N, Suzuki K, Shishido M, Okamatsu M, Mase M (2009) Use of reverse transcriptase PCR to subtype N1 to N9 neuraminidase genes of avian influenza viruses. J Clin Microbiol 47:2301–2303
Hoffmann E, Stech J, Guan Y, Webster RG, Perez DR (2001) Universal primer set for the full-length amplification of all influenza A viruses. Arch Virol 146:2275–2289
Edgar RC (2004) MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res 32:1792–1797
Kumar S, Stecher G, Li M, Knyaz C, Tamura K (2018) MEGA X: molecular evolutionary genetics analysis across computing platforms. Mol Biol Evol 35:1547–1549
Underwood PA (1982) Mapping of antigenic changes in the haemagglutinin of Hong Kong influenza (H3N2) strains using a large panel of monoclonal antibodies. J Gen Virol 62(Pt 1):153–169
Tian H, Zhou S, Dong L, Van Boeckel TP, Cui Y, Newman SH, Takekawa JY, Prosser DJ, Xiao X, Wu Y, Cazelles B, Huang S, Yang R, Grenfell BT, Xu B (2015) Avian influenza H5N1 viral and bird migration networks in Asia. Proc Natl Acad Sci USA 112:172–177
Koehler AV, Pearce JM, Flint PL, Franson JC, Ip HS (2008) Genetic evidence of intercontinental movement of avian influenza in a migratory bird: the northern pintail (Anas acuta). Mol Ecol 17:4754–4762
Si Y, Skidmore AK, Wang T, de Boer WF, Debba P, Toxopeus AG, Li L, Prins HH (2009) Spatio-temporal dynamics of global H5N1 outbreaks match bird migration patterns. Geospat Health 4:65–78
Zost SJ, Parkhouse K, Gumina ME, Kim K, Diaz Perez S, Wilson PC, Treanor JJ, Sant AJ, Cobey S, Hensley SE (2017) Contemporary H3N2 influenza viruses have a glycosylation site that alters binding of antibodies elicited by egg-adapted vaccine strains. Proc Natl Acad Sci USA 114:12578–12583
Acknowledgements
This was a collaborative research effort between the National Institute of Hygiene and Epidemiology (Vietnam), Nagasaki University (Japan), and Tottori University (Japan). This research was supported by AMED under Grant Number JP19fm0108001 (Japan Initiative for Global Research Network on Infectious Diseases (J-GRID)), and in part by a grant from MEXT for the Joint Research Program of the Research Center for Zoonosis Control, Hokkaido University, Japan.
Author information
Authors and Affiliations
Contributions
KS (first author) was responsible for the conception and design of the study, the acquisition, collation, and interpretation of the data, and the writing of the submitted article. MK was involved in the phylogenic/antigenic analyses and the animal infection experiment. KH, TTHU., and HLKN performed the virus surveillance and isolation study. HI, MQL, and TI (corresponding author) were involved in the coordination of the study, the import formality (Vietnam to Japan), and the submission process.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Edited by Simon D. Scott.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Supplementary file2 (PDF 110 kb)
Fig. S1 Phylogenetic trees of the N2 NA (A), N3 NA (B), N8 NA (C), PB2 (D), PB1 (E), PA (F), NP (G), and NS (H) gene segments of H3 AIV. The H3 virus isolates in our surveillance in Vietnam are shown in red. Wildfowl-originating viruses and H5 HPAIVs are shown in green and blue, respectively. The compressed taxa on the tree are shown by the italicized common denominators. The reference strains used for antigenic analysis in the present study are marked with filled circles. The partial sequences of each segment indicated in Table S1 were used for the analysis. Horizontal distances are proportional to the minimum number of nucleotide differences required to join the nodes and sequences. Numbers at the nodes indicate confidence levels in a bootstrap analysis with 1,000 replications. Bootstrap values of more than 60% are shown at each branch. The strains together with the virus isolates sharing a distinct common node [shown via asterisks (*)] were assumed to be within the same group.
Fig. S2 Alignment of the deduced H3 HA amino acid sequences of the strains used for antigenic analysis (H3 HA numbering). Group numbers are based on the H3 HA gene phylogeny (Fig. 1A). Amino acids constituting five antigenic sites (A—E) are boxed. Amino acid residues different from those in the group I LBM48 strain are shown in red.
Rights and permissions
About this article
Cite this article
Soda, K., Kashiwabara, M., Miura, K. et al. Characterization of H3 subtype avian influenza viruses isolated from poultry in Vietnam. Virus Genes 56, 712–723 (2020). https://doi.org/10.1007/s11262-020-01797-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11262-020-01797-7