Abstract
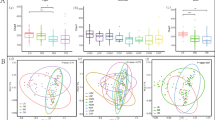
Until now, there has been little research on the intestinal microbial community of horseshoe crabs. To fill this gap, we investigated the microbiome composition of the Chinese horseshoe crab, Tachypleus tridentatus, and the mangrove horseshoe crab, Carcinoscorpius rotundicauda. We sequenced the 16S rRNA gene of intestinal bacterial species and compared the microbial community structure and diversity. Next, we show that the total effective bacterial sequence was 36,865 reads, and the average annotated operational taxonomic unit (OTU) number was 240. Through hierarchical clustering analysis and principal coordinate analysis samples from two horseshoe crab species, we found that the intestinal flora of the same horseshoe crab species was relatively concentrated, while the microbiome of a different horseshoe crab species were significantly separated. Cluster analysis showed that two samples, one from Chinese horseshoe crabs and one from mangrove horseshoe crabs, had similar microbial community structure, while other samples were relatively discrete. The gut microbiota of the mangrove horseshoe crab were dominated by the phyla Tenericutes (42.71%), Firmicutes (24.27%), and Proteobacteria (20.39%), while the top three phyla in the Chinese horseshoe crab intestinal tract were Tenericutes (57.19%), Proteobacteria (22.14%), and Bacteroidetes (7.38%). To intuitively understand the similarity and overlap of the OTU composition of each group, we performed Venn diagram analysis. The two species shared 284 OTUs, accounting for 81.8% of the total. This indicates that although there is high similarity between mangrove and Chinese horseshoe crab in gastrointestinal microbial community structure, there are also some differences, which deserve further discussion.




Similar content being viewed by others
Availability of Data and Materials
The sequence data supporting the result of this article in the NCBI (National Center for Biotechnology Information) under accession number PRJNA642919.
References
Walls EA, Berkson J, Smith SA (2002) The horseshoe crab, Limulus polyphemus: 200 million years of existence, 100 years of study. Rev Fish Sci 10:39–73
Rudkin DM, YoungGA NGS (2008) The oldest horseshoe crab: a new xiphosurid from Late Ordovician Konservat-Lagerstätten deposits, Manitoba, Canada. Palaeontology 51:1–9
Sekiguchi K (1988) Biology of horseshoe crabs. Science House, Tokyo, pp 10–21
Liao YY, Liu JX (2006) Species and distribution of horseshoe crab in Asia sea area. J Trop Oceanogr 25:85–90 (In Chinese, with English abstract)
Liao YY, Hsieh HL, Xu SQ, Zhong QP, Lei J, Liang MZ, Fang HY, Xu LL, Lin WY, Xiao XB, Chen CP, Cheung SG, Kwan BK (2019) Wisdom of Crowds reveals decline of Asian horseshoe crabs in Beibu Gulf, China. Oryx 53:222–229
Gauvry G (2015) Current horseshoe crab harvesting practices cannot support global demand for TAL/LAL: the pharmaceutical and medical device industries’ role in the sustainability of horseshoe crabs. Changing global perspectives on horseshoe crab biology, conservation and management. Springer, Cham, pp 475–482
Coyte KZ, Schluter J, Foster KR (2015) The ecology of the microbiome: networks, competition, and stability. Science 350:663–666
Endesfelder D, Zu Castell W, Ardissone A, Davis-Richardson AG, Achenbach P, Hagen M, Pflueger M, Gano KA, Fagen JR, Drew JC, Brown CT, Kolaczkowski B, Atkinson M, Schatz D, Bonifacio E, Triplett EW, Ziegler AG (2014) Compromised gut microbiota networks in children with anti-islet cell autoimmunity. Diabetes 63:2006–2014
Ratana-Arporn P, Jommark N (2014) Efficacy of neutral electrolyzed water for reducing pathogenic bacteria contaminating shrimp. J Food Prot 77:2176–2180
O'Hara AM, Shanahan F (2006) The gut flora as a forgotten organ. EMBO Rep 7:688–693
Fava F, Rizzetto L, Tuohy KM (2019) Gut microbiota and health: connecting actors across the metabolic system. Proc Nutr Soc 78:177–188
Chi C, Liu JY, Fei SZ, Zhang C, Chang YQ, Liu XL, Wang GX (2014) Effect of intestinal autochthonous probiotics isolated from the gut of sea cucumber (Apostichopus japonicus) on immune response and growth of A. japonicus. Fish Shellfish Immunol 38:367–373
Xiong JB, Wang K, Wu JF, Qiuqian LL, Yang KJ, Qian YX, Zhang DM (2015) Changes in intestinal bacterial communities are closely associated with shrimp disease severity. Appl Microbiol Biotechnol 99:6911–6919
Zhu JY, Dai WF, Qiu QF, Dong CM, Zhang JJ, Xiong JB (2016) Contrasting ecological processes and functional compositions between intestinal bacterial community in healthy and diseased shrimp. Microb Ecol 72:975–985
Magoč T, Salzberg SL (2011) FLASH: fast length adjustment of short reads to improve genome assemblies. Bioinformatics 27:2957–2963
Bokulich NA, Subramanian S, Faith JJ, Gevers D, Gordon JI, Knight R, Mills DA, Caporaso JG (2013) Quality-filtering vastly improves diversity estimates from Illuminaamplicon sequencing. Nat Methods 10:57–59
Caporaso JG, Kuczynski J, Stombaugh J, Bittinger K, Bushman FD, Costello EK, Fierer N, Peña AG, Goodrich JK, Gordon JI, Huttley GA, Kelley S, Knights D, Koenig J, Ley RE, Lozupone CA, McDonald DM, Muegge BD, Pirrung M, Reeder J, Sevinsky JR, Turnbaugh PJ, Walters WA, Widmann J, Yatsunenko T, Zaneveld J, Knight R (2010) QIIME allows analysis of high-throughput community sequencing data. Nat Methods 7:335–336
Edgar RC, Haas BJ, Clemente JC, Quince C, Knight R (2011) UCHIME improves sensitivity and speed of chimera detection. Bioinformatics 27(16):2194–2200
Haas BJ, Gevers D, Earl AM, Feldgarden M, Ward DV, Giannoukos G, Ciulla D, Tabbaa D, Highlander SK, Sodergren E, Methé B, DeSantis TZ, Human Microbiome Consortium, Petrosino JF, Knight R, Birren BW (2011) Chimeric 16S rRNA sequence formation and detection in Sanger and 454-pyrosequenced PCR amplicons. Genome Res 21(3):494–504
Edgar RC (2013) UPARSE: highly accurate OTU sequences from microbial amplicon reads. Nat Methods 10:996–998
Wang Q, Garrity GM, Tiedje JM, Cole JR (2007) Naive Bayesian classifier for rapid assignment of rRNA sequences into the new bacterial taxonomy. Appl Environ Microbiol 73:5261–5267
Quast C, Pruesse E, Yilmaz P, Gerken J, Schweer T, Yarza P, Peplies J, Glöckner FO (2013) The SILVA ribosomal RNA gene database project: improved data processing and web-based tools. Nucl Acids Res 41:D590–D596
Edgar RC (2004) MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res 32:1792–1797
Segata N, Izard J, Waldron L, Gevers D, Miropolsky L, Garrett WS, Huttenhower C (2011) Metagenomic biomarker discovery and explanation. Genome Biol 12(6):R60
Zoqratt MZHM, Eng WWH, Thai BT, Austin CM, Gan HM (2018) Microbiome analysis of Pacific white shrimp gut and rearing water from Malaysia and Vietnam: implications for aquaculture research and management. PeerJ 6:e5826
Saw JHW, Yuryev A, Kanbe M, Hou SB, Young AG, Aizawa S, Alam M (2012) Complete genome sequencing and analysis of Saprospira grandis str. Lewin, a predatory marine bacterium. Stand Genomic Sci 6:84
Xue SX, Xu W, Wei JL, Sun JS (2017) Impact of environmental bacterial communities on fish health in marine recirculating aquaculture systems. Vet Microbiol 203:34–39
Torres M, Reina JC, Fuentes-Monteverde JC, Fernández G, Rodríguez J, Jiménez C, Llamas I (2018) AHLlactonase expression in three marine emerging pathogenic Vibrio sp. reduces virulence and mortality in brine shrimp (Artemia salina) and Manila clam (Venerupis philippinarum). PLoS ONE 13:e0195176
Thompson FL, Iida T, Swings J (2004) Biodiversity of vibrios. Microbiol Mol Biol Rev 68:403–431
Ray AK, Ghosh K, Ringø E (2012) Enzyme-producing bacteria isolated from fish gut: a review. Aquac Nut 18:465–492
Li L, Kato C, Horikoshi K (1999) Microbial diversity in sediments collected from the deepest cold-seep area, the Japan Trench. Mar Biotechnol 1:391–400
Stevens H, Matthias S, Simon M, Brinkhoff T (2010) Phylogeny of proteobacteria and bacteroidetes from oxichabitats of a tidal flat ecosystem. FEMS Microbiol Ecol 54:351–365
Thomas F, Hehemann JH, Rebuffet E, Czjzek M, Michel G (2011) Environmental and gut bacteroidetes: the food connection. Front Microbiol 2:93
Kim YS, Milner JA (2007) Dietary modulation of colon cancer risk. J Nutr 137:2576–2579
Kim TY, Lee JJ, Kim BS, Choi SH (2017) Whole-body microbiota of sea cucumber (Apostichopus japonicus) from South Korea for improved seafood management. J Microbiol Biotechnol 27:1753–1762
Chai YH, Gao F, Wang JF, Zhou WL (2019) Intestinal microbiata in japonicus: regional difference and common feature. Oceanol Limnol Sin 50:1127–1137 (In Chinese, with English abstract)
Kwan BKY, Hsieh HL, Cheung SG, Shin PKS (2016) Present population and habitat status of potentially threatened Asian horseshoe crabs Tachypleus tridentatus and Carcinoscorpius rotundicauda in Hong Kong: a proposal for marine protected areas. Biodivers Conserv 25:673–692
Li HY (2008) The conservation of horseshoe crabs in Hong Kong. Doctoral dissertation, City University of Hong Kong, Hong Kong
Chen CP, Yang MC, Fan LF, Qiu GL, Liao YY, Hsieh HL (2015) Co-occurrence of juvenile horseshoe crabs Tachypleus tridentatus and Carcinoscorpius rotundicauda in an estuarine bay, southwestern China. Aquat Biol 24:117–126
Jawahir ARN, Samsur M, Shabdin ML, Adha ARK (2017) Distribution of two species of Asian horseshoe crabs at west coast of Sarawak's Waters, East Malaysia. Egypt J Aquat Res 43:135–140
Ringø E, Hoseinifar SH, Ghosh K, Doan HV, Beck BR, Song SK (2018) Lactic acid bacteria in finfish—an update. Front Microbiol 9:1818
Li T, He L, Liang QF (2013) Progress in the study of Rhizobium associated with non-leguminous plants. J Agric Sci Technol 15:97–102 (In Chinese, with English abstract)
Acknowledgements
This project was funded by the Guangxi Young and Middle-aged Teachers Basic Ability Improvement Project (KY2016YB488), Qinzhou Science and Technology Research and Development Plan (20198510, 20198520) and Guangxi Key Laboratory of Beibu Gulf Marine Biodiversity Conservation, Beibu Gulf University (2015ZC14). We are grateful to teachers from the Ocean College and Food Engineering College of Beibu Gulf University for their assistance in sample collection and data processing.
Author information
Authors and Affiliations
Contributions
PW and HW designed the research; PW, YN, and JH analyzed the data and wrote the manuscript; YN, JH, and ZD performed sample preparation; YW and YL revised the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no conflict of interests.
Ethical Approval
This study conformed to the guidelines for the care and use of experimental animals established by the Ministry of Science and Technology of the People’s Republic of China (Approval number: 2006-398). All of the procedures for experimental animal manipulation were approved by the Animal Research and Ethics Committee of the Beibu Gulf University.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Wang, P., Ning, Y., Huang, J. et al. Intestinal Tract Microbe Communities Associated with Horseshoe Crabs from Beibu Gulf, China. Curr Microbiol 77, 3330–3338 (2020). https://doi.org/10.1007/s00284-020-02140-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00284-020-02140-x