Abstract
Introduction
Since Populus has veritable value as timber, plywood, pulp, and paper, genomic research should create the sound basis for further breeding toward desirable wood quality attributes.
Materials and methods
In this study, we addressed the need for a research methodology that initially identifies and then characterize candidate genes encoding enzymes with wood property phenotypic traits, toward the aim of developing a genomics-based breeding technology.
Results
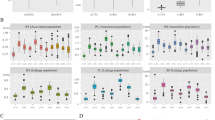
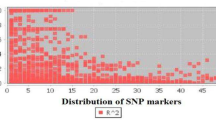
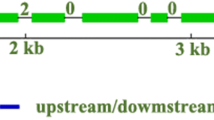
On 23 different poplar species/hybrid samples, we successfully amplified 55 primers designed on Populus trichocarpa L. Considering the number of polymorphic sites, out of 73,206 bp, 51 SNPs and 31 indel events were found. Non-synonymous single base mutations could be detected in number of 30, 21 out of 164 sequences were the number of minimum recombination events and 41 significant pairwise comparisons between loci could be detected.
Discussion and conclusion
Our results provide a roadmap for a future association genetic study between nucleotide diversity and precise evaluation of phenotype.
Similar content being viewed by others
References
Ache, P., Fromm, J., Hedrich, R. (2010) Potassium-dependent wood formation in poplar: seasonal aspects and environmental limitations. Plant Biol. 12, 259–267.
Arend, M., Stinzing, A., Wind, C., Langer, K., Latz, A., Ache, P., Hedrich, R. (2005) Polar-localised poplar K+ channel capable of controlling electrical properties of wood-forming cells. Planta 223, 140–148.
Bedon, F., Legay, S. (2011) Lignin synthesis, transcriptional regulation and potential for useful modification in plants. Plant. Sci. Rev. 6, 89.
Bradshaw, H. D., Stettler, R. E. (1993) Molecular genetics of growth and development in Populus. I. Triploidy in hybrid poplars. Theor. Appl. Gen. 86, 301–302.
Brunner, A. M., Busov, V. B., Strauss, S. H. (2004) Poplar genome sequence: functional genomics in an ecologically dominant plant species. Trends Plant Sci. 9, 49–56.
Carletti, G., Carra, A., Allegro, G., Vietto, L., Desiderio, F., Bagnaresi, P., Nervo, G. (2016) QTLs for woolly poplar aphid (Phloeomyzus passerinii L.) resistance detected in an interspecific Populus deltoides × P. nigra mapping population. PLoS ONE 11, 1–18.
Charlesworth, B. (1990) Mutation-selection balance and the evolutionary advantage of sex and recombination. Genet. Res. 55, 199–221.
Decroocq, V., Fave, M. G., Hagen, L., Bordenave, L., Decroocq, S. (2003) Development and transferability of apricot and grape EST microsatellite markers across taxa. Theor. Appl. Genet. 106, 912–922.
Denison, F. C., Paul, A. L., Zupanska, A. K., Ferl, R. J. (2011) 14-3-3 Proteins in plant physiology. Semin. Cell. Dev. Biol. 22, 720–727.
Dharmawardhana, P., Brunner, A. M., Strauss, S. H. (2010) Genome-wide transcriptome analysis of the transition from primary to secondary stem development in Populus trichocarpa. BMC Genomics 11, 150.
Drost, D. R., Puranik, S., Novaes, E., Novaes, C. R., Dervinis, C., Gailing, O., Kirst, M. (2015) Genetical genomics of Populus leaf shape variation. BMC Plant Biol. 15, 1–10.
Du, F. K., Xu, F., Qu, H., Feng, S., Tang, J., Wu, R. (2013) Exploiting the transcriptome of euphrates poplar, Populus euphratica (Salicaceae) to develop and characterize new EST-SSR markers and construct an EST-SSR database. PloS one 8, 1–11.
Dumolin, S., Demesure, B., Petit, R. J. (1995) Inheritance of chloroplast and mitochondrial genomes in pedunculate oak investigated with an efficient PCR method. Theor. Appl. Genet. 91, 1253–1256.
Feng, S. P., Li, W. G., Huang, H. S., Wang, J. Y., Wu, Y. T. (2009) Development, characterization and cross-species/genera transferability of EST-SSR markers for rubber tree (Hevea brasiliensis). Mol. Breed. 23, 85–97.
Fromm, J. (2010) Wood formation of trees in relation to potassium and calcium nutrition. Tree Physiol. 30, 1140–1147.
Fromm, J., Lautner, S. (2016) Abiotic stresses on secondary xylem formation. In: Kim, Y. S., Funada, R., Singh, A. P. (eds) Secondary Xylem Biology, Elsevier Academic Press, Amsterdam, Netherlands, pp. 59–71.
Garcia, K., Zimmermann, S. D. (2014) The role of mycorrhizal associations in plant potassium nutrition. Front. Plant Sci. 5, 1–9.
Geisler-Lee, J., Geisler, M., Coutinho, P. M., Segerman, B., Nishikubo, N, Takahashi, J., Sundberg, B. (2006) Poplar carbohydrate-active enzymes. Gene identification and expression analyses. Plant Physiol. 140, 946–962.
Hall, T. A. (1999) BioEdit: a user-friendly biological sequence alignment editor and analysis program for Windows 95/98/NT. Nucl. Acid Sci. 41, 95–98.
Halpin, C. (2004) Re-designing lignin for industry and agriculture. Biotechnol. Genet. Eng. 21, 229–248.
Henry, I. M., Zinkgraf, M. S., Groover, A. T., Comai, L. (2015) A system for dosage-based functional genomics in poplar. Plant Cell 27, 2370–2383.
Isabel, N., Lamothe, M., Thompson, S. L. (2013) A second-generation diagnostic single nucleotide polymorphism (SNP)-based assay, optimized to distinguish among eight poplar (Populus L.) species and their early hybrids. Tree Genet. Genomes 9, 621–626.
Joshi, C. P., DiFazio, S. P., Kole, C. (eds) (2011) Genetics, Genomics and Breeding of Poplar. Science Publisher, Clemson.
Langer, K., Levchenko, V., Fromm, J., Geiger, D., Steinmeyer, R., Lautner, S., Hedrich, R. (2004) The poplar K+ channel KPT1 is associated with K+ uptake during stomatal opening and bud development. Plant J. 37, 828–838. doi: 10.1111/j.0960-7412.2003.02008.x.
Li, L., Lu, S., Chiang, V. (2006) A genomic and molecular view of wood formation. Crit. Rev. Plant Sci. 25, 215–233.
Loziuk, P. L., Hecht, E. S., Muddiman, D. C. (2017) N-linked glycosite profiling and use of Skyline as a platform for characterization and relative quantification of glycans in differentiating xylem of Populus trichocarpa. Anal. Bioanal. Chem. 409, 487–497.
Lu, S., Li, Q., Wei, H., Chang, M. J., Tunlaya-Anukit, S., Kim, H., Yeh, T. F. (2013) Ptr-miR397a is a negative regulator of laccase genes affecting lignin content in Populus trichocarpa. Proc. Natl. Acad. Sci. U. S. A. 110, 10848–10853.
Luo, R., Chen, P. W., Wagenbach, M., Jian, X., Jenkins, L., Wordeman, L., Randazzo, P. A. (2016) Direct functional interaction of the kinesin-13 family member kinesin-like protein 2A (Kif2A) and Arf GAP with GTP-binding protein-like, ankyrin repeats and PH domains 1 (AGAP1). J. Biol. Chem. 291, 25761.
Mellerowicz, E. J., Sundberg, B. (2008) Wood cell walls: biosynthesis, developmental dynamics and their implications for wood properties. Curr. Opin. Plant Biol. 11, 293–300.
Mutwil, M., Debolt, S., Persson, S. (2008) Cellulose synthesis: a complex complex. Curr. Opin. Plant Biol. 11, 252–257.
Nersissian, A. M., Valentine, J. S., Immoos, C., Hill, M. G., Hart, P. J., Williams, G., Herrmann, R. G. (1998) Uclacyanins, stellacyanins, and plantacyanins are distinct subfamilies of phytocyanins: plant-specific mononuclear blue copper proteins. Protein Sci. 7, 1915–1929.
Oakley, R. V., Wang, Y. S., Ramakrishna, W., Harding, S. A., Tsai, C. J. (2007) Differential expansion and expression of alfa-and beta-tubulin gene families in Populus. Plant Physiol. 145, 961–973.
Olson, M. S., Robertson, A. L., Takebayashi, N., Silim, S., Schroeder, W. R., Tiffin, P. (2010) Nucleotide diversity and linkage disequilibrium in balsam poplar (Populus balsamifera). New Phytol. 186, 526–536.
Rademacher, P., Báder, M., Németh, R., Rousek, R., Paril, P., Baar, J., Hornicek, S., Dejmal, A., Domeny, J., Kudela, J., Kutnar, A., Neyses, B., Sandberg, D. (2017) European co-operation in wood research– from native wood to engineered materials. Part 2: densification modification in product development. In: Gurau, L., Campean, M., Ispas, M. (eds) Proceedings of the International Conference Wood Science and Engineering in the Third Millenium (ICWSE 2017).Transilvania University, Brasov, Romania, pp. 469–478.
Peakall, R., Gilmore, S., Keys, W., Morgante, M., Rafalski, A. (1996) Cross-species amplification of soybean (Glycine max) simple sequence repeats (SSRs) within the genus and other legume genera: implications for the transferability of SSRs in plants. Mol. Biol. Evol. 15, 1275–1287.
Rajinikanth, M., Harding, S. A., Tsai, C. J. (2007) The glycine decarboxylase complex multienzyme family in Populus. J. Exp. Bot. 58, 1761–1770.
Ralph, J., Schatz, P. F., Lu, F., Kim, H., Akiyama, T., Nelsen, S. F. (2009) Quinone Methides in Lignification. Wiley-Blackwell, Hoboken, NJ.
Ranjan, P., Yin, T., Zhang, X., Kalluri, U. C., Yang, X., Jawdy, S., Tuskan, G. A. (2010) Bioinformatics-based identification of candidate genes from QTLs associated with cell wall traits in Populus. Bioenergy Res. 3, 172–182.
Ratke, C., Pawar, P. M. A., Balasubramanian, V. K., Naumann, M., Duncranz, M. L., Derba-Maceluch, M., Mellerowicz, E. J. (2015) Populus GT43 family members group into distinct sets required for primary and secondary wall xylan biosynthesis and include useful promoters for wood modification. Plant Biotechnol. J. 13, 26–37.
Rozas, J., Rozas, R. (1995) DnaSP, DNA sequence polymorphism: an interactive program for estimating population genetics parameters from DNA sequence data. CABIOS 11, 621–625.
Sasaki, T., Fukuda, H., Oda, Y. (2017) Cortical microtubule disorderingl is required for secondary cell wall patterning in xylem vessels. Plant Cell 29, 3123–3139.
Savolainen, O., Pyhajarvi, T. (2007) Genomic diversity in forest trees. Curr. Opin. Plant Biol. 10, 162–167.
Sawada, D., Kalluri, U. C., O’Neill, H., Urban, V., Langan, P., Davison, B., Pingali, S. V. (2018) Tension wood structure and morphology conducive for better enzymatic digestion. Biotechnol. Biofuels 11, 1–9.
Sterky, F., Regan, S., Karlsson, J., Hertzberg, M., Rohde, A., Holmberg, A., Van Montagu, M. (1998) Gene discovery in the wood-forming tissues of poplar: analysis of 5, 692 expressed sequence tags. Proc. Natl. Acad. Sci. U. S. A. 95, 13330–13335.
Takabe, K., Takeuchi, M., Sato, T., Ito, M., Fujita, M. (2001) lmmunocytochemical localization of enzymes involved in lignification of the cell wall. J. Plant Res. 114, 509–515.
Tenney, A. E., Wu, J. Q., Langten, L., Klueh, P., Quatrano, R., Brent, M. R. (2007) A tale of two templates: automatically resolving double traces has many applications, including efficient PCR-based elucidation of alternative splices. Genome Res. 17, 212–218.
Vander Mijnsbrugge, K., Meyermans, H., Van Montagu, M., Bauw, G., Boerjan, W. (2000) Wood formation in poplar: identification, characterization, and seasonal variation of xylem proteins. Planta 210, 589–598.
Villalobos, D. P., Diaz-Moreno, S. M., Said, E. S. S., Cañas, R. A., Osuna, D., Van Kerckhoven, S. H., Cantón, F. R. (2012) Reprogramming of gene expression during compression wood formation in pine: coordinated modulation of s-adenosylmethionine, lignin and lignan related genes. BMC Plant Biol. 12, 100.
Wright, S. I., Andolfatto, P. (2008) The impact of natural selection on the genome: emerging patterns in Drosophila and Arabidopsis. Annu. Rev. Ecol. Evol. Syst. 39, 193–213.
Zhong, R., Ye, Z. H. (2007) Regulation of cell wall biosynthesis. Curr. Opin. Plant Biol. 10, 564–572.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Köbölkuti, Z.A., Cseke, K., Benke, A. et al. Allelic variation in candidate genes associated with wood properties of cultivated poplars (Populus). BIOLOGIA FUTURA 70, 286–294 (2019). https://doi.org/10.1556/019.70.2019.32
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1556/019.70.2019.32