Abstract
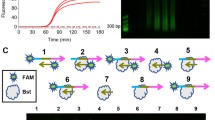
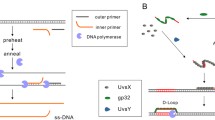
In recent years, methods of isothermal amplification of nucleic acids using polymerases with a strand-displacement activity have become widespread for identification of specific nucleotide sequences. Bst exo– polymerase is the most popular of these polymerases, although it is inclined to nonspecific amplification (the so-called multimerization), which leads to the accumulation of by-products constructed of tandem nucleotide repeats. In this study, we evaluated the efficiency of multimerization depending on the reaction conditions and proposed some methods for its elimination. The highest efficiency of multimerization was found in the case of Bst 2.0 polymerase in Isothermal buffer, whereas the Bst-like Gss polymerase provided the formation of multimerization products only in Isothermal buffer and at the latest stages of the reaction. The optimal method for elimination of multimerization was the use of Gss polymerase and Thermopol buffer, or Bst LF polymerase and Isothermal II buffer, or Bst 3.0 polymerase and Thermopol buffer, or Bst 3.0 polymerase in Isothermal buffer and Mn2+ ions as a cofactor. In these cases specific isothermal amplification of the target DNA may take place and provide accurate and reliable results.



Similar content being viewed by others
REFERENCES
Fire, A. and Xu, S.Q., Proc. Natl. Acad. Sci. U. S. A., 1995, vol. 92, pp. 4641–4645.
Ali, M.M., Li, F., Zhang, Zh., Zhang, K., Kang, D.K., Ankrum, J.A., Le, X.C., and Zhao, W., Chem. Soc. Rev., 2014, vol. 43, pp. 3324–3341.
Mohsen, M.G. and Kool, E.T., Acc. Chem. Res., 2016, vol. 49, pp. 2540–2550.
Notomi, T., Okayama, H., Masubuchi, H., Yonekawa, T., Watanabe, K., Amino, N., and Hase, T., Nucleic Acids Res., 2000, vol. 28. E63.
Wong, Y.P., Othman, S., Lau, Y.L., Radu, S., and Chee, H.Y., J. Appl. Microbiol., 2018, vol. 124, pp. 626–643.
Compton, J., Nature, 1991, vol. 350, pp. 91–92.
Walter, N.G. and Strunk, G., Proc. Natl. Acad. Sci. U. S. A., 1994, vol. 91, pp. 7937–7941.
Kim, J. and Easley, C.J., Bioanalysis, 2011, vol. 3, pp. 227–239.
Zhao, Y., Chen, F., Li, Q., Wang, L., and Fan, C., Chem. Rev., 2015, vol. 115, pp. 12491–12545.
Fakruddin, M., Mannan, K.S., Chowdhury, A., Mazumdar, R.M., Hossain, M.N., Islam, S., and Chowdhury, M.A., J. Pharm. Bioallied Sci., 2013, vol. 5, pp. 245–252.
Deng, H. and Gao, Z., Anal. Chim. Acta, 2015, vol. 853, pp. 30–45.
Mayboroda, O., Katakis, I., and O’Sullivan, C.K., Anal. Biochem., 2018, vol. 545, pp. 20–30.
Giuffrida, M.C. and Spoto, G., Biosens. Bioelectron., 2017, vol. 90, pp. 174–186.
Qi, H., Yue, S., Bi, S., Ding, C., and Song, W., Biosens. Bioelectron., 2018, vol. 110, pp. 207–217.
Cao, H., Zhou, X., and Zeng, Y., Sens. Actuators, B, 2019, vol. 279, pp. 447–457.
Treerattrakoon, K., Jiemsakul, T., Tansarawiput, C., Pinpradup, P., Iempridee, T., Luksirikul, P., Khoothiam, K., Dharakul, T., and Japrung, D., Anal. Biochem., 2019, vol. 577, pp. 89–97.
Ma, Y., Zhang, B., Wang, M., Ou, Y., Wang, J., and Li, S., Biomed. Res. Int., 2016, p. 2906484.
Rastgoo, N., Sadeghizadeh, M., Bambaei, B., and Hosseinkhani, S., J. Biotechnol., 2009, vol. 144, pp. 245–252.
Çağlayan, M. and Bilgin, N., Appl. Biochem. Biotechnol., 2011, vol. 165, pp. 1188–200.
Oscorbin, I.P., Belousova, E.A., Boyarskikh, U.A., Zakabunin, A.I., Khrapov, E.A., and Filipenko, M.L., Nucleic Acids Res., 2017, vol. 45, pp. 9595–9610.
Oscorbin, I.P., Boyarskikh, U.A., and Filipenko, M.L., Mol. Biotechnol., 2015, vol. 57, pp. 947–959.
Zyrina, N.V., Antipova, V.N., and Zheleznaya, L.A., FEMS Microbiol. Lett., 2014, vol. 351, pp. 1–6.
Hafner, G.J., Yang, I.C., Wolter, L.C., Stafford, M.R., and Giffard, P.M., BioTechniques, 2001, vol. 30, pp. 852–867.
Gu, L., Yan, W., Liu, L., Wang, S., Zhang, X., and Lyu, M., Pharmaceuticals, 2018, vol. 11, p. 35.
Wang, G., Ding, X., Hu, J., Wu, W., Sun, J., and Mu, Y., Sci. Rep., 2017, vol. 7. E13928.
Oscorbin, I.P., Belousova, E.A., Zakabunin, A.I., Boyarskikh, U.A., and Filipenko, M.L., BioTechniques, 2016, vol. 61, pp. 20–25.
Gil’vanov, A.R., Sakhabutdinova, A.R., Chemeris, A.V., and Garafutdinov, R.R., Biomika, 2018, vol. 11, pp. 268–273.
Güixens-Gallardo, P., Hocek, M., and Perlíková, P., Bioorg. Med. Chem. Lett., 2016, vol. 26, pp. 288–291.
Hays, H. and Berdis, A.J., Biochemistry, 2002, vol. 41, pp. 4771–4778.
Frank, E.G. and Woodgate, R.J., Biol. Chem., 2007, vol. 282, pp. 24689–24696.
Xu, W., Zhao, W., Morehouse, N., Tree, M.O., and Zhao, L., J. Mol. Biol., 2019, vol. 431, pp. 673–686.
Ralec, C., Henry, E., Lemor, M., Killelea, T., and Henneke, G., Nucleic Acids Res., 2017, vol. 45, pp. 12425–12440.
Vashishtha, A.K. and Konigsberg, W.H., Biochemistry, 2018, vol. 55, pp. 2661–2670.
ACKNOWLEDGMENTS
The studies were conducted using the equipment of the Center of Collective Use Biomika and a unique scientific system KODINK.
Funding
The work was supported by Russian State Federal budget (project no. АААА-А16-116020350032-1) and partially supported by Russian State-funded Budget project no. АААА-А17-117020210023-1 to ICBFM SB RAS.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
COMPLIANCE WITH ETHICAL STANDARDS
In this work, humans or animals were not involved as subjects of studies.
Conflict of Interests
The authors notified about the absence of conflict of interest.
Additional information
Translated by E. Shirokova
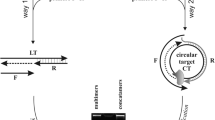
Abbreviations: CT, circular DNA templates; DMTr, dimethoxytrityl; DTT, dithiothreitol; FAM, 5'-carboxyfluoresceine; LT, linear single-stranded DNA templates; RCA, rolling circle amplification.
Corresponding author: phone/fax: +7 (347) 235-6088; e-mail: sakhabutdinova.a.r@gmail.com.
Rights and permissions
About this article
Cite this article
Sakhabutdinova, A.R., Mirsaeva, L.R., Oscorbin, I.P. et al. Elimination of DNA Multimerization Arising from Isothermal Amplification in the Presence of Bst Exo– DNA Polymerase. Russ J Bioorg Chem 46, 52–59 (2020). https://doi.org/10.1134/S1068162020010082
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1134/S1068162020010082