Abstract
Fourier transform infrared (FTIR) spectroscopy has proven to be a non-invasive tool to analyse cells without the hurdle of employing exogenous dyes or probes. Nevertheless, the study of single live bacteria in their aqueous environment has long remained a big challenge, due to the strong infrared absorption of water and the small size of bacteria compared to the micron-range infrared wavelengths of the probing photons. To record infrared spectra of bacteria in an aqueous environment, at different spatial resolutions, two setups were developed. A custom-built attenuated total reflection inverted microscope was coupled to a synchrotron-based FTIR spectrometer, using a germanium hemisphere. With such a setup, a projected spot size of 1 × 1 μm2 was achieved, which allowed spectral acquisition at the single-cell level in the 1800–1300 cm−1 region. The second setup used a demountable liquid micro-chamber with a thermal source-powered FTIR microscope, in transmission geometry, for probing clusters of a few thousands of live cells in the mid-IR region (4000–975 cm−1). Both setups were applied for studying two strains of a model lactic acid bacterium exhibiting different cryo-resistances. The two approaches allowed the discrimination of both strains and revealed population heterogeneity among bacteria at different spatial resolutions. The multivariate analysis of spectra indicated that the cryo-sensitive cells presented the highest cell heterogeneity and the highest content of proteins with the α-helix structure. Furthermore, the results from clusters of bacterial cells evidenced phosphate and peptidoglycan vibrational bands associated with the cell envelope, as potential markers of resistance to environmental conditions.

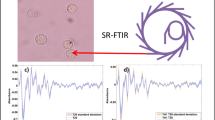
Graphical Abstract







Similar content being viewed by others
References
Holman H-YN, Martin MC, McKinney WR. Synchrotron-based FTIR spectromicroscopy: cytotoxicity and heating considerations. J Biol Phys. 2003;29:275–86.
Miller LM, Dumas P. From structure to cellular mechanism with infrared microspectroscopy. Curr Opin Struct Biol. 2010;20:649–56. https://doi.org/10.1016/j.sbi.2010.07.007.
AlRabiah H, Correa E, Upton M, Goodacre R. High-throughput phenotyping of uropathogenic E. coli isolates with Fourier transform infrared spectroscopy. Analyst. 2013;138:1363. https://doi.org/10.1039/c3an36517d.
Winder CL, Goodacre R. Comparison of diffuse-reflectance absorbance and attenuated total reflectance FT-IR for the discrimination of bacteria. Analyst. 2004;129:1118. https://doi.org/10.1039/b408169b.
Udelhoven T, Naumann D, Schmitt J. Development of a hierarchical classification system with artificial neural networks and FT-IR spectra for the identification of bacteria. Appl Spectrosc. 2000;54:1471–9. https://doi.org/10.1366/0003702001948619.
Naumann D. Infrared spectroscopy in microbiology. In: Encyclopedia of analytical chemistry: John Wiley & Sons, Ltd.; 2006.
Alvarez-Ordóñez A, Mouwen DJM, López M, Prieto M. Fourier transform infrared spectroscopy as a tool to characterize molecular composition and stress response in foodborne pathogenic bacteria. J Microbiol Methods. 2011;84:369–78. https://doi.org/10.1016/j.mimet.2011.01.009.
Saulou C, Jamme F, Maranges C, Fourquaux I, Despax B, Raynaud P, et al. Synchrotron FTIR microspectroscopy of the yeast Saccharomyces cerevisiae after exposure to plasma-deposited nanosilver-containing coating. Anal Bioanal Chem. 2010;396:1441–50. https://doi.org/10.1007/s00216-009-3316-5.
Lu X, Liu Q, Wu D, Al-Qadiri HM, Al-Alami NI, Kang D-H, et al. Using of infrared spectroscopy to study the survival and injury of Escherichia coli O157:H7, Campylobacter jejuni and Pseudomonas aeruginosa under cold stress in low nutrient media. Food Microbiol. 2011;28:537–46. https://doi.org/10.1016/j.fm.2010.11.002.
Passot S, Gautier J, Jamme F, Cenard S, Dumas P, Fonseca F. Understanding the cryotolerance of lactic acid bacteria using combined synchrotron infrared and fluorescence microscopies. Analyst. 2015;140:5920–8. https://doi.org/10.1039/C5AN00654F.
Saulou C, Jamme F, Girbal L, Maranges C, Fourquaux I, Cocaign-Bousquet M, et al. Synchrotron FTIR microspectroscopy of Escherichia coli at single-cell scale under silver-induced stress conditions. Anal Bioanal Chem. 2013;405:2685–97. https://doi.org/10.1007/s00216-013-6725-4.
Gazi E, Dwyer J, Lockyer NP, Miyan J, Gardner P, Hart C, et al. Fixation protocols for subcellular imaging by synchrotron-based Fourier transform infrared microspectroscopy. Biopolymers. 2005;77:18–30. https://doi.org/10.1002/bip.20167.
Lyng F, Gazi E, Gardner P. Preparation of tissues and cells for infrared and Raman spectroscopy and imaging. In: Biomedical applications of synchrotron infrared microspectroscopy, Royal Society of Chemistry. Dublin: D. Moss; 2011. p. 147–85.
Vaccari L, Birarda G, Businaro L, Pacor S, Grenci G. Infrared microspectroscopy of live cells in microfluidic devices (MD-IRMS): toward a powerful label-free cell-based assay. Anal Chem. 2013;84:4768–75.
Xie X, Zubarev RA. Effects of low-level deuterium enrichment on bacterial growth. PLoS One. 2014;9:e102071. https://doi.org/10.1371/journal.pone.0102071.
Holman H-YN, Miles R, Hao Z, Wozei E, Anderson LM, Yang H. Real-time chemical imaging of bacterial activity in biofilms using open-channel microfluidics and synchrotron FTIR spectromicroscopy. Anal Chem. 2009;81:8564–70. https://doi.org/10.1021/ac9015424.
Tobin MJ, Puskar L, Barber RL, Harvey EC, Heraud P, Wood BR, et al. FTIR spectroscopy of single live cells in aqueous media by synchrotron IR microscopy using microfabricated sample holders. Vib Spectrosc. 2010;53:34–8. https://doi.org/10.1016/j.vibspec.2010.02.005.
Vaccari L, Birada G, Grenci G, Pacor S, Businaro L. Synchrotron radiation infrared microspectroscopy of single living cells in microfluidic devices: advantages, disadvantages and future perspectives. J Phys Conf Ser. 2012;359:012007. https://doi.org/10.1088/1742-6596/359/1/012007.
Doherty J, Raoof A, Hussain A, Wolna M, Cinque G, Brown M, et al. Live single cell analysis using synchrotron FTIR microspectroscopy: development of a simple dynamic flow system for prolonged sample viability. Analyst. 2019;144:997–1007. https://doi.org/10.1039/C8AN01566J.
Chan KLA, Fale PLV, Atharawi A, Wehbe K, Cinque G. Subcellular mapping of living cells via synchrotron microFTIR and ZnS hemispheres. Anal Bioanal Chem. 2018;410:6477–87. https://doi.org/10.1007/s00216-018-1245-x.
Chan KLA, Altharawi A, Fale P, Song CL, Kazarian SG, Cinque G, et al. Transmission Fourier transform infrared spectroscopic imaging, mapping, and synchrotron scanning microscopy with zinc sulfide hemispheres on living mammalian cells at sub-cellular resolution. Appl Spectrosc. 2020;74:544–52. https://doi.org/10.1177/0003702819898275.
Doherty J, Zhang Z, Wehbe K, Cinque G, Gardner P, Denbigh J. Increased optical pathlength through aqueous media for the infrared microanalysis of live cells. Anal Bioanal Chem. 2018;410:5779–89. https://doi.org/10.1007/s00216-018-1188-2.
Marcotte L, Therien-Aubin H, Sandt C, Barbeau J, Lafleur M. Solute size effects on the diffusion in biofilms of Streptococcus mutans. Biofouling. 2004;20:189–201. https://doi.org/10.1080/08927010400010494.
Delille A, Quiles F, Humbert F. In situ monitoring of the nascent Pseudomonas fluorescens biofilm response to variations in the dissolved organic carbon level in low-nutrient water by attenuated total reflectance-Fourier transform infrared spectroscopy. Appl Environ Microbiol. 2007;73:5782–8. https://doi.org/10.1128/AEM.00838-07.
Comeau JWD, Pink J, Bezanson E, Douglas CD, Pink D, Smith-Palmer T. A comparison of Pseudomonas Aeruginosa biofilm development on ZnSe and TiO 2 using attenuated total reflection Fourier transform infrared spectroscopy. Appl Spectrosc. 2009;63:1000–7. https://doi.org/10.1366/000370209789379259.
Quilès F, Humbert F, Delille A. Analysis of changes in attenuated total reflection FTIR fingerprints of Pseudomonas fluorescens from planktonic state to nascent biofilm state. Spectrochim Acta A. 2010;75:610–6. https://doi.org/10.1016/j.saa.2009.11.026.
Kuimova MK, Chan KLA, Kazarian SG. Chemical imaging of live cancer cells in the natural aqueous environment. Appl Spectrosc. 2009;63:164–71. https://doi.org/10.1366/000370209787391969.
Kazarian SG, Chan KLA. ATR-FTIR spectroscopic imaging: recent advances and applications to biological systems. Analyst. 2013;138:1940–51. https://doi.org/10.1039/C3AN36865C.
Meneghel J, Passot S, Dupont S, Fonseca F. Biophysical characterization of the Lactobacillus delbrueckii subsp. bulgaricus membrane during cold and osmotic stress and its relevance for cryopreservation. Appl Microbiol Biotechnol. 2017;101:1427–41. https://doi.org/10.1007/s00253-016-7935-4.
Meneghel J, Passot S, Cenard S, Réfrégiers M, Jamme F, Fonseca F. Subcellular membrane fluidity of Lactobacillus delbrueckii subsp. bulgaricus under cold and osmotic stress. Appl Microbiol Biotechnol. 2017;101:6907–17. https://doi.org/10.1007/s00253-017-8444-9.
Kong J, Yu S. Fourier transform infrared spectroscopic analysis of protein secondary structures. Acta Biochim Biophys Sinica. 2007;39:549–59. https://doi.org/10.1111/j.1745-7270.2007.00320.x.
Delcour J, Ferain T, Deghorain M, Palumbo E, Hols P. The biosynthesis and functionality of the cell-wall of lactic acid bacteria. Anton Leeuw Int J Gen Mol Microbiol. 1999;76:159–84.
Kleerebezem M, Hols P, Bernard E, Rolain T, Zhou M, Siezen RJ, et al. The extracellular biology of the lactobacilli. FEMS Microbiol Rev. 2010;34:199–230. https://doi.org/10.1111/j.1574-6976.2009.00208.x.
Chapot-Chartier M-P. Interactions of the cell-wall glycopolymers of lactic acid bacteria with their bacteriophages. Front Microbiol. 2014;5:236. https://doi.org/10.3389/fmicb.2014.00236.
Barth A. Infrared spectroscopy of proteins. Biochim Biophys Acta Bioenerg. 2007;1767:1073–101. https://doi.org/10.1016/j.bbabio.2007.06.004.
Yin X, Salemi MR, Phinney BS, Gotcheva V, Angelov A, Marco ML. Proteomes of Lactobacillus delbrueckii subsp. bulgaricus LBB.B5 incubated in milk at optimal and low temperatures. mSystems. 2017;2:e00027-17. https://doi.org/10.1128/mSystems.00027-17.
De Angelis M, Calasso M, Cavallo N, Di Cagno R, Gobbetti M. Functional proteomics within the genus Lactobacillus. Proteomics. 2016;16:946–62. https://doi.org/10.1002/pmic.201500117.
Streit F, Delettre J, Corrieu G, Béal C. Acid adaptation of Lactobacillus delbrueckii subsp. bulgaricus induces physiological responses at membrane and cytosolic levels that improves cryotolerance. J Appl Microbiol. 2008;105:1071–80. https://doi.org/10.1111/j.1365-2672.2008.03848.x.
Serror P, Dervyn R, Ehrlich SD, Maguin E. csp -like genes of Lactobacillus delbrueckii ssp. bulgaricus and their response to cold shock. FEMS Microbiol Lett. 2003;226:323–30. https://doi.org/10.1016/S0378-1097(03)00594-9.
Schmid A, Kortmann H, Dittrich PS, Blank LM. Chemical and biological single cell analysis. Curr Opin Biotech. 2010;21:12–20. https://doi.org/10.1016/j.copbio.2010.01.007.
Wang D, Bodovitz S. Single cell analysis: the new frontier in ‘omics’. Trends Biotechnol. 2010;28:281–90. https://doi.org/10.1016/j.tibtech.2010.03.002.
Lencastre Fernandes R, Nierychlo M, Lundin L, Pedersen AE, Puentes Tellez PE, Dutta A, et al. Experimental methods and modeling techniques for description of cell population heterogeneity. Biotechnol Adv. 2011;29:575–99. https://doi.org/10.1016/j.biotechadv.2011.03.007.
Brehm-Stecher BF, Johnson EA. Single-cell microbiology: tools, technologies, and applications. Microbiol Mol Biol Rev. 2004;68:538–59. https://doi.org/10.1128/MMBR.68.3.538-559.2004.
Naumann D, Barnickel G, Bradaczek H, Labischinski H, Giesbrecht P. Infrared spectroscopy, a tool for probing bacterial peptidoglycan: potentialities of infrared spectroscopy for cell wall analytical studies and rejection of models based on crystalline chitin. Eur J Biochem. 1982;125:505–15. https://doi.org/10.1111/j.1432-1033.1982.tb06711.x.
Schuster KC, Urlaub E, Gapes JR. Single-cell analysis of bacteria by Raman microscopy: spectral information on the chemical composition of cells and on the heterogeneity in a culture. J Microbiol Methods. 2000;42:29–38. https://doi.org/10.1016/S0167-7012(00)00169-X.
Choo-Smith LP, Maquelin K, van Vreeswijk T, Bruining HA, Puppels GJ, Thi NAN, et al. Investigating microbial (micro)colony heterogeneity by vibrational spectroscopy. Appl Environ Microbiol. 2001;67:1461–9. https://doi.org/10.1128/aem.67.4.1461-1469.2001.
Acknowledgments
We are grateful to the beamline staff for their cooperative involvement in designing and implementing the new experimental device and for providing assistance during the experimental work and for data treatment.
Funding
This work was supported by the French National Research Institute for Agriculture, Food and the Environment (INRAE) and the French National Research Agency (ANR) under the Investing in the Future Program, Grant No. ANR-10-IDEX-0003-02. It was performed at the French national synchrotron facility SOLEIL (Gif-sur-Yvette, France) at the SMIS beamline (Proposal No. 201408998 and 20150220).
Author information
Authors and Affiliations
Contributions
JM, SP, FF and PD conceived the experimental procedure; JM, SP and FF determined the best samples to be studied; SL and PD designed and constructed the setups; JM, SP, PL and FF conducted the experiments; FJ developed an in-house MATLAB script for water subtraction; JM, SP and FF analysed the data; JM, SP and FF contributed to the thorough interpretation and manuscript writing; and JM, SP, FJ, FF and PD provided the critical assessment of the scientific content.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Consent for publication
All authors had full access to the data and approved the manuscript before submission.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
ESM 1
(PDF 931 kb)
Rights and permissions
About this article
Cite this article
Meneghel, J., Passot, S., Jamme, F. et al. FTIR micro-spectroscopy using synchrotron-based and thermal source-based radiation for probing live bacteria. Anal Bioanal Chem 412, 7049–7061 (2020). https://doi.org/10.1007/s00216-020-02835-x
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00216-020-02835-x