Abstract
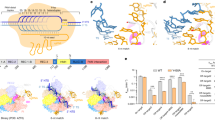
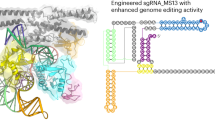
Cas9 nucleases recognize and cleave their target DNA through base pairing of a guide RNA with a spacer adjacent to a protospacer adjacent motif (PAM). Streptococcus thermophilus Cas9 (St1Cas9), a smaller Cas9 orthologue than Streptococcus pyogenes Cas9, enables robust genome editing in diverse organisms. Here we report high-resolution structures of St1Cas9 in complex with a single-guide RNA and different PAM-containing DNAs. All of the structures represent an HNH catalytic state that is rarely observed in other Cas9 structures, clearly depicting the active conformation. A unique wing region in the REC domain forms intensive interactions with the HNH domain, playing a key role in regulating St1Cas9 DNA cleavage activity and probably stabilizing the active conformation. Furthermore, St1Cas9 applies a strategy distinct from those of other Cas9 orthologues for PAM recognition. Structure-guided engineering of St1Cas9 substantially expanded its targeting scope. These molecular-level characterizations will facilitate the rational engineering of St1Cas9.

This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
Conservation analysis of the active-site residues of the HNH domains was performed from the Cas9 protein family (InterPro accession: IPR028629). The structures of St1Cas9 in complex with different PAMs have been deposited in the Protein Data Bank under the accession codes: St1Cas9/GGAA (6M0V), St1Cas9/AGAA (6M0W) and St1Cas9/AGGA (6M0X). The data that support the plots within this paper and other findings of this study are available from the corresponding author on reasonable request. Source data are provided with this paper.
References
Mojica, F. J., Diez-Villasenor, C., Garcia-Martinez, J. & Soria, E. Intervening sequences of regularly spaced prokaryotic repeats derive from foreign genetic elements. J. Mol. Evol. 60, 174–182 (2005).
Makarova, K. S., Grishin, N. V., Shabalina, S. A., Wolf, Y. I. & Koonin, E. V. A putative RNA-interference-based immune system in prokaryotes: computational analysis of the predicted enzymatic machinery, functional analogies with eukaryotic RNAi, and hypothetical mechanisms of action. Biol Direct 1, 7 (2006).
Barrangou, R. et al. CRISPR provides acquired resistance against viruses in prokaryotes. Science 315, 1709–1712 (2007).
Garneau, J. E. et al. The CRISPR/Cas bacterial immune system cleaves bacteriophage and plasmid DNA. Nature 468, 67–71 (2010).
Sapranauskas, R. et al. The Streptococcus thermophilus CRISPR/Cas system provides immunity in Escherichia coli. Nucleic Acids Res. 39, 9275–9282 (2011).
Gasiunas, G., Barrangou, R., Horvath, P. & Siksnys, V. Cas9-crRNA ribonucleoprotein complex mediates specific DNA cleavage for adaptive immunity in bacteria. Proc. Natl Acad. Sci. USA 109, E2579–2586 (2012).
Jinek, M. et al. A programmable dual-RNA-guided DNA endonuclease in adaptive bacterial immunity. Science 337, 816–821 (2012).
Cong, L. et al. Multiplex genome engineering using CRISPR/Cas systems. Science 339, 819–823 (2013).
Mali, P. et al. RNA-guided human genome engineering via Cas9. Science 339, 823–826 (2013).
Jiang, W., Bikard, D., Cox, D., Zhang, F. & Marraffini, L. A. RNA-guided editing of bacterial genomes using CRISPR–Cas systems. Nat. Biotechnol. 31, 233–239 (2013).
Chen, W., Zhang, Y., Yeo, W. S., Bae, T. & Ji, Q. Rapid and efficient genome editing in Staphylococcus aureus by using an engineered CRISPR/Cas9 system. J. Am. Chem. Soc. 139, 3790–3795 (2017).
Bao, Z. et al. Homology-integrated CRISPR–Cas (HI-CRISPR) system for one-step multigene disruption in Saccharomyces cerevisiae. ACS Synth. Biol. 4, 585–594 (2015).
Fu, Y. et al. High-frequency off-target mutagenesis induced by CRISPR–Cas nucleases in human cells. Nat. Biotechnol. 31, 822–826 (2013).
Sternberg, S. H., Redding, S., Jinek, M., Greene, E. C. & Doudna, J. A. DNA interrogation by the CRISPR RNA-guided endonuclease Cas9. Nature 507, 62–67 (2014).
Ran, F. A. et al. In vivo genome editing using Staphylococcus aureus Cas9. Nature 520, 186–191 (2015).
Rock, J. M. et al. Programmable transcriptional repression in mycobacteria using an orthogonal CRISPR interference platform. Nat. Microbiol. 2, 16274 (2017).
Muller, M. et al. Streptococcus thermophilus CRISPR–Cas9 systems enable specific editing of the human genome. Mol. Ther. 24, 636–644 (2016).
Kim, E. et al. In vivo genome editing with a small Cas9 orthologue derived from Campylobacter jejuni. Nat. Commun. 8, 14500 (2017).
Ibraheim, R. et al. All-in-one adeno-associated virus delivery and genome editing by Neisseria meningitidis Cas9 in vivo. Genome Biol. 19, 137 (2018).
Kleinstiver, B. P. et al. Engineered CRISPR–Cas9 nucleases with altered PAM specificities. Nature 523, 481–485 (2015).
Esvelt, K. M. et al. Orthogonal Cas9 proteins for RNA-guided gene regulation and editing. Nat. Methods 10, 1116–1121 (2013).
Wang, H., La Russa, M. & Qi, L. S. CRISPR/Cas9 in genome editing and beyond. Annu. Rev. Biochem. 85, 227–264 (2016).
Barrangou, R. & Horvath, P. A decade of discovery: CRISPR functions and applications. Nat. Microbiol. 2, 17092 (2017).
Hille, F. et al. The biology of CRISPR–Cas: backward and forward. Cell 172, 1239–1259 (2018).
Hirano, H. et al. Structure and engineering of Francisella novicida Cas9. Cell 164, 950–961 (2016).
Acharya, S. et al. Francisella novicida Cas9 interrogates genomic DNA with very high specificity and can be used for mammalian genome editing. Proc. Natl Acad. Sci. USA 116, 20959–20968 (2019).
Agudelo, D. et al. Versatile and robust genome editing with Streptococcus thermophilus CRISPR1-Cas9. Genome Res. 30, 107–117 (2020).
Kleinstiver, B. P. et al. Broadening the targeting range of Staphylococcus aureus CRISPR–Cas9 by modifying PAM recognition. Nat Biotechnol. 33, 1293–1298 (2015).
Hu, J. H. et al. Evolved Cas9 variants with broad PAM compatibility and high DNA specificity. Nature 556, 57–63 (2018).
Nishimasu, H. et al. Engineered CRISPR–Cas9 nuclease with expanded targeting space. Science 361, 1259–1262 (2018).
Miller, S. M. et al. Continuous evolution of SpCas9 variants compatible with non-G PAMs. Nat. Biotechnol. 38, 471–481 (2020).
Walton, R. T., Christie, K. A., Whittaker, M. N. & Kleinstiver, B. P. Unconstrained genome targeting with near-PAMless engineered CRISPR–Cas9 variants. Science 368, 290–296 (2020).
Kleinstiver, B. P. et al. High-fidelity CRISPR–Cas9 nucleases with no detectable genome-wide off-target effects. Nature 529, 490–495 (2016).
Slaymaker, I. M. et al. Rationally engineered Cas9 nucleases with improved specificity. Science 351, 84–88 (2016).
Chen, J. S. et al. Enhanced proofreading governs CRISPR–Cas9 targeting accuracy. Nature 550, 407–410 (2017).
Casini, A. et al. A highly specific SpCas9 variant is identified by in vivo screening in yeast. Nat. Biotechnol. 36, 265–271 (2018).
Tan, Y. et al. Rationally engineered Staphylococcus aureus Cas9 nucleases with high genome-wide specificity. Proc. Natl Acad. Sci. USA 116, 20969–20976 (2019).
Bratovic, M. et al. Bridge helix arginines play a critical role in Cas9 sensitivity to mismatches. Nat. Chem. Biol. 16, 587–595 (2020).
Anders, C., Niewoehner, O., Duerst, A. & Jinek, M. Structural basis of PAM-dependent target DNA recognition by the Cas9 endonuclease. Nature 513, 569–573 (2014).
Jinek, M. et al. Structures of Cas9 endonucleases reveal RNA-mediated conformational activation. Science 343, 1247997 (2014).
Nishimasu, H. et al. Crystal structure of Cas9 in complex with guide RNA and target DNA. Cell 156, 935–949 (2014).
Jiang, F., Zhou, K., Ma, L., Gressel, S. & Doudna, J. A. STRUCTURAL BIOLOGY. A Cas9-guide RNA complex preorganized for target DNA recognition. Science 348, 1477–1481 (2015).
Nishimasu, H. et al. Crystal structure of Staphylococcus aureus Cas9. Cell 162, 1113–1126 (2015).
Zhu, X. et al. Cryo-EM structures reveal coordinated domain motions that govern DNA cleavage by Cas9. Nat. Struct. Mol. Biol. 26, 679 (2019).
Zuo, Z. et al. Structural and functional insights into the bona fide catalytic state of Streptococcus pyogenes Cas9 HNH nuclease domain. eLife 8, e46500 (2019).
Sun, W. et al. Structures of Neisseria meningitidis Cas9 complexes in catalytically poised and anti-CRISPR-inhibited states. Mol. Cell 76, 938 (2019).
Fuchsbauer, O. et al. Cas9 allosteric inhibition by the anti-CRISPR protein AcrIIA6. Mol. Cell 76, 922–937 (2019).
Yamada, M. et al. Crystal structure of the minimal Cas9 from Campylobacter jejuni reveals the molecular diversity in the CRISPR–Cas9 systems. Mol. Cell 65, 1109–1121 (2017).
Hirano, S. et al. Structural basis for the promiscuous PAM recognition by Corynebacterium diphtheriae Cas9. Nat. Commun. 10, 1968 (2019).
Palermo, G. et al. Key role of the REC lobe during CRISPR–Cas9 activation by ‘sensing’, ‘regulating’, and ‘locking’ the catalytic HNH domain. Q. Rev. Biophys. 51, e91 (2018).
Guo, M. H. et al. Structural insights into a high fidelity variant of SpCas9. Cell Res. 29, 183–192 (2019).
Jiang, F. & Doudna, J. A. CRISPR–Cas9 structures and mechanisms. Annu. Rev. Biophys. 46, 505–529 (2017).
Komor, A. C., Kim, Y. B., Packer, M. S., Zuris, J. A. & Liu, D. R. Programmable editing of a target base in genomic DNA without double-stranded DNA cleavage. Nature 533, 420–424 (2016).
Nishida, K. et al. Targeted nucleotide editing using hybrid prokaryotic and vertebrate adaptive immune systems. Science 353, aaf8729 (2016).
Gaudelli, N. M. et al. Programmable base editing of A*T to G*C in genomic DNA without DNA cleavage. Nature 551, 464–471 (2017).
Kim, Y. B. et al. Increasing the genome-targeting scope and precision of base editing with engineered Cas9-cytidine deaminase fusions. Nat. Biotechnol. 35, 371–376 (2017).
Komor, A. C. et al. Improved base excision repair inhibition and bacteriophage Mu Gam protein yields C:G-to-T:A base editors with higher efficiency and product purity. Sci. Adv. 3, eaao4774 (2017).
Li, X. et al. Base editing with a Cpf1-cytidine deaminase fusion. Nat. Biotechnol. 36, 324–327 (2018).
Gibson, D. G. et al. Enzymatic assembly of DNA molecules up to several hundred kilobases. Nat Methods 6, 343–345 (2009).
Minor, W., Cymborowski, M., Otwinowski, Z. & Chruszcz, M. HKL-3000: the integration of data reduction and structure solution–from diffraction images to an initial model in minutes. Acta Crystallogr. D 62, 859–866 (2006).
Bailey, S. The Ccp4 Suite—programs for protein crystallography. Acta Crystallogr. D 50, 760–763 (1994).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D 60, 2126–2132 (2004).
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D 66, 213–221 (2010).
Katoh, K. & Standley, D. M. MAFFT multiple sequence alignment software version 7: improvements in performance and usability. Mol. Biol. Evol. 30, 772–780 (2013).
Kumar, S., Stecher, G. & Tamura, K. MEGA7: molecular evolutionary genetics analysis version 7.0 for bigger datasets. Mol. Biol. Evol. 33, 1870–1874 (2016).
Wang, Y. et al. CRISPR–Cas9 and CRISPR-assisted cytidine deaminase enable precise and efficient genome editing in Klebsiella pneumoniae. Appl. Environ. Microbiol. 84, e01834-18 (2018).
Kluesner, M. G. et al. EditR: a method to quantify base editing from Sanger sequencing. CRISPR J. 1, 239–250 (2018).
Acknowledgements
We thank staff from BL19U1 beamlines of the National Facility for Protein Science Shanghai at Shanghai Synchrotron Radiation Facility for help with data collection. This work was financially supported by the National Natural Science Foundation of China (grant nos. 21922705, 91753127 and 31700123), National Key R&D Program of China Grant (no. 2017YFA0506800) and the Shanghai Science and Technology Committee Rising-Star Program (grant no. 19QA1406000) to Q.J.
Author information
Authors and Affiliations
Contributions
Q.J. conceived and directed the project. Y.Z. and Q.J. designed the research. Y.Z. performed most of the experiments. H.Z., Z.W. and N.T. helped express and purify St1Cas9 proteins. H.Z., Yujue Wang, W.C., Yannan Wang and Yu Wang helped perform the cytosine base editing assay in K. pneumoniae. H.Z., Yujue Wang, W.C., Yannan Wang, Z.W., N.T. and Yu Wang helped with other experiments. X.X. and S.Z. performed the bioinformatics analysis. Y.Z. and Q.J. analysed the data. Y.Z., J.G. and Q.J. carried out crystallographic studies. Y.Z. and Q.J. wrote the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Bacterial survival assay of St1Cas9 with full-length or truncated sgRNA.
a, Scheme of St1Cas9-mediated bacterial survival assay in Klebsiella pneumoniae cells. b, The dhaB3 spacer-introduced pCasKP-St1Cas9 plasmids containing the full-length sgRNA (middle) or the sgRNA with the deletion of stem loop2 (right) efficiently killed K. pneumoniae cells. The empty pCasKP-St1Cas9 was transformed into K. pneumoniae cells as a control (left).
Extended Data Fig. 2 The unique wing region is only present in St1Cas9 among the currently determined Cas9 structures.
a, The REC lobe of St1Cas9 in this study. The wing region is highlighted by the pink cycle. b–g, The REC lobes of SpCas9 (b), FnCas9 (c), SaCas9 (d), Nm1Cas9 (e), CdCas9 (f) and CjCas9 (g), respectively.
Extended Data Fig. 3 Interactions between St1Cas9 and the nucleic acids.
St1Cas9 residues that interact with the sgRNA-dsDNA duplex via their main chains are shown in parentheses. Water-mediated hydrogen-bonding interactions are not shown.
Extended Data Fig. 4 Distinct conformations of the Cas9 orthologs.
a, Post-catalytic state of DNA-bound SpCas9. Zoom-in view of the active sites of the SpCas9 HNH domain is shown in the right figure. b and c, Inactive states of DNA-bound SaCas9 (b) and DNA-bound FnCas9 (c). d, Catalytic state of DNA-bound Nm1Cas9. Zoom-in view of the active sites of the Nm1Cas9 HNH domain is shown in the left figure. e, Post-catalytic state of DNA-bound Nm1Cas9. f, Catalytic state of DNA-bound St1Cas9. All distances were measured between the Cα atom of the catalytic residue H and the scissile phosphate of TS nucleotides 3 and 4 in the catalytic or inactive state Cas9 structures (b–d, f), or between the Cα atom of the catalytic residue H and the 5’-PO3 of TS nucleotide 4 in the post-catalytic state Cas9 structures (a, e). The positions of the catalytic residue H were shown as yellow stars.
Extended Data Fig. 5 Catalytic pockets of the HNH domains.
a–c, Close-up view of Mg2+-binding catalytic pockets of the HNH domains of St1Cas9 (a), Nm1Cas9 (b) and AnCas9 (c). Mg2+ and water molecules were coloured green and red, respectively. d, In vitro DNA cleavage assay of the wild type and various mutants of St1Cas9. The linearized plasmid containing a target sequence was used as the substrate and the full-length sgRNA was used as the guide RNA in the assay.
Extended Data Fig. 6 DNA binding activities of the wild type St1Cas9 and the variant with the truncation of the wing region.
Electrophoretic mobility shift assay was performed using catalytic inactive dCas9-sgRNA complexes and the FAM-labelled target dsDNA.
Extended Data Fig. 7 Phylogenetic tree of putative wing region containing Cas9s.
Each sequence represents sequences with >70% sequence identity.
Extended Data Fig. 8 In vitro cleavage activities of wild type St1Cas9 and the variant with the truncation of the wing region.
a, The spacer sequences of the linearized plasmids with single or double mismatches at positions 20-16. b, In vitro DNA cleavage assay of wild type St1Cas9. The linearized plasmids containing double mismatches at positions 20-16 were used as the substrates. c, In vitro DNA cleavage assay of wild type St1Cas9. The linearized plasmids containing single mismatches at positions 20-16 were used as the substrates. d, In vitro DNA cleavage assay of wild type St1Cas9 and the variant with the truncation of the wing region. The linearized plasmids containing single mismatches at positions 20 or 19 were used as the substrates. The reaction condition of wild-type St1Cas9 was 5 nM linearized plasmid, 120 nM Cas9, and 240 nM sgRNA. The reaction condition of the St1Cas9 variant with the truncation of the wing region was 5 nM linearized plasmid, 500 nM Cas9, and 1 µM sgRNA.
Extended Data Fig. 9 In vitro cleavage activities of the wild type and various mutants of St1Cas9.
a, In vitro DNA cleavage assay of the wild type and various K1086 mutants of St1Cas9. The linearized plasmids containing the 5’-AAANAA-3’ PAM sequences were used as the substrates. b, In vitro DNA cleavage assay of the wild type and K1086L mutant of St1Cas9. The linearized plasmids containing the 5’-AANTAA-3’ PAM sequences were used as the substrates. c, In vitro DNA cleavage assay of the wild type and various engineered mutants of St1Cas9. The linearized plasmids containing the 5’-AARTAA-3’ PAM sequences were used as the substrates.
Extended Data Fig. 10 Determination of the editing window of St1Cas9-mediated cytosine base editing system in K. pneumoniae.
a, Scheme of St1Cas9-mediated cytosine base editing. b, The editing results of various C-rich spacers in the KP_1.6366 strain. The Cs were coloured red and the target PAM sites were coloured blue. The experiment was performed in triplicate. Error bars report standard deviations of the mean (s.d.).
Supplementary information
Supplementary Information
Supplementary Figs. 1–4 and Tables 1–4.
Source data
Source Data Fig. 1
Unprocessed gels.
Source Data Fig. 2
Unprocessed gels.
Source Data Fig. 3
Unprocessed gels.
Source Data Fig. 4
Unprocessed gels.
Source Data Fig. 5
Unprocessed gels.
Source Data Fig. 5
Base editing in K. pneumoniae.
Source Data Extended Data Fig. 1
Unprocessed gels.
Source Data Extended Data Fig. 2
Unprocessed gels.
Source Data Extended Data Fig. 3
Unprocessed gels.
Source Data Extended Data Fig. 4
Unprocessed gels.
Rights and permissions
About this article
Cite this article
Zhang, Y., Zhang, H., Xu, X. et al. Catalytic-state structure and engineering of Streptococcus thermophilus Cas9. Nat Catal 3, 813–823 (2020). https://doi.org/10.1038/s41929-020-00506-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41929-020-00506-9
This article is cited by
-
Revolutionizing Agriculture: Harnessing CRISPR/Cas9 for Crop Enhancement
Indian Journal of Microbiology (2024)
-
Structural basis for mismatch surveillance by CRISPR–Cas9
Nature (2022)
-
R-loop formation and conformational activation mechanisms of Cas9
Nature (2022)
-
Structural biology of CRISPR–Cas immunity and genome editing enzymes
Nature Reviews Microbiology (2022)
-
Programmed genome editing by a miniature CRISPR-Cas12f nuclease
Nature Chemical Biology (2021)