Abstract
Background
Alu elements are most abundant retrotransposons with > 1.2 million copies in the primate genome. AluYb8 subfamily was diverged from AluY lineage, and has accumulated eight diagnostic mutations and 7-bp duplication during primate evolution. A total of 1851 AluYb copies are present in the human genome, and most of them are human-specific. On the other hand, only a few AluYb8 copies were identified in the chimpanzee genome by previous studies on AluYb8. The significantly different number of species-specific AluYb8 elements between human and chimpanzee might result from the incompletion of chimpanzee reference genome sequences at the time of the previous study.
Objective
AluYb8 elements could generate genomic structural variations in the chimpanzee genome. This study aimed to identify and characterize chimpanzee-specific AluYb elements using the most updated chimpanzee reference genome sequences (Jan. 2018, panTro6).
Methods
To identify chimpanzee-specific AluYb8, we carried out genomic comparison with non-chimpanzee primate genome using the UCSC table browser. In addition, chimpanzee-specific AluYb8 candidates were manually inspected and experimentally verified using PCR and Sanger sequencing.
Results
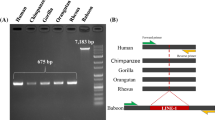
Among a total of 231 chimpanzee-specific AluYb8 candidates, 11 of the candidates are chimpanzee-specific AluYb8, and 29 elements are shared between the chimpanzee and non-chimpanzee primate genomes. Through the sequence analysis of AluYb8 and other Alu subfamilies, we were able to observe various diagnostic mutations and variable length duplications in 7-bp duplication region of AluYb8 element. In addition, we further validated two of the chimpanzee-specific AluYb8 elements (CS8 and CS20) that were not previously discovered by display PCR and Sanger sequencing. Interestingly, we identified a AluYb8 insertion-mediated deletion (CS8 locus) in the chimpanzee genome.
Conclusion
Our study found that AluYb8 elements are much more abundant in the human genome than chimpanzee genome, and that it could be due to the absence of hyperactive “master” AluYb8 elements in the chimpanzee genome.



Similar content being viewed by others
References
Ahmed M, Li W, Liang P (2013) Identification of three new Alu Yb subfamilies by source tracking of recently integrated Alu Yb elements. Mob DNA 4:25
Batzer MA, Deininger PL (2002) Alu repeats and human genomic diversity. Nat Rev Genet 3:370–379
Batzer MA, Deininger PL, Hellmann-Blumberg U, Jurka J, Labuda D, Rubin CM, Schmid CW, Zietkiewicz E, Zuckerkandl E (1996) Standardized nomenclature for Alu repeats. J Mol Evol 42:3–6
Callinan PA, Hedges DJ, Salem AH, Xing J, Walker JA, Garber RK, Watkins WS, Bamshad MJ, Jorde LB, Batzer MA (2003) Comprehensive analysis of Alu-associated diversity on the human sex chromosomes. Gene 317:103–110
Carter AB, Salem AH, Hedges DJ, Keegan CN, Kimball B, Walker JA, Watkins WS, Jorde LB, Batzer MA (2004) Genome-wide analysis of the human Alu Yb-lineage. Hum Genom 1:167–178
Chimpanzee S, Analysis C (2005) Initial sequence of the chimpanzee genome and comparison with the human genome. Nature 437:69–87
Feusier J, Witherspoon DJ, Scott Watkins W, Goubert C, Sasani TA, Jorde LB (2017) Discovery of rare, diagnostic AluYb8/9 elements in diverse human populations. Mob DNA 8:9
Hall TA (1999) BioEdit: a user-friendly biological sequence alignment editor and analysis program for Windows 95/98/NT. Nucleic Acids Symp Ser 41:95–98
Han K, Xing J, Wang H, Hedges DJ, Garber RK, Cordaux R, Batzer MA (2005) Under the genomic radar: the stealth model of Alu amplification. Genome Res 15:655–664
Hedges DJ, Callinan PA, Cordaux R, Xing J, Barnes E, Batzer MA (2004) Differential alu mobilization and polymorphism among the human and chimpanzee lineages. Genome Res 14:1068–1075
Jurka J (1993) A new subfamily of recently retroposed human Alu repeats. Nucleic Acids Res 21:2252
Kim S, Cho CS, Han K, Lee J (2016) Structural variation of Alu element and human disease. Genom Inform 14:70–77
Konkel MK, Walker JA, Batzer MA (2010) LINEs and SINEs of primate evolution. Evol Anthropol 19:236–249
Konkel MK, Walker JA, Hotard AB, Ranck MC, Fontenot CC, Storer J, Stewart C, Marth GT, Genomes C, Batzer MA (2015) Sequence analysis and characterization of active human Alu subfamilies based on the 1000 Genomes Pilot Project. Genome Biol Evol 7:2608–2622
Kumar S, Stecher G, Tamura K (2016) MEGA7: molecular evolutionary genetics analysis version 7.0 for bigger datasets. Mol Biol Evol 33:1870–1874
Lander ES, Linton LM, Birren B, Nusbaum C, Zody MC, Baldwin J, Devon K, Dewar K, Doyle M, FitzHugh W et al (2001) Initial sequencing and analysis of the human genome. Nature 409:860–921
Lee J, Cordaux R, Han K, Wang J, Hedges DJ, Liang P, Batzer MA (2007) Different evolutionary fates of recently integrated human and chimpanzee LINE-1 retrotransposons. Gene 390:18–27
Lee S, Tang W, Liang P, Han K (2019) A comprehensive analysis of chimpanzee (Pan troglodytes)-specific LINE-1 retrotransposons. Gene 693:46–51
Ullu E, Tschudi C (1984) Alu sequences are processed 7SL RNA genes. Nature 312:171–172
Untergasser A, Cutcutache I, Koressaar T, Ye J, Faircloth BC, Remm M, Rozen SG (2012) Primer3–new capabilities and interfaces. Nucleic Acids Res 40:e115
Acknowledgments
This research was supported by Basic Science Research Capacity Enhancement Project through Korea Basic Science Institute (National research Facilities and Equipment Center) grant funded by the Ministry of Education (Grant No. 2019R1A6C1010033). The present research was supported by the research fund of Dankook university in 2019 for the University Innovation Support Program. This work was supported by the Korea Foundation for the Advancement of Science &Creativity (KOFAC), and funded by the Korean Government (MOE) in 2019. The authors would like to thank Cherljoon Lee, Yujun Park, Yunseok Oh, Hyunjune Park, and Seungwon Yang (Dankook University) for their assistance with the initial data collection and analysis.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that there is no conflict of interest regarding the publication of this paper.
Ethical approval
Experiments with the chimpanzee and gorilla were carried out in accordance with the guidelines and regulations approved by the Animal Experimentation Committees of Kyoto University.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
13258_2020_989_MOESM1_ESM.pptx
Supplementary Figure 1. Diagnostic nucleotide position of 11 AluYb8 elements and 29 AluYb-like elements. Blue and Yellow boxes indicate diagnostic mutation position of AluYb8 elements and 7-bp duplication regions, respectively
13258_2020_989_MOESM2_ESM.pptx
Supplementary Figure 2. Sequence alignment of CS8 in human, chimpanzee, and gorilla. Black upper case letters indicate shared flanking unique sequences. The red and blue characters indicate chimpanzee-specific AluYb8 sequences and deleted counterpart sequences in the human and gorilla, respectively
Rights and permissions
About this article
Cite this article
Kim, S., Kim, D.H., Imai, H. et al. A comprehensive analysis of chimpanzee (Pan Troglodytes)-specific AluYb8 element. Genes Genom 42, 1207–1213 (2020). https://doi.org/10.1007/s13258-020-00989-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13258-020-00989-7