Abstract
Targeted downregulation of select endogenous plant genes is known to confer disease or pest resistance in crops and is routinely accomplished via transgenic modification of plants for constitutive gene silencing. An attractive alternative to the use of transgenics or pesticides in agriculture is the use of a ‘green’ alternative known as RNAi, which involves the delivery of siRNAs that downregulate endogenous genes to confer resistance. However, siRNA is a molecule that is highly susceptible to enzymatic degradation and is difficult to deliver across the lignin-rich and multi-layered plant cell wall that poses the dominant physical barrier to biomolecule delivery in plants. We have demonstrated that DNA nanostructures can be utilized as a cargo carrier for direct siRNA delivery and gene silencing in mature plants. The size, shape, compactness and stiffness of the DNA nanostructure affect both internalization into plant cells and subsequent gene silencing efficiency. Herein, we provide a detailed protocol that can be readily adopted with standard biology benchtop equipment to generate geometrically optimized DNA nanostructures for transgene-free and force-independent siRNA delivery and gene silencing in mature plants. We further discuss how such DNA nanostructures can be rationally designed to efficiently enter plant cells and deliver cargoes to mature plants, and provide guidance for DNA nanostructure characterization, storage and use. The protocol described herein can be completed in 4 d.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
All materials are available from commercial sources or can be derived using methods described in this study. All data and controls relevant to the protocol have been included in the Supplementary Information. Raw data such as unprocessed image files can be obtained from the corresponding author upon reasonable request.
References
Napoli, C., Lemieux, C. & Jorgensen, R. Introduction of a chimeric chalcone synthase gene into petunia results in reversible co-suppression of homologous genes in trans. Plant Cell 2, 279–289 (1990).
Davis, M. E. et al. Evidence of RNAi in humans from systemically administered siRNA via targeted nanoparticles. Nature 464, 1067–U1140 (2010).
Fire, A. et al. Potent and specific genetic interference by double-stranded RNA in Caenorhabditis elegans. Nature 391, 806–811 (1998).
Cunningham, F. J., Goh, N. S., Demirer, G. S., Matos, J. L. & Landry, M. P. Nanoparticle-mediated delivery towards advancing plant genetic engineering. Trends Biotechnol. 36, 882–897 (2018).
Demirer, G. S. & Landry, M. P. Delivering genes to plants. Chem. Eng. Prog. 113, 40–45 (2017).
Pereira, A. Plant Reverse Genetics: Methods and Protocols (Humana Press, 2011).
Zhang, J. et al. Vacuum and co-cultivation agroinfiltration of (germinated) seeds results in tobacco rattle virus (TRV) mediated whole-plant virus-induced gene silencing (VIGS) in wheat and maize. Front. Plant Sci. 8, 393 (2017).
da Cunha, N. B. et al. The next generation of antimicrobial peptides (AMPs) as molecular therapeutic tools for the treatment of diseases with social and economic impacts. Drug Discov. Today 22, 234–248 (2017).
Xu, P., Zhang, Y. J., Kang, L., Roossinck, M. J. & Mysore, K. S. Computational estimation and experimental verification of off-target silencing during posttranscriptional gene silencing in plants. Plant Physiol. 142, 429–440 (2006).
Kodama, H. & Komamine, A. RNAi and Plant Gene Function Analysis: Methods and Protocols (Springer, 2011).
Silva, A. T., Nguyen, A., Ye, C., Verchot, J. & Moon, J. H. Conjugated polymer nanoparticles for effective siRNA delivery to tobacco BY-2 protoplasts. BMC Plant Biol. 10, 291 (2010).
Mitter, N. et al. Clay nanosheets for topical delivery of RNAi for sustained protection against plant viruses. Nat. Plants 3, 16207 (2017).
Demirer, G. S. et al. Carbon nanocarriers deliver siRNA to intact plant cells for efficient gene knockdown. Sci. Adv. 6, eaaz0495 (2020).
Sun, W. J. et al. Cocoon-like self-degradable DNA nanoclew for anticancer drug delivery. J. Am. Chem. Soc. 136, 14722–14725 (2014).
Ruan, W. M. et al. DNA nanoclew templated spherical nucleic acids for siRNA delivery. Chem. Commun. 54, 3609–3612 (2018).
Hu, Q. Q., Li, H., Wang, L. H., Gu, H. Z. & Fan, C. H. DNA nanotechnology-enabled drug delivery systems. Chem. Rev. 119, 6459–6506 (2019).
Kostiainen, M. & Linko, V. DNA nanostructures as innovative vehicles for smart drug delivery. Eur. J. Hum. Genet. 27, 768–769 (2019).
Jiang, Q., Liu, S. L., Liu, J. B., Wang, Z. G. & Ding, B. Q. Rationally designed DNA-origami nanomaterials for drug delivery in vivo. Adv. Mater. 31, 1804781–1804786 (2019).
Sun, W. et al. Self-assembled DNA nanoclews for the efficient delivery of CRISPR–Cas9 for genome editing. Ang. Chem. Int. Ed. 54, 12029–12033 (2015).
Madhanagopal, B. R., Zhang, S., Demirel, E., Wady, H. & Chandrasekaran, A. R. DNA nanocarriers: programmed to deliver. Trends Biochem. Sci. 43, 997–1013 (2018).
Linko, V., Ora, A. & Kostiainen, M. A. DNA nanostructures as smart drug-delivery vehicles and molecular devices. Trends Biotechnol. 33, 586–594 (2015).
Li, J., Fan, C., Pei, H., Shi, J. & Huang, Q. Smart drug delivery nanocarriers with self-assembled DNA nanostructures. Adv. Mater. 25, 4386–4396 (2013).
Zhang, H. et al. Programming chain-growth copolymerization of DNA hairpin tiles for in-vitro hierarchical supramolecular organization. Nat. Commun. 10, 1006 (2019).
Li, J. et al. Self-assembled multivalent DNA nanostructures for noninvasive intracellular delivery of immunostimulatory CpG oligonucleotides. Acs Nano 5, 8783–8789 (2011).
Zhang, H. et al. DNA nanostructures coordinate gene silencing in mature plants. Proc. Natl Acad. Sci. USA 116, 7543–7548 (2019).
Bastings, M. M. C. et al. Modulation of the cellular uptake of DNA origami through control over mass and shape. Nano Lett. 18, 3557–3564 (2018).
Webster, D. E. & Thomas, M. C. Post-translational modification of plant-made foreign proteins; glycosylation and beyond. Biotechnol. Adv. 30, 410–418 (2012).
Withers, J. & Dong, X. N. Post-translational regulation of plant immunity. Curr. Opin. Plant Biol. 38, 124–132 (2017).
Bila, H., Kurisinkal, E. E. & Bastings, M. M. C. Engineering a stable future for DNA-origami as a biomaterial. Biomater Sci. 7, 532–541 (2019).
Altpeter, F. et al. Advancing crop transformation in the era of genome editing. Plant Cell 28, 1510–1520 (2016).
Wang, P., Lombi, E., Zhao, F. J. & Kopittke, P. M. Nanotechnology: a new opportunity in plant sciences. Trends Plant Sci. 21, 699–712 (2016).
Baulcombe, D. RNA silencing in plants. Nature 431, 356–363 (2004).
Meister, G. & Tuschl, T. Mechanisms of gene silencing by double-stranded RNA. Nature 431, 343–349 (2004).
Johansen, L. K. & Carrington, J. C. Silencing on the spot. Induction and suppression of RNA silencing in the Agrobacterium-mediated transient expression system. Plant Physiol. 126, 930–938 (2001).
Dalakouras, A. et al. Induction of silencing in plants by high-pressure spraying of in vitro-synthesized small RNAs. Front. Plant Sci. 7, 1327 (2016).
Tang, W., Weidner, D. A., Hu, B. Y., Newton, R. J. & Hu, X. H. Efficient delivery of small interfering RNA to plant cells by a nanosecond pulsed laser-induced stress wave for posttranscriptional gene silencing. Plant Sci. 171, 375–381 (2006).
Cheon, S. H. et al. Effective delivery of siRNA to transgenic rice cells for enhanced transfection using PEI-based polyplexes. Biotechnol. Bioproc. E 22, 577–585 (2017).
Unnamalai, N., Kang, B. G. & Lee, W. S. Cationic oligopeptide-mediated delivery of dsRNA for post-transcriptional gene silencing in plant cells. FEBS Lett. 566, 301–310 (2004). 307.
Numata, K., Ohtani, M., Yoshizumi, T., Demura, T. & Kodama, Y. Local gene silencing in plants via synthetic dsRNA and carrier peptide. Plant Biotechnol. J. 12, 1027–1034 (2014).
Bao, W. L., Wang, J. Y., Wang, Q., O’Hare, D. & Wan, Y. L. Layered double hydroxide nanotransporter for molecule delivery to intact plant cells. Sci. Rep. 6, 26738 (2016).
Schwartz, S. H., Hendrix, B., Hoffer, P., Sanders, R. A. & Zheng, W. Carbon dots for efficient siRNA delivery and gene silencing in plants. Preprint at https://www.biorxiv.org/content/10.1101/722595v1 (2019).
Winfree, E., Liu, F., Wenzler, L. A. & Seeman, N. C. Design and self-assembly of two-dimensional DNA crystals. Nature 394, 539–544 (1998).
Dirks, R. M. & Pierce, N. A. Triggered amplification by hybridization chain reaction. Proc. Natl Acad. Sci. USA 101, 15275–15278 (2004).
Lin, M. H. et al. Programmable engineering of a biosensing interface with tetrahedral DNA nanostructures for ultrasensitive DNA detection. Angew. Chem. Int. Ed. 54, 2151–2155 (2015).
Nicot, N., Hausman, J. F., Hoffmann, L. & Evers, D. Housekeeping gene selection for real-time RT-PCR normalization in potato during biotic and abiotic stress. J. Exp. Bot. 56, 2907–2914 (2005).
Kim, H. et al. ‘Shotgun DNA synthesis’ for the high-throughput construction of large DNA molecules. Nucleic Acids Res. 40, e140 (2012).
Castro, C. E. et al. A primer to scaffolded DNA origami. Nat. Methods 8, 221–229 (2011).
Douglas, S. M. et al. Rapid prototyping of 3D DNA-origami shapes with caDNAno. Nucleic Acids Res. 37, 5001–5006 (2009).
Liu, Q. et al. Carbon nanotubes as molecular transporters for walled plant cells. Nano Lett. 9, 1007–1010 (2009).
Tang, W. et al. Post-transcriptional gene silencing induced by short interfering RNAs in cultured transgenic plant cells. Genomics Proteomics Bioinformatics 2, 97–108 (2004).
Schmittgen, T. D. & Livak, K. J. Analyzing real-time PCR data by the comparative CT method. Nat. Prot. 3, 1101–1108 (2008).
Thomas, N. C. et al. Roq1 confers resistance to Xanthomonas, Pseudomonas syringae and Ralstonia solanacearum in tomato. Preprint at https://www.biorxiv.org/content/10.1101/813758v2 (2020).
Schultink, A., Qi, T. C., Lee, A., Steinbrenner, A. D. & Staskawicz, B. Roq1 mediates recognition of the Xanthomonas and Pseudomonas effector proteins XopQ and HopQ1. Plant J. 92, 787–795 (2017).
Chuang, C. F. & Meyerowitz, E. M. Specific and heritable genetic interference by double-stranded RNA in Arabidopsis thaliana. Proc. Natl Acad. Sci. USA 97, 4985–4990 (2000).
Schweizer, P., Pokorny, J., Schulze-Lefert, P. & Dudler, R. Double-stranded RNA interferes with gene function at the single-cell level in cereals. Plant J. 24, 895–903 (2000).
Klahre, U., Crete, P., Leuenberger, S. A., Iglesias, V. A. & Meins, F. High molecular weight RNAs and small interfering RNAs induce systemic posttranscriptional gene silencing in plants. Proc. Natl Acad. Sci. USA 99, 11981–11986 (2002).
Kwak, S.-Y. et al. Chloroplast-selective gene delivery and expression in planta using chitosan-complexed single-walled carbon nanotube carriers. Nat. Nanotechnol. 14, 447–455 (2019).
Demirer, G. S. et al. High aspect ratio nanomaterials enable delivery of functional genetic material without DNA integration in mature plants. Nat. Nanotechnol. 14, 456–464 (2019).
Acknowledgements
Hu. Z. acknowledges the support of the Chinese National Natural Science Foundation (21605153). The authors acknowledge support from a Burroughs Wellcome Fund Career Award at the Scientific Interface (CASI), a Stanley Fahn PDF Junior Faculty Grant under award no. PF-JFA-1760, a Beckman Foundation Young Investigator Award, a USDA AFRI award, a grant from the Gordon and Betty Moore Foundation, a USDA NIFA award, a USDA-BBT EAGER award, support from the Chan-Zuckerberg Foundation and an FFAR New Innovator Award (to M.P.L.). G.S.D. is supported by a Schlumberger Foundation Faculty for the Future Fellowship. The authors also acknowledge support from UC Berkeley Molecular Imaging Center (supported by the Gordon and Betty Moore Foundation), the QB3 Shared Stem Cell Facility and the Innovative Genomics Institute (IGI).
Author information
Authors and Affiliations
Contributions
Hu. Z. and M.P.L designed the experiments, and C.F. helped with the experimental design. Hu. Z. and Ho. Z. performed the simulations and experiments. Hu. Z. and G.S.D analyzed the data and created the figures. Hu. Z., Ho. Z., G.S.D, and E.G.-G. wrote the manuscript. All authors revised and approved the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Protocols thanks Veikko Linko and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Related links
Key references using this protocol
Zhang, H. et al. Proc. Natl Acad. Sci. USA 116, 7543–7548 (2019): https://doi.org/10.1073/pnas.1818290116
Zhang, H. et al. Nat. Commun. 10, 1006 (2019): https://doi.org/10.1038/s41467-019-09004-4
Integrated supplementary information
Supplementary Figure 1 Representative gel image for characterization of the formation of DNA nanostructures.
a,b,10% native PAGE gel showing the formation of tetrahedron (a) and DHT monomer A (b). Lane1: marker; lane 2: DNA nanostructure.
Supplementary Figure 2 Infiltration of leaves with 1-ml syringe.
Introduce a tiny puncture into the Nb plant leaf with a pipette tip (left) on the leaf abaxial surface. Center the syringe tip at the puncture area (middle), and gently push the syringe plunger until all the liquid is infiltrated (right).
Supplementary Figure 3 Confirmation of strong GFP expression in transgenic mGFP5 Nb plants.
a, Representative confocal images of mGFP5 Nb leaves. Scale bar: 100 µm. b, To validate GFP expression in mGFP5 Nb, a western gel shows the correct GFP band size (~27 kDa) for GFP extracted from mGFP5 Nb leaves.
Supplementary Figure 4 Internalization of Cy3-labeled DHT monomer into three different plant species.
Cy3-labeled HT monomer can internalize into tobacco, arugula and watercress leaf cells. Scale bars, 100 µm.
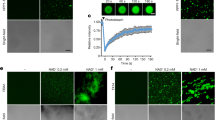
Supplementary Figure 5 Representative confocal images to test the internalization mechanism of the Cy3-labeled DHT monomer into Nb leaves.
a, Temperature dependence of internalization: we observe internalization predominately when plants infiltrated with Cy3-DHT monomers are incubated at 21 °C, but not when incubated at 4 °C, suggesting that the internalization process is energy dependent. Scale bars, 50 µm. b, Leaves pretreated with 33 µM wortmannin (incubated for half an hour before DNA nanostructure infiltration), a chemical inhibitor of endocytosis, show greatly reduced Cy3-DHT nanostructure internalization compared with non-treated counterparts, suggesting that the nanostructures enter the plant cells through receptor-mediated endocytosis.
Supplementary Figure 6 DNA nanostructure-induced GFP silencing is transient.
a, Representative confocal images of mGFP5 Nb leaves 7 d post-infiltration with PBS (control), siRNA–tetrahedron nanostructures, or siRNA–DHT monomer nanostructures, showing GFP fluorescence recovery. Scale bars, 100 µm. b, Quantitative fluorescence intensity analysis of confocal images. n.s.= non significant (s.d., n = 15). c, Statistical analysis and representative western gel of GFP extracted from nanostructure-treated leaves 72 h post-infiltration. n.s.= non significant (s.d., n = 3).
Supplementary Figure 7 Gene silencing pathways for siRNA-linked nanostructures.
a, qPCR of leaves infiltrated with free siRNA, siRNA-loaded nanostring, DHT-c, tetrahedron or DHT-s 1d post-infiltration. ****P < 0.0001 in one-way ANOVA. Error is SEM (n = 4). b, Scheme showing the proposed silencing pathways induced by different siRNA-loaded DNA nanostructures.
Supplementary Figure 8 DNA nanostructures without siRNA loading cause no change in GFP mRNA.
qPCR results from GFP5 Nb leaves infiltrated with DHT monomer alone 2 d post-infiltration showing no mRNA change. Control samples are non-infiltrated leaves. Error bars indicate s.e.m. (n = 4).
Supplementary Figure 9 DNA nanostructure-infiltrated Nb leaves show no stress response as measured by qPCR.
qPCR analysis of NbrbohB, a known plant stress gene, to test the toxicity of DNA nanostructures used to deliver siRNA. Nanostructures do not upregulate the plant stress gene. Control samples are PBS buffer-infiltrated leaves. Error bars indicate s.e.m. (n = 3).
Supplementary Figure 10 Silencing of endogenous functional Nb ROQ1 gene with siRNA loaded on tetrahedron DNA nanostructures.
Free ROQ1 siRNA does not induce significant silencing of the ROQ1 gene, whereas 100 nM ROQ1 siRNA loaded on the tetrahedron nanostructure yields a near 50% decrease of ROQ1 mRNA as assessed by qPCR of infiltrated Nb leaves compared with the non-treated control leaves. ** P = 0.0010 in one-way ANOVA. Error bars indicate s.e.m. (n = 3).
Supplementary Figure 11 DNA nanostructures are gradually degraded in plant cell lysate.
a, 10% native PAGE gel showing incubation of DHT monomer with plant cell lysate at different time points. Lane 1: marker; lane 2: DHT monomer only as control; lanes 3–8: DHT monomer incubated with plant cell lysate at 0 h, 12 h, 24 h, 48 h, 72 h and 96 h at room temperature. b, Normalized band intensity analysis of the gels in a where 100% band intensity is defined as the DHT monomer control alone without the plant cell lysate. Upward deviations from 100% are due to nonspecific protein adsorption to nanostructures that slightly increase the band width and optical density.
Supplementary information
Supplementary Information
Supplementary Figures 1–11, Tables 1 and 2 and Discussion.
Rights and permissions
About this article
Cite this article
Zhang, H., Zhang, H., Demirer, G.S. et al. Engineering DNA nanostructures for siRNA delivery in plants. Nat Protoc 15, 3064–3087 (2020). https://doi.org/10.1038/s41596-020-0370-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41596-020-0370-0
This article is cited by
-
RNAi-based drug design: considerations and future directions
Nature Reviews Drug Discovery (2024)
-
An Overview of Targeted Genome Editing Strategies for Reducing the Biosynthesis of Phytic Acid: an Anti-nutrient in Crop Plants
Molecular Biotechnology (2024)
-
DNA origami
Nature Reviews Methods Primers (2021)
-
Citric acid/β-alanine carbon dots as a novel tool for delivery of plasmid DNA into E. coli cells
Scientific Reports (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.