Abstract
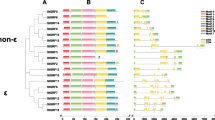
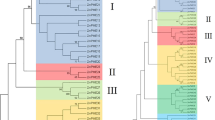
Pectin methylesterase (PME) is a type of protein encoded by a multi-gene family and participates in seed germination, fruit ripening and stress response. To further study the function of the PME gene family in potato, this paper screened and identified potato PME gene family by bioinformatics methods, and analysed the structure, physicochemical properties, conserved domains and evolutionary relationships of the gene sequences. The results showed that the identified 54 potato PME genes can be divided into Group I (only PME domain) and Group II (both PME domain and PMEI domain). The phylogenetic tree constructed by the maximum likelihood method indicated that potato PME genes consisted of five subfamilies. The protein sequences of potato PME genes were compared with clustalx2.1 software. Most of these sequences were identified to have five conserved structural regions including Gx Yx E (Region I), QAVAxR (Region II), QDTL (Region III), GxxDFIFG (Region IV) and YLGRxWx (Region V). Conserved motifs of potato PME amino acid sequences were analysed by MEME online software. Motif1 contains three of the five conserved structural regions and motif2 and motif8 each contain one, indicating that these sequences are more conservative. The structure of 54 potato PME gene sequences was analysed by the GSDS website. It was found that there were only two parts, exon or coding sequence (CDS) between untranslated regions (UTRs), without intron structures in these sequences, indicating that the structure is relatively simple. It will provide a reference for the study on the function of potato PME gene.




Similar content being viewed by others
References
Bailey TL, Williams N, Misleh C, Li WW (2006) MEME: discovering and analyzing DNA and protein sequence motifs. Nucleic Acids Res 34:W369–W373
Bosch M, Hepler PK (2005) Pectin methylesterases and pectin dynamics in pollen tubes. Plant Cell 17:3219–3226
Butenko MA, Patterson SE, Grini PE, Stenvik GE, Amundsen SS, Mandal A, Aalen RB (2003) INFLORESCENCE DEFICIENT IN ABSCISSION controls floral organ abscission in Arabidopsis and identifies a novel family of putative ligands in plants. Plant Cell 15:2296–2307
Collmer AC, Keen NT (2003) The role of pectic enzymes in plant pathogenesis. Annu Rev Phytopathol 24:383–409
Coutinho PM, Stam M, Blanc E, Henrissat B (2003) Why are there so many carbohydrate-active enzyme-related genes in plants? Trends Plant Sci 8:563–565
Dorokhov YL, Frolova OY, Skurat EV, Ivanov PA, Gasanova TV, Sheveleva AA, Ravin NV, Mäkinen KM, Klimyuk VI, Skryabin KG, Gleba YY, Atabekov JG (2006) A novel function for a ubiquitous plant enzyme pectin methylesterase: the enhancer of RNA silencing. FEBS Lett 580:3872–3878
Eticha D, Stass A, Horst WJ (2005) Cell-wall pectin and its degree of methylation in the maize root-apex: significance for genotypic differences in aluminum resistance. Plant Cell Environ 28:1410–1420
Gao J, Wang Z, Xiong B, Shi D, Zhang T, Zeng H, Liao L, Cao S, Gu X, Li Q (2015) Correlations of albedo fracture toughness with cell wall substances and enzyme activities in kiyomi fruits. Food Sci 36:131–135 (In Chinese with English abstract)
Gasteiger E, Gattiker A, Hoogland C, Ivanyi I, Appel RD, Bairoch A (2003) ExPASy: the proteomics server for in-depth protein knowledge and analysis. Nucleic Acids Res 31:3784–3788
González-Carranza ZH, Whitelaw CA, Swarup R, Roberts JA (2002) Temporal and spatial expression of a polygalacturonase during leaf and flower abscission in oilseed rape and Arabidopsis. Plant Physiol 128:534–543
Gupta R, Brunak S (2002) Prediction of glycosylation across the human proteome and the correlation to protein function [C]. Kauai, awaii, SA: Pac Symp Biocomput 310–322
Hu B, Jin JP, Guo AY, Zhang H, Luo J, Gao G (2015) GSDS 2.0: an upgraded gene feature visualization server. Bioinformatics 31:1296–1297
Ivica L, Tobias D, Peer B (2012) SMART 7: recent updates to the protein domain annotation resource. Nucleic Acids Res 40:302–305
Jenkins J, Mayans O, Smith D, Worboys K, Pickersgill RW (2001) Three-dimensional structure of Erwinia chrysanthemi pectin methylesterase reveals a novel esterase active site. J Mol Biol 305:951–960
Jiang L, Wang Q, Liu-Zhong YE, Lin Y (2005) Pollen morphology of Artemisia L.and its systematic significance. Wuhan University J Natural Sci 10:448–454
Louvet R, Cavel E, Gutierrez L, Guenin S, Roger D, Gillet F, Guerineau F, Pelloux J (2006) Comprehensive expression profiling of the pectin methylesterase gene family during silique development in Arabidopsis thaliana. Planta 224:782–791
Marchler-Bauer A, Lu S, Anderson JB, Chitsaz F, Derbyshire MK, DeWeese-Scott C, Fong JH, Geer LY, Geer RC, Gonzales NR, Gwadz M, Hurwitz DI, Jackson JD, Ke Z, Lanczycki CJ, Lu F, Marchler GH, Mullokandov M, Omelchenko MV, Robertson CL, Song JS, Thanki N, Yamashita RA, Zhang D, Zhang N, Zheng C, Bryant SH (2011) CDD: a conserved domain database for the functional annotation of proteins. Nucleic Acids Res 39:D225–D229
Markovic O, Janecek S (2004) Pectin methylesterases: sequence-structural features and phylogenetic relationships. Carbohydr Res 339:2281–2295
Moustacas AM, Nari J, Borel M, Noat G, Ricard J (1991) Pectin methylesterase, metal ions, and plant cell-wall extension. The role of metal ions in plant cell-wall extension. Biochem J 279:351–354
Pelloux J, Rustérucci C, Mellerowicz EJ (2007) New insights into pectin methylesterase structure and function. Trends Plant Sci 12:267–277
Punta M, Coggill PC, Eberhardt RY, Mistry J, Tate J, Boursnell C, Pang N, Forslund K, Ceric G, Clements J, Heger A, Holm L, Sonnhammer ELL, Eddy SR, Bateman A, Finn RD (2012) The Pfam protein families database. Nucleic Acids Res 40:D290–D301
Rengel Z, Reid RJ (1997) Uptake of Al across the plasma membrane of plant cells. Plant Soil 192:31–35
Schmohl N, Pilling J, Fisahn J, Horst WJ (2000) Pectin methylesterase modulates aluminum sensitivity in Zea mays and Solanum tuberosum. Physiol Plant 109:419–427
Talor GJ (1988) The physiology of aluminum phytotoxicity. In: Sigel H, Sigel A (eds) Metal ions in biological systems, vol 24. Dekker, New York, pp 123–163
Thompson JD, Gibson TJ, Plewniak F, Jeanmougin F, Higgins DG (1997) The CLUSTAL_XWindows interface: flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucleic Acids Res 25:4876–4882
Tian GW, Chen MH, Zaltsman A, Citovsky V (2006) Pollen-specific pectin methylesterase involved in pollen tube growth. Dev Biol 294:83–91
Udall JA, Swanson JM, Haller K, Rapp RA, Sparks ME, Hatfield J, Yu Y, Wu Y, Dowd C, Arpat AB, Sickler BA, Wilkins TA, Guo JY, Chen XY, Scheffler J, Taliercio E, Turley R, McFadden H, Payton P, Klueva N, Allen R, Zhang D, Haigler C, Wilkerson C, Suo J, Schulze SR, Pierce ML, Essenberg M, Kim H, Llewellyn DJ, Dennis ES, Kudrna D, Wing R, Paterson AH, Soderlund C, Wendel JF (2006) A global assembly of cotton ESTs. Genome Res 16:441–450
Willats WG, Orfila C, Limberg G, Buchholt HC, van Alebeek GJ, Voragen AG, Marcus SE, Christensen TM, Mikkelsen JD, Murray BS, Knox JP (2001) Modulation of the degree and pattern of methyl-esterification of pectic homogalacturonan in plant cell walls. Implications for pectin methyl esterase action, matrix properties, and cell adhesion. J Biol Chem 276:19404–19413
Yang XY, Zeng ZH, Yan JY, Fan W, Bian HW, Zhu MY, Yang JL, Zheng SJ (2012) Association of specific pectin methylesterases with Al-induced root elongation inhibition in rice. Phyiol plant 148:502–511
Zhang L, Xue JA, Yu HQ, Li QY, Li RZ (2012) The expression and function study of pectin methylesterase genes which regulate and control the petal falling in Arabidopsis. Plant Physiol J 48:350–358 (In Chinese with English abstract)
Zhang T, Zhang S, Pei L, Yu T, Chen M, Li L, Zhou Y, Ma Y, Xu Q, Xu Z (2014) Genome-wide analysis of ANK gene family and expression pattern of the ANK25 gene in snap bean. J Plant Genetic Res 15:1334–1341 (In Chinese with English abstract)
Funding
This study was supported by grants from the National Natural Science Foundation of China (Grant Nos. 31660352, 31660350), Innovation Project of Guangxi Postgraduate Education (YCBZ2019015), Guangxi Innovation Team Project of Tubers of Modern Agricultural Industrial Technology System of China (nycytxgxcxtd-11-01), and the Science & Technology Development Fund of Guangxi Academy of Agricultural Sciences (Grant Nos. Guinongke2017JZ11).
Author information
Authors and Affiliations
Corresponding authors
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Li, Y., He, H. & He, LF. Genome-Wide Analysis of the Pectin Methylesterase Gene Family in Potato. Potato Res. 64, 1–19 (2021). https://doi.org/10.1007/s11540-020-09453-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11540-020-09453-1