Abstract
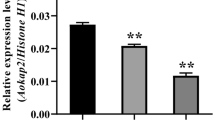
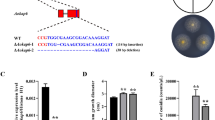
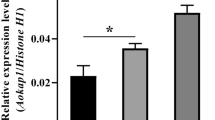
Glycerol dehydrogenase has been identified and characterized functionally in many species. However, little is known about glycerol dehydrogenase genes and their functions in Aspergillus oryzae. Here, a total of 45 glycerol dehydrogenase genes in Aspergillus oryzae were identified and renamed from AoGld1 to AoGld45 according to their chromosome distribution. They were classified into three groups based on phylogenetic analysis. Synteny analysis revealed that thirteen AoGld genes are conserved among Aspergillus species. Promoter analysis displayed that AoGld3 and AoGld13 harbored multiple binding elements of GATA-type transcription factors and zinc-finger protein msnA that were involved in nitrogen and kojic acid metabolism, respectively. Moreover, the AoGld3 deletion strain Δgld3 was generated by the CRISPR/Cas9 system, which had no visible growth defects compared with the control wild-type strain under the control and osmotic stress treatments. However, disruption of AoGld3 led to the inhibition of kojic acid production, and the expression of kojA, kojR was down-regulated in the Δgld3 strain. Furthermore, when kojA or kojR was overexpressed in the Δgld3 strain, the yield of kojic acid was restored, suggesting that AoGld3 is involved in kojic acid production through affecting the expression of kojR and kojA. Taken together, these findings provide new insights into our understanding of glycerol dehydrogenase and establish foundation for further study of their roles in Aspergillus oryzae.







Similar content being viewed by others
References
Aguilera J, Prieto JA (2001) The Saccharomyces cerevisiae aldose reductase is implied in the metabolism of methylglyoxal in response to stress conditions. Curr Genet 39:273–283. https://doi.org/10.1007/s002940100213
Amako K et al (2006) NADP+-dependent D-arabinose dehydrogenase shows a limited contribution to erythroascorbic acid biosynthesis and oxidative stress resistance in Saccharomyces cerevisiae. Biosci Biotechnol Biochem 70:3004–3012. https://doi.org/10.1271/bbb.60399
Arakawa GY et al (2019) A unique Zn(II)2-Cys6-type protein, KpeA, is involved in secondary metabolism and conidiation in Aspergillus oryzae. Fungal Genet Biol 127:35–44. https://doi.org/10.1016/j.fgb.2019.02.004
Bentley R (2006) From miso, sake and shoyu to cosmetics: a century of science for kojic acid. Nat Prod Rep 23:1046–1062. https://doi.org/10.1039/b603758p
Chang Q, Griest TA, Harter TM, Mark Petrash J (2007) Functional studies of aldo-keto reductases in Saccharomyces cerevisiae. BBA Mol Cell Res 1773:321–329. https://doi.org/10.1016/j.bbamcr.2006.10.009
Chang PK et al (2011) Loss of msnA, a putative stress regulatory gene, in Aspergillus parasiticus and Aspergillus flavus increased production of conidia, aflatoxins and kojic acid. Toxins 3:82–104. https://doi.org/10.3390/toxins3010082
Christensen T, Hynes MJ, Davis MA (1998) Role of the regulatory gene areA of Aspergillus oryzae in nitrogen metabolism. Appl Environ Micro 64:3232–3237
Cordier H, Mendes F, Vasconcelos I, Francois JM (2007) A metabolic and genomic study of engineered Saccharomyces cerevisiae strains for high glycerol production. Metab Eng 9:364–378. https://doi.org/10.1016/j.ymben.2007.03.002
de Vries RP et al (2003) Glycerol dehydrogenase, encoded by gldB is essential for osmotolerance in Aspergillus nidulans. Mol microbiol 49:131–141. https://doi.org/10.1046/j.1365-2958.2003.03554.x
Feng W, Liang J, Wang B, Chen J (2019) Improvement of kojic acid production in Aspergillus oryzae AR-47 mutant strain by combined mutagenesis. Bioprocess Biosyst Eng 42:753–761. https://doi.org/10.1007/s00449-019-02079-9
Ford G, Ellis EM (2002) Characterization of Ypr1p from Saccharomyces cerevisiae as a 2-methylbutyraldehyde reductase. Yeast 19:1087–1096. https://doi.org/10.1002/yea.899
Garay-Arroyo A, Covarrubias AA (1999) Three genes whose expression is induced by stress in Saccharomyces cerevisiae. Yeast 15:879–892
Izawa S, Sato M, Yokoigawa K, Inoue Y (2004) Intracellular glycerol influences resistance to freeze stress in Saccharomyces cerevisiae: analysis of a quadruple mutant in glycerol dehydrogenase genes and glycerol-enriched cells. Appl Microbiol Biot 66:108–114. https://doi.org/10.1007/s00253-004-1624-4
Janevska S, Tudzynski B (2018) Secondary metabolism in Fusarium fujikuroi: strategies to unravel the function of biosynthetic pathways. Appl Microbiol Biot 102:615–630. https://doi.org/10.1007/s00253-017-8679-5
Jiang W, Wang S, Wang Y, Fang B (2016) Key enzymes catalyzing glycerol to 1,3-propanediol. Biotechnol Biofuels 9:57. https://doi.org/10.1186/s13068-016-0473-6
Katayama T, Tanaka Y, Okabe T, Nakamura H, Fujii W, Kitamoto K, Maruyama J (2016) Development of a genome editing technique using the CRISPR/Cas9 system in the industrial filamentous fungus Aspergillus oryzae. Biotechnol Lett 38:637–642. https://doi.org/10.1007/s10529-015-2015-x
Kim ST, Huh WK, Lee BH, Kang SO (1998) D-arabinose dehydrogenase and its gene from Saccharomyces cerevisiae. BBA Protein Struct Mol Enzymol 1429:29–39. https://doi.org/10.1016/s0167-4838(98)00217-9
Klein M, Swinnen S, Thevelein JM, Nevoigt E (2017) Glycerol metabolism and transport in yeast and fungi: established knowledge and ambiguities. Environ Microbiol 19:878–893. https://doi.org/10.1111/1462-2920.13617
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2(T)(-Delta Delta C) method. Methods 25:402–408. https://doi.org/10.1006/meth.2001.1262
Marui J et al (2011) Kojic acid biosynthesis in Aspergillus oryzae is regulated by a Zn(II)2Cys6 transcriptional activator and induced by kojic acid at the transcriptional level. J Biosci Bioeng 112:40–43. https://doi.org/10.1016/j.jbiosc.2011.03.010
Maruyama J, Kitamoto K (2011) Targeted gene disruption in Koji mold Aspergillus oryzae. Methods Mol Biol 765:447–456. https://doi.org/10.1007/978-1-61779-197-0_27
Masuda CA, Previato JO, Miranda MN, Assis LJ, Penha LL, Mendonca-Previato L, Montero-Lomeli M (2008) Overexpression of the aldose reductase GRE3 suppresses lithium-induced galactose toxicity in Saccharomyces cerevisiae. FEMS Yeast Res 8:1245–1253. https://doi.org/10.1111/j.1567-1364.2008.00440.x
Matsuzawa T, Ohashi T, Hosomi A, Tanaka N, Tohda H, Takegawa K (2010) The gld1+ gene encoding glycerol dehydrogenase is required for glycerol metabolism in Schizosaccharomyces pombe. Appl Microbiol Biot 87:715–727. https://doi.org/10.1007/s00253-010-2586-3
Mohamad R et al (2010) Kojic acid: applications and development of fermentation process for production. Biotechnol Mol Biol Rev 5:24–37
Nevoigt E, Stahl U (1997) Osmoregulation and glycerol metabolism in the yeast Saccharomyces cerevisiae. FEMS Microbiol Rev 21:231–241. https://doi.org/10.1111/j.1574-6976.1997.tb00352.x
Norbeck J, Blomberg A (1997) Metabolic and regulatory changes associated with growth of Saccharomyces cerevisiae in 1.4 M NaCl. Evidence for osmotic induction of glycerol dissimilation via the dihydroxyacetone pathway. J Biol Chem 272:5544–5554. https://doi.org/10.1074/jbc.272.9.5544
Oda K, Kobayashi A, Ohashi S, Sano M (2014) Aspergillus oryzae laeA regulates kojic acid synthesis genes. Biosci Biotechnol Biochem 75:1832–1834. https://doi.org/10.1271/bbb.110235
Oechsner U, Magdolen V, Bandlow W (1988) A nuclear yeast gene (GCY) encodes a polypeptide with high homology to a vertebrate eye lens protein. FEBS Lett 238:123–128. https://doi.org/10.1016/0014-5793(88)80240-0
Petrash JM, Murthy BS, Young M, Morris K, Rikimaru L, Griest TA, Harter T (2001) Functional genomic studies of aldo-keto reductases. Chem Biol Interact 130–132:673–683. https://doi.org/10.1016/s0009-2797(00)00258-1
Redkar RJ, Locy RD, Singh NK (1995) Biosynthetic pathways of glycerol accumulation under salt stress in Aspergillus nidulans. Exp Mycol 19:241–246. https://doi.org/10.1006/emyc.1995.1030
Robinson CH (2001) Cold adaptation in Arctic and Antarctic fungi. New Phytol 151:341–353. https://doi.org/10.1046/j.1469-8137.2001.00177.x
Ruijter GJ, Visser J, Rinzema A (2004) Polyol accumulation by Aspergillus oryzae at low water activity in solid-state fermentation. Microbiology 150:1095–1101. https://doi.org/10.1099/mic.0.26723-0
Ruzheinikov SN et al (2001) Glycerol dehydrogenase. structure, specificity, and mechanism of a family III polyol dehydrogenase. Structure 9:789–802. https://doi.org/10.1016/s0969-2126(01)00645-1
Sano M (2016) Aspergillus oryzae nrtA affects kojic acid production. Biosci Biotechnol Biochem 80:1776–1780. https://doi.org/10.1080/09168451.2016.1176517
Schuurink R, Busink R, Hondmann DH, Witteveen CF, Visser J (1990) Purification and properties of NADP(+)-dependent glycerol dehydrogenases from Aspergillus nidulans and A. niger. J Gen Microbiol 136:1043–1050. https://doi.org/10.1099/00221287-136-6-1043
Seidl V, Seiboth B, Karaffa L, Kubicek CP (2004) The fungal STRE-element-binding protein Seb1 is involved but not essential for glycerol dehydrogenase (gld1) gene expression and glycerol accumulation in Trichoderma atroviride during osmotic stress. Fungal Genet Biol 41:1132–1140. https://doi.org/10.1016/j.fgb.2004.09.002
Terabayashi Y et al (2010) Identification and characterization of genes responsible for biosynthesis of kojic acid, an industrially important compound from Aspergillus oryzae. Fungal Genet Biol 47:953–961. https://doi.org/10.1016/j.fgb.2010.08.014
Traff KL, Otero Cordero RR, van Zyl WH, Hahn-Hagerdal B (2001) Deletion of the GRE3 aldose reductase gene and its influence on xylose metabolism in recombinant strains of Saccharomyces cerevisiae expressing the xylA and XKS1 genes. Appl Environ Microbiol 67:5668–5674. https://doi.org/10.1128/AEM.67.12.5668-5674.2001
Tudzynski B (2014) Nitrogen regulation of fungal secondary metabolism in fungi. Front Microbiol. https://doi.org/10.3389/fmicb.2014.00656
Williams TJ, Allen M, Tschitschko B, Cavicchioli R (2017) Glycerol metabolism of haloarchaea. Environ Microbiol 19:864–877. https://doi.org/10.1111/1462-2920.13580
Wong KH, Hynes MJ, Davis MA (2008) Recent advances in nitrogen regulation: a comparison between Saccharomyces cerevisiae and filamentous fungi. Eukaryot Cell 7:917–925. https://doi.org/10.1128/ec.00076-08
Zhang Z et al (2019) Identification and characterization of the ZRT, IRT-like protein (ZIP) family genes reveal their involvement in growth and kojic acid production in Aspergillus oryzae. J Ind Microbiol Biotechnol 46:1769–1780. https://doi.org/10.1007/s10295-019-02236-9
Acknowledgements
We thank Dr. Van-Tuan Tran for kindly providing the pEX1 vector. This work was supported by Science and Technology Research Project of Jiangxi Provincial Department of Education (GJJ190611, GJJ180630) and Doctoral Scientific Research Foundation of Jiangxi Science and Technology Normal University (2018BSQD030).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical approval
This article does not contain any studies with human participants or animals experiments.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Fan, J., Zhang, Z., Long, C. et al. Identification and functional characterization of glycerol dehydrogenase reveal the role in kojic acid synthesis in Aspergillus oryzae. World J Microbiol Biotechnol 36, 136 (2020). https://doi.org/10.1007/s11274-020-02912-4
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11274-020-02912-4