Abstract
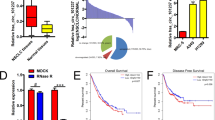
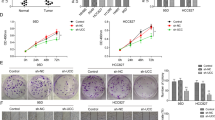
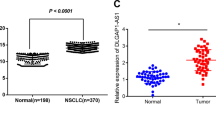
Circular RNAs (circRNAs) are involved in the regulation of many pathophysiological processes as non-coding RNAs. This study focuses on the role of circRACGAP1 in the development of non-small cell lung cancer (NSCLC). Expression patterns of circRACGAP1 and miR-144-5p in NSCLC tissues and cell lines were quantified by qRT-PCR analysis. Then, the function of circRACGAP1 on cell proliferation and tumorigenesis were confirmed in vitro and in vivo using CCK-8 assay, colony formation, EdU incorporation, and xenograft technique. The regulation of circRACGAP1 on Gefitinib resistance of NSCLC cells was evaluated by flow cytometry. The regulatory network of circRACGAP1/miR-144-5p/CDKL1 was verified by luciferase reporter assay and RNA pull-down. Western blotting analysis was performed to assess the biomarkers of cell cycle and apoptosis-associated proteins. CircRACGAP1 was highly expressed and miR-144-5p was inhibited both in NSCLC tissues and cell lines, suggesting their negative correlation in NSCLC. Knockdown of circRACGAP1 suppressed cell proliferation via arresting the cell cycle. miR-144-5p was identified as a downstream target to reverse circRACGAP1-mediated cell proliferation. miR-144-5p directly targeted the 3′-UTR of CDKL1 to regulate cell cycle of NSCLC cells. circRACGAP1 knockdown dramatically inhibited the tumor growth and enhanced the sensitivity of NSCLC to Gefitinib in vitro and in vivo. In summary, our study revealed a novel machinery of circRACGAP1/miR-144-5p/CDKL1 for the NSCLC tumorigenesis and development, providing potential diagnostic and therapeutic targets for NSCLC.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
References
Ferlay J, Soerjomataram I, Dikshit R, Eser S, Mathers C, Rebelo M, et al. Cancer incidence and mortality worldwide: sources, methods and major patterns in GLOBOCAN 2012. Int J Cancer. 2015;136:E359–86.
Wang L, Tong X, Zhou Z, Wang S, Lei Z, Zhang T, et al. Circular RNA hsa_circ_0008305 (circPTK2) inhibits TGF-beta-induced epithelial-mesenchymal transition and metastasis by controlling TIF1gamma in non-small cell lung cancer. Mol Cancer. 2018;17:140.
Zang H, Wang W, Fan S. The role of microRNAs in resistance to targeted treatments of non-small cell lung cancer. Cancer Chemother Pharm. 2017;79:227–31.
Feng J, Zhang X, Zhu H, Wang X, Ni S, Huang J. High expression of FoxP1 is associated with improved survival in patients with non-small cell lung cancer. Am J Clin Pathol. 2012;138:230–5.
DeSantis CE, Lin CC, Mariotto AB, Siegel RL, Stein KD, Kramer JL, et al. Cancer treatment and survivorship statistics, 2014. CA Cancer J Clin. 2014;64:252–71.
Tian J, Luo Y, Xiang J, Tang J. Combined treatment for non-small cell lung cancer and breast cancer patients with brain metastases with whole brain radiotherapy and temozolomide: a systematic review and meta-analysis. J Neurooncol. 2017;135:217–27.
Zhang M, Xin Y. Circular RNAs: a new frontier for cancer diagnosis and therapy. J Hematol Oncol. 2018;11:21.
Memczak S, Jens M, Elefsinioti A, Torti F, Krueger J, Rybak A, et al. Circular RNAs are a large class of animal RNAs with regulatory potency. Nature. 2013;495:333–8.
Jeck WR, Sorrentino JA, Wang K, Slevin MK, Burd CE, Liu J, et al. Circular RNAs are abundant, conserved, and associated with ALU repeats. RNA. 2013;19:141–57.
Rybak-Wolf A, Stottmeister C, Glazar P, Jens M, Pino N, Giusti S, et al. Circular RNAs in the mammalian brain are highly abundant, conserved, and dynamically expressed. Mol Cell. 2015;58:870–85.
Piwecka M, Glazar P, Hernandez-Miranda LR, Memczak S, Wolf SA, Rybak-Wolf A, et al. Loss of a mammalian circular RNA locus causes miRNA deregulation and affects brain function. Science. 2017;357. https://doi.org/10.1126/science.aam8526.
Li Z, Huang C, Bao C, Chen L, Lin M, Wang X, et al. Exon-intron circular RNAs regulate transcription in the nucleus. Nat Struct Mol Biol. 2015;22:256–64.
Li Y, Hu J, Li L, Cai S, Zhang H, Zhu X, et al. Upregulated circular RNA circ_0016760 indicates unfavorable prognosis in NSCLC and promotes cell progression through miR-1287/GAGE1 axis. Biochem Biophys Res Commun. 2018;503:2089–94.
Danhua Qu BY, Rui Xin, Tianggang MA. A novel circular RNA hsa_circ_0020123 exerts oncogenic properties through suppression of miR-144 in non-small cell lung cancer. Am J Cancer Res. 2018;8:1387–402.
Zou Q, Wang T, Li B, Li G, Zhang L, Wang B, et al. Overexpression of circ-0067934 is associated with increased cellular proliferation and the prognosis of non-small cell lung cancer. Oncol Lett. 2018;16:5551–6.
Yao JT, Zhao SH, Liu QP, Lv MQ, Zhou DX, Liao ZJ, et al. Over-expression of CircRNA_100876 in non-small cell lung cancer and its prognostic value. Pathol Res Pr. 2017;213:453–6.
Zhang S, Zeng X, Ding T, Guo L, Li Y, Ou S, et al. Microarray profile of circular RNAs identifies hsa_circ_0014130 as a new circular RNA biomarker in non-small cell lung cancer. Sci Rep. 2018;8:2878.
Liang Y, Liu M, Wang P, Ding X, Cao Y. Analysis of 20 genes at chromosome band 12q13: RACGAP1 and MCRS1 overexpression in nonsmall-cell lung cancer. Genes Chromosomes Cancer. 2013;52:305–15.
Tang L, Gao Y, Yan F, Tang J. Evaluation of cyclin-dependent kinase-like 1 expression in breast cancer tissues and its regulation in cancer cell growth. Cancer Biother Radiopharm. 2012;27:392–8.
Siegel RL, Miller KD, Jemal A. Cancer Statistics, 2017. CA Cancer J Clin. 2017;67:7–30.
Chen W, Zheng R, Baade PD, Zhang S, Zeng H, Bray F, et al. Cancer statistics in China, 2015. CA Cancer J Clin. 2016;66:115–32.
Rimkunas VM, Crosby KE, Li D, Hu Y, Kelly ME, Gu TL, et al. Analysis of receptor tyrosine kinase ROS1-positive tumors in non-small cell lung cancer: identification of a FIG-ROS1 fusion. Clin Cancer Res. 2012;18:4449–57.
Byers LA, Rudin CM. Small cell lung cancer: where do we go from here? Cancer. 2015;121:664–72.
Hu W, Bi ZY, Chen ZL, Liu C, Li LL, Zhang F, et al. Emerging landscape of circular RNAs in lung cancer. Cancer Lett. 2018;427:18–27.
Jiang MM, Mai ZT, Wan SZ, Chi YM, Zhang X, Sun BH, et al. Microarray profiles reveal that circular RNA hsa_circ_0007385 functions as an oncogene in non-small cell lung cancer tumorigenesis. J Cancer Res Clin Oncol. 2018;144:667–74.
Liu W, Xiao P, Wu H, Wang L, Kong D, Yu F. MicroRNA-98 plays a suppressive role in non-small cell lung cancer through inhibition of SALL4 protein expression. Oncol Res. 2017;25:975–88.
Wang R, Yan B, Li Z, Jiang Y, Mao C, Wang X, et al. Long non-coding RNA HOX transcript antisense RNA promotes expression of 14-3-3sigma in non-small cell lung cancer. Exp Ther Med. 2017;14:4503–8.
Dai X, Zhang N, Cheng Y, Yang T, Chen Y, Liu Z, et al. RNA-binding protein trinucleotide repeat-containing 6A regulates the formation of circular RNA 0006916, with important functions in lung cancer cells. Carcinogenesis. 2018;39:981–92.
Zhu X, Wang X, Wei S, Chen Y, Chen Y, Fan X, et al. hsa_circ_0013958: a circular RNA and potential novel biomarker for lung adenocarcinoma. FEBS J. 2017;284:2170–82.
Zhong Y, Du Y, Yang X, Mo Y, Fan C, Xiong F, et al. Circular RNAs function as ceRNAs to regulate and control human cancer progression. Mol Cancer. 2018;17:79.
Kim HH, Kuwano Y, Srikantan S, Lee EK, Martindale JL, Gorospe M. HuR recruits let-7/RISC to repress c-Myc expression. Genes Dev. 2009;23:1743–8.
Begum S, Yiu A, Stebbing J, Castellano L. Novel tumour suppressive protein encoded by circular RNA, circ-SHPRH, in glioblastomas. Oncogene. 2018;37:4055–7.
Meng J, Chen S, Han JX, Qian B, Wang XR, Zhong WL, et al. Twist1 regulates vimentin through Cul2 circular RNA to promote EMT in hepatocellular carcinoma. Cancer Res. 2018;78:4150–62.
Wu J, Jiang Z, Chen C, Hu Q, Fu Z, Chen J, et al. CircIRAK3 sponges miR-3607 to facilitate breast cancer metastasis. Cancer Lett. 2018;430:179–92.
Ma Y, Zhang X, Wang YZ, Tian H, Xu S. Research progress of circular RNAs in lung cancer. Cancer Biol Ther. 2019;20:123–9.
Wan L, Zhang L, Fan K, Cheng ZX, Sun QC, Wang JJ. Circular RNA-ITCH suppresses lung cancer proliferation via inhibiting the wnt/beta-catenin pathway. Biomed Res Int. 2016;2016:1579490.
Luo YH, Zhu XZ, Huang KW, Zhang Q, Fan YX, Yan PW, et al. Emerging roles of circular RNA hsa_circ_0000064 in the proliferation and metastasis of lung cancer. Biomed Pharmacother. 2017;96:892–8.
Matsushita R, Seki N, Chiyomaru T, Inoguchi S, Ishihara T, Goto Y, et al. Tumour-suppressive microRNA-144-5p directly targets CCNE1/2 as potential prognostic markers in bladder cancer. Br J Cancer. 2015;113:282–9.
Yamada Y, Arai T, Kojima S, Sugawara S, Kato M, Okato A, et al. Regulation of antitumor miR-144-5p targets oncogenes: Direct regulation of syndecan-3 and its clinical significance. Cancer Sci. 2018;109:2919–36.
Song L, Peng L, Hua S, Li X, Ma L, Jie J, et al. miR-144-5p enhances the radiosensitivity of non-small-cell lung cancer cells via targeting ATF2. Biomed Res Int. 2018;2018:5109497.
Yang XM, Cao XY, He P, Li J, Feng MX, Zhang YL, et al. Overexpression of Rac GTPase activating protein 1 contributes to proliferation of cancer cells by reducing hippo signaling to promote cytokinesis. Gastroenterology. 2018;155:1233–49. e1222
Pei J, Xiao W, Zhu D, Ji X, Shi L, Deng X. LncRNA DSCAM-AS1 promotes proliferation, migration and invasion of colorectal cancer cells via modulating miR-144-5p/CDKL1. Life Sci. 2019:117050.
Shang J, Chen WM, Wang ZH, Wei TN, Chen ZZ, Wu WB. CircPAN3 mediates drug resistance in acute myeloid leukemia through the miR-153-5p/miR-183-5p-XIAP axis. Exp Hematol. 2019;70:42–54. e3.
Liu YT, Han XH, Xing PY, Hu XS, Hao XZ, Wang Y, et al. Circular RNA profiling identified as a biomarker for predicting the efficacy of Gefitinib therapy for non-small cell lung cancer. J Thorac Dis. 2019;11:1779–87.
Chandran RK, Geetha N, Sakthivel KM, Aswathy CG, Gopinath P, Raj TVA, et al. Genomic amplification of BCR-ABL1 fusion gene and its impact on the disease progression mechanism in patients with chronic myelogenous leukemia. Gene. 2019;686:85–91.
Author information
Authors and Affiliations
Contributions
Guarantor of integrity of the entire study: PL; Study concepts: C-HH, PL; Study design: PL, C-HH; Definition of intellectual content: ML, HX; Literature research: ML, HX, FW, H-XZ; Clinical studies: ML, HX, Z-KX; Experimental studies: ML, HX, BL, Z-KX; Data acquisition: ML, HX, BL, Z-KX; Data analysis: ML, HX, BL, Z-KX; Statistical analysis: FW, H-XZ; Manuscript preparation: HX, FW, Z-KX; Manuscript editing: ML, PL, H-XZ; Manuscript review: PL.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical approval
This study was approved by the Ethics Committee of the Second Xiangya Hospital of Central South University.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
About this article
Cite this article
Lu, M., Xiong, H., Xia, ZK. et al. circRACGAP1 promotes non-small cell lung cancer proliferation by regulating miR-144-5p/CDKL1 signaling pathway. Cancer Gene Ther 28, 197–211 (2021). https://doi.org/10.1038/s41417-020-00209-0
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41417-020-00209-0
This article is cited by
-
A novel circ_MACF1/miR-942-5p/TGFBR2 axis regulates the functional behaviors and drug sensitivity in gefitinib-resistant non-small cell lung cancer cells
BMC Pulmonary Medicine (2022)
-
The emerging roles of circular RNA-mediated autophagy in tumorigenesis and cancer progression
Cell Death Discovery (2022)