Abstract
SUMO E3 ligases specify protein substrates for SUMOylation. The SUMO E3 ligases PIAS1 and TIF1γ target the transcriptional regulator SnoN for SUMOylation leading to suppression of epithelial–mesenchymal transition (EMT). Whether and how TIF1γ and PIAS1 might coordinate SnoN SUMOylation and regulation of EMT remained unknown. Here, we reveal that SnoN associates simultaneously with both TIF1γ and PIAS1, leading to a trimeric protein complex. Hence, PIAS1 and TIF1γ collaborate to promote the SUMOylation of SnoN. Importantly, loss of function studies of PIAS1 and TIF1γ suggest that these E3 ligases act in an interdependent manner to suppress EMT of breast cell-derived tissue organoids. Collectively, our findings unveil a novel mechanism by which SUMO E3 ligases coordinate substrate SUMOylation with biological implications.
Similar content being viewed by others
Introduction
Conjugation of proteins with the small ubiquitin like modifier (SUMO) protein bears important consequences for protein substrates, regulating their biochemical, biological, and pathological properties [1, 2]. SUMOylation is a multistep process involving the SUMO E1 activating enzyme, the SUMO E2 conjugating enzyme Ubc9, and a SUMO E3 ligase [3, 4]. SUMO E3 ligases play a key role in specifying protein substrates for SUMOylation [5], but the mechanisms underlying specifying SUMOylation remain incompletely understood.
The protein inhibitors for activated stats (signal transducer and activator of transcription) (PIASs) have been the subject of intense investigation [6]. Members of the PIAS SUMO E3 ligases, PIAS1-4, specify a diverse set of proteins for SUMOylation [6, 7]. Interestingly, some of the SUMO substrates regulated by the PIAS family of SUMO E3 ligases have been demonstrated to be also targeted by other SUMO E3 ligases [4, 8,9,10]. However, it is not clear how the function of distinct SUMO E3 ligases targeting common protein substrates for SUMOylation is coordinated.
The transcriptional regulator SnoN exerts diverse biological responses in various cell types including proliferating and postmitotic cells [4, 11,12,13,14,15]. SnoN function is regulated by SUMOylation [4, 8, 13]. The SUMO E3 ligase PIAS1 associates with and promotes the SUMOylation of SnoN [4]. Interestingly, recent studies have revealed that TIF1γ interacts with and promotes SUMOylation of the SnoN1 isoform [8, 16]. SUMOylation mediates the ability of SnoN to suppress epithelial–mesenchymal transition (EMT), a fundamental cellular process in development that is deregulated in disease states including cancer [4, 8, 13, 16, 17]. That PIAS1 and TIF1γ modify SnoN to suppress EMT raises the key question of whether PIAS1 and TIF1γ act independently or collaboratively in promoting SUMOylation of SnoN and regulating EMT.
In this study, we have uncovered that SnoN triggers the assembly of a PIAS1–SnoN-TIF1γ- multiprotein complex that promotes SnoN SUMOylation. Accordingly, PIAS1 and TIF1γ require each other to suppress EMT in three-dimensional organoids of non-transformed and transformed mammary epithelial cells.
Material and methods
Plasmids
CMV-based plasmids expressing FLAG- or HA-tagged wild-type TIF1γ, FLAG-TIF1γ CS, in which Cysteines 125 and 128 within the ring domain are converted to two serine residues, FLAG-TIF1γΔC, in which the C terminal starting from amino acid 889 is deleted, FLAG-PIAS1, FLAG-PIAS1CS, in which Cysteine 350 is converted to a serine residue, FLAG-SnoN, HA-SnoN, MYC-SnoN, FLAG-SnoNres, green fluorescence protein (GFP) in fusion with the TIF1γ interaction peptide motif (TIPtide) in SnoN1 corresponding to amino acids 452–467, are described [8, 9, 13, 15, 17, 18]. Mouse U6 RNA polymerase III promoter-based RNA interference-CMV-GFP plasmids, co-expressing enhanced green fluorescent protein (EGFP), to express short hairpin (sh) RNAs targeting TIF1γ, PIAS1, and SnoN are described [4, 8, 13, 18]. Restriction endonuclease-digested SnoN2, SnoNKdR, and PIAS1 were used as inserts to ligate with Restriction endonuclease-digested Renilla Luciferase (RLuc) tagged SnoN (RLuc-SnoN) vector [19], to generate plasmids encoding the proteins RLuc-SnoN2, RLuc-SnoNKdR, and RLuc-PIAS1, respectively. A pCAGip vector encoding a puromycin resistance marker alone or together with SnoNKdR, SUMO–SnoN, PIAS1, or PIAS1CS as a bicistronic transcript containing an internal ribosomal entry site was used to stably express the resistance marker alone (vector control) or together with the specific protein of interest, respectively, in cells [4, 18].
Cell lines and transfections
The human embryonic kidney 293T epithelial cell, human MDA-MB-231 breast cancer cells, and NAMRU murine mammary gland (NMuMG) epithelial cells were cultured in Dulbecco’s modified Eagle’s medium with high glucose and L-glutamine (Invitrogen, Canada) supplemented with 10% fetal bovine serum (FBS, Invitrogen), and 10 mg/ml recombinant human insulin (Invitrogen, Canada) in the case of the NMuMG cells. The human MCF7 breast cancer cells were cultured in minimum essential medium (Invitrogen, Canada) supplemented with 1× nonessential amino acids (Invitrogen, Canada), 1× sodium pyruvate (Invitrogen, Canada), 10% FBS, and 10 mg/ml recombinant human insulin. The 293T, MDA-MB231, NMuMG, and MCF7 cells were from the American Tissue Cell Collection. Cells were maintained in a humidified incubator at 37 °C in 95% air and 5% CO2 and all the cells were confirmed to be free of pathogenic mycoplasma strains by a PCR-ELISA kit. 293T were transfected by the calcium-phosphate precipitation method [20]. The MDA-MB-231, NMuMG, and MCF7 cells were transfected using Lipofectamine 3000 reagent (Invitrogen, Canada) as per the manufacturer’s instructions.
Co-immunoprecipitation analyses
Thirty percent confluent cells were transfected with expression plasmids as indicated. Two days post transfection, cells were lysed in TNTE (50 mM Tris, 150 mM NaCl, and 1 mM EDTA) buffer-containing 0.5% Triton X-100, protease inhibitors, and phosphatase inhibitors [8, 17, 21]. Cell extracts were centrifuged at 14,000 × g for 10 min at 4 °C and the supernatants were immunoprecipitated with mouse anti-FLAG (Sigma, Canada) or anti-HA (Biolegend, USA) antibodies. The input and immunoprecipitated protein complexes were resolved using SDS-PAGE followed by transfer to nitrocellulose membranes. The membranes were incubated with mouse anti-FLAG, mouse anti-HA, mouse anti-GFP (Santa Cruz, Canada), rabbit anti-PIAS1 (Abcam, USA), rabbit anti-SnoN (Proteintech, USA), mouse anti-TIF1γ (Santa Cruz, Canada) or mouse anti-actin (Santa Cruz, Canada), as the primary antibody, and HRP-conjugated goat anti-mouse or donkey anti-rabbit IgG (Jackson Laboratories) as secondary antibodies. Immunoprecipitates and total protein lysates containing RLuc alone or in fusion with SnoN or PIAS1 were subjected to RLuc assays using the RLuc kit (Promega, USA) and the Orion II luminometer (Berthold Detection Systems, Germany) detection system. Prior to RLuc analysis, immunoprecipitates were resuspended in TNE buffer-containing 0.1% Triton X-100 [19]. In the sequential immunoprecipitation assay, TIF1γ was immunoprecipitated using anti-FLAG antibody followed by elution of the FLAG immuno-complex using FLAG elution peptide (Millipore Sigma, Canada). The eluate was further subjected to anti-HA immunoprecipitation [22].
In vivo SUMOylation assays
Thirty percent confluent cells were transfected with a control vector, or a plasmid encoding MYC-SnoN, HA-hSUMO1 (referred to as SUMO hereafter), alone or together, without or with plasmids to express or knockdown TIF1γ or PIAS1 as shown. Two days post transfection, cells were lysed in TNTE (50 mM Tris, 150 mM NaCl, and 1 mM EDTA) buffer-containing 0.5% Triton X-100, 0.1% SDS, protease inhibitors, and phosphatase inhibitors, alone or together with 20 mM of n-ethylmaleimide (NEM) (Calbiochem, Canada), an isopeptidase inhibitor to suppress the activity of the SENP deSUMOylases upon lysis to help preserve the abundance of SUMOylated species of SnoN [4, 8, 17, 21]. The cell extracts were centrifuged at 14,000 × g for 10 min at 4 °C and the supernatant was immunoprecipitated using mouse anti-MYC antibody (Santa Cruz, Canada) at 4 °C, after saving 10% of the supernatant for Bradford protein quantification and protein expression analysis (Bio-Rad Laboratories, Canada). The immunoprecipitated sample and input lysates were immunoblotted with mouse anti-HA (Biolegend, Canada), rabbit anti-PIAS1 (Abcam, USA), rabbit anti-SnoN (Proteintech, USA), mouse anti-TIF1γ (Santa Cruz, Canada) or mouse anti-actin (Santa Cruz, Canada), as the primary antibody, and HRP-conjugated goat anti-mouse or donkey anti-rabbit IgG (Jackson Laboratories) as secondary antibodies, followed by Enhanced Chemiluminescence (ECL) (Millipore) reagent incubation and light signal detection using a VersaDoc 5000 Imager (Bio-Rad Laboratories). Densitometry was performed using Quantity One software (Bio-Rad Laboratories).
Three-dimensional cultures
Three-dimensional cultures of cells were prepared in 8-well chamber slides (Millicell EZ Slide, Millipore) or 96-well flat-bottom, ultra-low attachment plates (BD Biosciences, ON, Canada). 8-well chamber slide wells or each well of the 96-well dish were precoated with 75 or 50 μl, respectively, of 3 mg/ml final concentration Matrigel (BD Biosciences) cushions in antibiotic–antimycotic-containing growth medium. The Matrigel bed was allowed to solidify by incubating at 37 °C in 5% CO2 containing tissue culture incubator. 700 or 400 cells per well of 8-well chamber or 96-well dish respectively, were resuspended in 75 or 50 μl, respectively, of 5 mg/ml Matrigel and layered on top of the Matrigel bed and allowed to solidify by incubating at 37 °C in 5% CO2 containing tissue culture incubator. Complete medium was layered on top after 1 h. The next day and every third day the three-dimensional cell-derived organoids received 100pM TGF, 10 μM KI, alone or together until 8 days in culture. Differential interference contrast images of six representative organoids from each well were captured at 30× objective (Olympus IX70) following an overall estimate of the total number (~50) and assessment of growth phenotypes (acini vs. filled for NMuMG and MCF7 organoids and non-deformed vs. deformed for MDA-MB-231 organoids).
Indirect immunofluorescence
The 3D-organoids were fixed with 4% formaldehyde, permeabilized using 0.5% ice-cold Triton X-100 solution for time, and blocked using 10% BSA in phosphate-buffered saline at 37 °C for 1 h. The organoids were then subjected to indirect immunofluorescence staining using a rabbit E-cadherin antibody (Cell Signaling Technology, Canada) as the primary antibody and goat anti-rabbit antibody conjugated to Cy5 dye (Jackson Lab, Burlington, ON, Canada) as the secondary antibody along with the DNA fluorescent dye Hoechst 33342 (Invitrogen) to visualize cells’ nuclei. Fluorescence images of the multicellular colonies were captured using a fluorescent microscope with a 40× objective lens (Olympus Fluoview FV1000 microscope, Canada). For each biological replicate, the exposure time for each E-cadherin and Hoechst-specific fluorescence signals was kept constant for all the different conditions tested. After an overall observation of each well, images of 2–3 colonies per experimental condition were captured.
Statistical analyses
Replicates are used to account for inter-experimental variations in measurements of a particular parameter, which may otherwise lead to erroneous hypothesis testing and parameter estimation. Biological replicates refer to the use of biologically distinct samples or cell culture populations in a specific type of experiment, and is used to alleviate random biological variations which may be a source of noise or be part of the subject of study [23]. A minimum of three biological replicates were used in each type of experiment in this study to facilitate statistical inference. The Student’s t test (for two groups) or one-way analysis of variance followed by Tukey–Kramer post test (for more than two groups) using InStat (Graphpad InStat, USA) were performed to evaluate the statistical significance of data from biological replicates per experiment where appropriate. Values of P ≤ 0.05 were considered statistically significant. Data are presented graphically as mean ± standard error of the mean (S.E.M.) for experiments with a minimum of three biological replicates.
Results
SnoN promotes a TIF1γ–SnoN–PIAS1 multiprotein complex
PIAS1 and TIF1γ promote SUMOylation of SnoN [4, 8, 16]. A key question raised by these studies is whether and how the activity of PIAS1 and TIF1γ as SUMO E3 ligases is coordinated. To address this question, we first determined if PIAS1 and TIF1γ interact. Co-immunoprecipitation analyses showed that TIF1γ and PIAS1 formed a complex (Fig. 1a–d). SnoN associates with each of TIF1γ and PIAS1 [4, 8]. The 16 amino acid motif termed TIPtide within the SnoN isoform SnoN1 specifies its association with TIF1γ [8]. The isoform SnoN2 lacks the TIPtide motif [11]. We found that PIAS1 interacted similarly with SnoN and SnoN2, suggesting PIAS1 may interact with SnoN via a distinct region than the TIPtide motif (Fig. 1e). These data raised the question of whether SnoN impacts the interaction of TIF1γ and PIAS1. We found that expression of exogenous SnoN promoted the TIF1γ–PIAS1 association (Fig. 1a, b). Conversely, knockdown of endogenous SnoN reduced the TIF1γ–PIAS1 interaction (Fig. 1c, d). Together, these data suggest that TIF1γ and PIAS1 associate in a potentially SnoN-dependent manner.
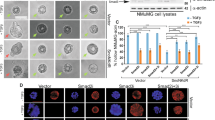
a Lysates of 293T cells transfected with vector control or a plasmid containing cDNA encoding HA-TIF1γ, FLAG-PIAS1, alone or in combination, together with a plasmid containing cDNA encoding Renilla luciferase (RLuc) (−) alone, or RLuc–SnoN fusion (+), were subjected to PIAS1 immunoprecipitation (α-FLAG IP) followed by TIF1γ (α-HA), SnoN (α-SnoN), and PIAS1 (α-FLAG) immunoblotting (IB). Lysates were subjected to TIF1γ (α-HA), SnoN (α-SnoN), and PIAS1 (α-FLAG) immunoblotting (IB) as the input. b Bar graph represents relative mean ± SEM of coimmunoprecipitated TIF1γ normalized to the protein abundance of PIAS1 in the corresponding immunoprecipitated sample and TIF1γ in the input in the absence or presence of expressed SnoN from three biological replicates including from the data shown in the last two lanes of 1A (Student’s t test: ***P ≤ 0.001). c Lysates of 293T cells transfected with HA-PIAS1 and FLAG-TIF1γ together with a control RNAi vector (−) or one expressing short hairpin RNA (shRNA) targeting SnoN (SnoNi), were subjected to TIF1γ immunoprecipitation using a mouse α-FLAG antibody (+) or a mouse IgG (−) as a negative control, followed by PIAS1 (α-PIAS1), SnoN (α-SnoN), and TIF1γ (α-FLAG) immunoblotting (IB) (right hand side panel). Lysates were subjected to immunoblotting for TIF1γ (α-FLAG), SnoN (α-SnoN), and PIAS1 (α-PIAS1) immunoblotting as the input (left hand side panel). d Bar graph represents relative mean ± SEM of coimmunoprecipitated PIAS1, normalized to the protein abundance of immunoprecipitated TIF1γ and the input PIAS1, in the absence (−) or presence (+) of shRNA targeting endogenous SnoN (SnoNi), from four biological replicates including the one with the data shown in lanes 2 and 4 of right-hand side panel in c (Student’s t test: ***P ≤ 0.001). e Bar graph depicts relative input-normalized SnoN (mean ± SEM, n = 3 biological replicates) interacting with PIAS1 obtained by luciferase measurements of 80% of α-FLAG immunoprecipitation (IP) of lysates of 293T cells co-expressing FLAG-PIAS1 and RLuc alone (−), or in fusion with SnoN or SnoN2. Four percent of lysates (input) and 20% of the PIAS1 immunocomplexes were subjected to luciferase measurements and PIAS1 (α-FLAG) immunoblotting (IB), respectively. f Top panel: bar graph depicts relative input-normalized PIAS1 interaction with TIF1γ (mean ± SEM, n = 4 biological replicates) obtained by subjecting lysates of 293T cells, transfected with control vector (−) or one expressing FLAG-TIF1γ, HA-SnoN, alone or combined, together with a plasmid expressing RLuc only (−) or in fusion with PIAS1 (+), to TIF1γ immunoprecipitation (α-FLAG IP) followed by subjecting 10% of the immunocomplexes and 0.5% of the lysates to luciferase measurements, with the latter serving as the input values (ANOVA: **P ≤ 0.01; ***P ≤ 0.001). Lower panels: representative scans from one of the biological replicates in which 10% of the TIF1γ immunocomplexes were subjected to TIF1γ (α-FLAG) and SnoN (α-SnoN) immunoblotting (IB), and where lysates were subjected to TIF1γ (α-FLAG), SnoN (α-SnoN), and actin (α-actin) immunoblotting as the input, with the latter as a loading control. g Upper panel: bar graph depicts relative input-normalized PIAS1 interaction with SnoN (mean ± SEM, n = 4 biological replicates) obtained by SnoN immunoprecipitation (α-HA) of FLAG peptide eluates of the TIF1γ (FLAG) immunocomplexes of cells transfected and immunoprecipitated as described in a, followed by subjecting the SnoN immunocomplexes to luciferase measurements, with the input from a serving as the input values (ANOVA: ***P ≤ 0.001). Lower panels: representative scans from one of the three biological replicates showing TIF1γ (α-FLAG) and SnoN (α-SnoN) immunoblotting (IB) of 10% of the SnoN immunocomplexes.
The finding that SnoN promoted the interaction between TIF1γ and PIAS1 raised the possibility of a TIF1γ–SnoN–PIAS1 multiprotein complex. To address this question, we performed sequential co-immunoprecipitation assays. Lysates of cells transfected with the vector control or a plasmid expressing TIF1γ, PIAS1, and SnoN in different combinations were subjected to TIF1γ immunoprecipitation (α-FLAG IP), followed by SnoN immunoprecipitation (α-HA IP) of the eluates of the TIF1γ immunocomplexes. PIAS1, expressed as a RLuc-tagged protein, in each of these two immunoprecipitations was quantified using Renilla luciferase activity [19, 24]. As expected, SnoN coimmunoprecipitated with TIF1γ. The proportion of PIAS1 in TIF1γ immunocomplexes was enhanced by increased abundance of SnoN in these immunocomplexes (Fig. 1f). Importantly, PIAS1 and TIF1γ coimmunoprecipitated with SnoN in TIF1γ immuno-complex-containing eluates supporting the existence of a PIAS1–SnoN–TIF1γ trimeric complex (Fig. 1g). Collectively, these data suggest that SnoN supports the assembly of PIAS1–SnoN–TIF1γ multimeric protein complex in mammalian cells.
PIAS1 and TIF1γ cooperate to regulate SnoN SUMOylation
We next performed in vivo SUMOylation assays to address the question of the functional significance of formation of the PIAS1–SnoN–TIF1γ protein complex [4, 17]. As expected, coexpression of SnoN with SUMO led to NEM-sensitive SUMO-immunoreactive protein species in SnoN immunoprecipitations (Fig. 2a, b) [4]. Expression of exogenous PIAS1 or TIF1γ increased the proportion of SUMOylated SnoN (Fig. 2a, b). Combined expression of PIAS1 and TIF1γ further enhanced the proportion of SUMOylated SnoN (Fig. 2a, b). Knockdown of endogenous PIAS1 or TIF1γ reduced the proportion of SUMOylated SnoN in cells (Fig. 2a, b). Knockdown of endogenous TIF1γ and PIAS1 led to reciprocal reduction of the ability of expressed PIAS1 and TIF1γ to promote the proportion of SUMOylated SnoN (Fig. 2a, b), suggesting that PIAS1 and TIF1γ cooperate to promote SUMOylation of SnoN. Collectively, our data suggest that SnoN may act as an adapter to promote a TIF1γ–PIAS1 complex to promote its own SUMOylation.
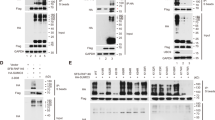
a NEM-treated lysates of 293T cells transfected with control plasmids, alone or together with plasmids encoding MYC-SnoN or HA-SUMO, alone or together, alone or along with different combinations of plasmids encoding FLAG-TIF1γ, FLAG-PIAS1, short hairpin RNA (sh) targeting PIAS1 (PIAS1i), or shRNA targeting TIF1γ (TIF1γi), were subjected to SnoN immunoprecipitation (α-MYC IP) followed by SUMO (α-HA) and SnoN (α-SnoN) immunoblotting. Lysates were subjected to TIF1γ, SnoN, PIAS1, and actin immunoblotting as the input, with the latter as a loading control. b The bar graph represents the mean ± SEM of proportion of SUMOylated SnoN relative to unmodified SnoN quantified from HA and SnoN immunoblots of SnoN immunoprecipitation and expressed relative to the proportion of SUMOylated SnoN in lysates of cells transfected with MYC/SnoN and HA/SUMO along with empty vectors as a control (lane 4). The data are from three biological replicates including from the replicate whose data are shown in a (ANOVA: **P ≤ 0.01; ***P ≤ 0.001).
PIAS1 and TIF1γ regulate morphogenesis of mammary epithelial organoids
The finding that SnoN promotes a TIF1γ–PIAS1 SUMO E3 ligase complex raised the question of the biological significance of the interaction. EMT is a fundamental cellular process in tissue and organ development and contributes to homeostasis [25, 26]. The secreted protein transforming growth factor beta (TGFβ) induces EMT in the developing and adult organism [27, 28]. Both PIAS1 and TIF1γ act via SUMOylated SnoN to suppress TGFβ-induced EMT, raising the question of whether PIAS1–TIF1γ reciprocally regulate each other’s role in EMT. NMuMG murine mammary epithelial cells represent a widely used model to investigate EMT. Three-dimensional culture approaches, where cells are grown in the context of an extracellular matrix (ECM) support system, provide suitable in vivo insight on the regulation of cellular processes like EMT [29]. Isolated non-transformed epithelial cells, e.g., NMuMG cells, cultured in the context the ECM components contained in the Matrigel, replicate forming aggregates that organize into multicellular spherical organoids with hollow centers or lumens (Fig. S1B) [8, 21]. A key feature of these acini is the apical (toward lumen) to basal (towards the ECM) polarity, and which is lost in EMT. EMT-related changes manifest as lumen filling and may include budding and deformation of epithelial cell-derived organoids [8, 21]. The epithelial marker E-cadherin in these epithelial cell-derived organoids localizes at the basolateral junction (Fig. S1D). A hallmark of EMT is the mislocalization and loss of E-cadherin, which can be detected by indirect immunofluorescence [4, 8, 21].
We used a Matrigel-based three-dimensional culture system, to address the question of whether TIF1γ and PIAS1 coordinately regulate TGFβ-induced EMT [8, 21]. PIAS1 suppression of TGFβ-induced EMT has only been reported in conventional two-dimensional cultures [4]. We, thus, first determined if PIAS1–SnoN SUMOylation axis regulates EMT in epithelial cell-derived organoids. We found that expression of exogenous PIAS1 suppressed TGFβ-induced acini filling and deformation of three-dimensional NMuMG cell-derived organoids, which was reversed by expression of the SUMO-loss of function SnoNKdR (Fig. S1A–C). Conversely, expression of the SUMO E3 ligase inactive PIASCS promoted acini filling and deformation of the organoids even in the absence of TGFβ, and which was reversed by expression of the SUMO gain of function SUMO–SnoN, a stable linear protein fusion of SUMO and SnoN (Fig. S1E–G). Indirect immunofluorescence analyses demonstrated that PIAS1 acts via SUMOylated SnoN to suppress TGFβ-induced mislocalization and loss of E-cadherin in NMuMG cell-derived organoids (Fig. S1D, H). Together, these data show that similar to TIF1γ [8], PIAS1 suppresses TGFβ-induced EMT in the mammary epithelial cell-derived organoids in a SUMOylated SnoN-dependent manner.
Next, we addressed the question of whether PIAS1 and TIF1γ coordinately suppress EMT in NMuMG cell-derived organoids. We found that knockdown of endogenous TIF1γ and PIAS1 abrogated the ability of exogenous PIAS1 and TIF1γ, respectively, to suppress TGFβ-induced EMT as assessed by acini filling, and E-cadherin loss and mislocalization of the organoids (Fig. 3a–d), suggesting that endogenous and exogenous PIAS1 and TIF1γ intersect to suppress TGFβ-induced EMT in the mammary epithelial-derived organoids. In other experiments, we found that expression of the SUMO E3 ligase inactive TIF1γΔC, which is deleted in C-terminal PHD and Bromo domains, inhibited exogenous PIAS1 suppression of TGFβ-induced EMT in NMuMG cell-derived organoids. Conversely, the SUMO E3 ligase inactive PIAS1CS reduced exogenous TIF1γ suppression of TGFβ-induced EMT in the NMuMG-derived organoids (Fig. 3e–h). Collectively, these data suggest that a TIF1γ–PIAS1 complex acts in a SUMO E3 ligase-dependent manner to suppress EMT in epithelial-derived organoids.
a PIAS1, TIF1γ, and actin immunoblotting of lysates of NMuMG cells expressing PIAS1, TIF1γ, PIAS1i, and TIF1γi alone or together, and which were used to generate three-dimensional organoids shown in b–d. b Representative DIC light microscopy micrographs of untreated or 100pM TGFβ-treated 8-day-old organoids derived from NMuMG cells transfected and assessed as in a. Green and red arrows indicate acinar and filled organoids, respectively. c Bar graph depicts mean ± SEM proportion of acinar organoids expressed as a percentage of total colonies counted for each experimental condition from three biological replicates including the replicates with data shown in b. d Representative fluorescence microscopy scans of E-cadherin-(red) and nuclei-(blue) stained fixed 8-day-old three-dimensional organoids derived from NMuMG cells transfected and assessed in a, b. e PIAS1, FLAG, and actin immunoblotting of lysates of NMuMG cells expressing PIAS1, FLAG-TIF1γ, PIAS1CS, and FLAG-TIF1γΔC alone or together, and which were used to generate three-dimensional organoids shown in f–h. f Representative DIC light microscopy micrographs of untreated or 100pM TGFβ-treated 8-day-old organoids derived from NMuMG cells transfected and assessed as in e. Green and red arrows indicate acinar and filled organoids, respectively. g Bar graph depicts mean ± SEM proportion of acinar organoids expressed as a percentage of total colonies counted for each experimental condition from three biological replicates including the replicates with data shown in f. h Representative fluorescence microscopy scans of E-cadherin-(red) and nuclei-(blue) stained fixed 8-day-old three-dimensional organoids derived from NMuMG cells transfected and assessed in e, f. Statistical difference, ANOVA: **P ≤ 0.01; ***P ≤ 0.001. Scale bar indicates 50 μm.
Next, we addressed the question of whether SUMOylated SnoN plays a role in the TIF1γ–PIAS1-mediated suppression of EMT. The TIPtide motif specifies SnoN interaction with TIF1γ but not with PIAS1 (Fig. 1e and [8]). We confirmed that TIPtide competes with SnoN for association with TIF1γ (Fig. 4a). Accordingly, we found that expression of TIPtide promoted EMT in NMuMG cell-derived organoids even in the absence of TGFβ, as demonstrated by lumen filling and disorganization of the acini, and E-cadherin loss and mislocalization in these organoids (Fig. 4b–e). In addition, TIPtide suppressed TIF1γ reversal of TGFβ-induced EMT (Fig. 4b–e). Remarkably, TIPtide opposed PIAS1 suppression of TGFβ-induced EMT in the NMuMG cell-derived organoid (Fig. 4b–e). Collectively, these data suggest that the TIF1γ–PIAS1 SUMO E3 ligase complex acts via SnoN SUMOylation to suppress EMT in the mammary epithelial-derived organoid.
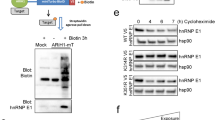
a Lysates of 293T cells expressing HA-TIF1γ, FLAG-SnoN, and GFP-TIPtide alone or together were subjected to SnoN immunoprecipitation (FLAG IP) followed by TIF1γ (α-HA) and SnoN (α-FLAG) immunoblotting. Lysates were subjected to immunoblotting for TIF1γ (α-HA), SnoN (α-FLAG), TIPtide (α-GFP), and actin (α-actin) as the input, with the latter as a loading control. b TIF1γ, PIAS1, GFP, and actin immunoblotting of lysates of NMuMG cells expressing TIF1γ, PIAS1, and GFP-TIPtide alone or together, and which were used to generate three-dimensional organoids shown in c–e. c Representative DIC light microscopy micrographs of untreated or 100pM TGFβ-treated 8-day-old organoids derived from NMuMG cells transfected and assessed as in b. Green and red arrows indicate acinar and filled organoids, respectively. d Bar graph depicts mean ± SEM proportion of acinar organoids expressed as a percentage of total colonies counted for each experimental condition from three biological replicates including the replicates with data shown in c. e Representative fluorescence microscopy scans of E-cadherin-(red) and nuclei-(blue) stained fixed 8-day-old three-dimensional organoids derived from NMuMG cells transfected and assessed in b, c. Statistical difference, ANOVA: ***P ≤ 0.001. Scale bar indicates 50 μm.
A TIF1γ–PIAS1 SUMO E3 ligase complex regulates EMT in breast cancer cell organoids
EMT plays key roles in cancer invasiveness and metastasis including in breast cancer [25] raising the question whether the PIAS1–TIF1γ SUMO E3 ligase complex plays a role in cancer. PIAS1 acts via SUMOylation of SnoN to suppress invasive behaviour of the human triple negative breast cancer MDA-MB-231 cell-derived organoids [17]. Using a three-dimensional culture model system, we addressed, first, the question of whether TIF1γ regulates breast cancer cell invasive behaviour. MDA-MB-231 cells grown in the context of ECM, e.g., Matrigel, form multicellular structures that are mostly smooth-surfaced solid/lacking lumens spherically-shaped organoids, a characteristic of transformed epithelial cells (Figs. 5, 6, S2, S3) [17, 18, 30]. TGFβ leads to invasiveness, budding, and disruption of the 3D-MDA-MB-231 cell-derived organoids (Figs. 5, 6, S2, S3) [17, 18, 30]. We found that knockdown of endogenous TIF1γ promoted disruption and invasiveness of the MDA-MB231 cell-derived organoids even in the absence of TGFβ, suggesting that endogenous TIF1γ suppresses the invasive growth of MDA-MB-231 cell-derived organoids (Fig. S2A–C). Consistently, expression of exogenous TIF1γ suppressed, while the SnoN-interaction defective TIF1γCS or the SUMO E3 ligase inactive TIF1γΔC, promoted the invasive growth of MDA-MB-231 cell-derived organoids (Fig. S2D–F). Knockdown of endogenous SnoN inhibited exogenous TIF1γ suppression of the invasive behaviour of 3D-organoids in the absence or presence of TGFβ (Fig. S2G–I). Together, these data suggested that endogenous TIF1γ acts in a SUMO E3 ligase and endogenous SnoN-dependent manner to suppress TGFβ-induced disruptive morphogenetic behaviour of the MDA-MB-231 cell-derived organoids. Accordingly, we found that expression of SUMO–SnoN suppressed the ability of knockdown of endogenous TIF1γ or expression of TIF1γΔC to promote disruption of organoids (Fig. S3A–F), while expression of SnoNKdR reduced TIF1γ suppression of the invasive phenotype of the spheroids (Fig. S3G–I). Collectively, these data show that the SUMO E3 ligase TIF1γ acts via SnoN SUMOylation to suppress TGFβ-induced invasive growth of the triple negative breast cancer cell-derived organoids.
a PIAS1, TIF1γ, and actin immunoblotting of lysates of MDA-MB-231 cells expressing PIAS1, TIF1γ, PIAS1i, and TIF1γi alone or together, and which were used to generate three-dimensional organoids shown in b–d. b Representative DIC light microscopy micrographs of untreated or 100pM TGFβ-treated 8-day-old organoids derived from MDA-MB-231 cells transfected and assessed as in a. Green and red arrows indicate non-deformed and disrupted organoids, respectively. c Bar graph depicts mean ± SEM proportion of non-deformed organoids expressed as a percentage of total colonies counted for each experimental condition from three biological replicates including the replicates with data shown in b. d PIAS1, FLAG, and actin immunoblotting of lysates of MDA-MB-231 cells expressing PIAS1, FLAG-TIF1γ, PIAS1CS, and FLAG-TIF1γΔC alone or together, and which were used to generate three-dimensional organoids shown in e–g. e Representative DIC light microscopy micrographs of untreated or 100pM TGFβ-treated 8-day-old organoids derived from MDA-MB-231 cells transfected and assessed as in d. Green and red arrows indicate non-deformed and disrupted organoids, respectively. f Bar graph depicts mean ± SEM proportion of non-deformed organoids expressed as a percentage of total colonies counted for each experimental condition from five biological replicates including the replicates with data shown in e. g Representative fluorescence microscopy scans of E-cadherin-(red) and nuclei-(blue) stained fixed 8-day-old three-dimensional organoids derived from MDA-MB-231 cells transfected and assessed in d, e. Statistical difference, ANOVA: ***P ≤ 0.001. Scale bar indicates 50 μm.
a TIF1γ, PIAS1, GFP, and actin immunoblotting of lysates of MDA-MB-231 cells expressing TIF1γ, PIAS1, and GFP-TIPtide alone or together, and which were used to generate three-dimensional organoids shown in b–d. b Representative DIC light microscopy micrographs of untreated or 100pM TGFβ-treated 8-day-old organoids derived from MDA-MB-231 cells transfected and assessed as in a. Green and red arrows indicate non-deformed and disrupted organoids, respectively. c Bar graph depicts mean ± SEM proportion of non-deformed organoids expressed as a percentage of total colonies counted for each experimental condition from five biological replicates including the replicates with data shown in b. d Representative fluorescence microscopy scans of E-cadherin-(red) and nuclei-(blue) stained fixed 8-day-old three-dimensional organoids derived from NMuMG cells transfected and assessed in b, c. Statistical difference, ANOVA: *P ≤ 0.05; ***P ≤ 0.001. Scale bar indicates 50 μm. The scale bar indicated in the top left image (first image) in d reflects the relative 50 μm ruler of that image and that of other images that do not contain a scale bar. Any other images that display a different length of the 50 μm scale bar reflects images that are scaled down or up relative to the first image.
Next, we addressed the question of whether the TIF1γ–PIAS1 SUMO E3 ligase complex regulates the invasive behaviour of MDA-MB-231 cell-derived organoids. Expression of exogenous PIAS1 or TIF1γ suppressed, while knockdown of endogenous PIAS1 or TIF1γ promoted the invasive growth of three-dimensional MDA-MB-231 cell-derived organoids (Figs. S2, 5, and [17, 18]). Importantly, we found that knockdown of endogenous TIF1γ and PIAS1 abrogated exogenous PIAS1 and TIF1γ, respectively, suppression of TGFβ-induced invasive phenotype of the MDA-MB-231 cell-derived organoids (Fig. 5a–c), suggesting that endogenous PIAS1 and TIF1γ cooperate to suppress TGFβ-induced invasive/disruptive behaviour of the MDA-MB-231 cell-derived spheroids. In other analyses, we found that expression of the SUMO E3 ligase inactive TIF1γΔC and PIAS1CS inhibited the ability of PIAS1 and TIF1γ, respectively, to suppress TGFβ-induced invasive characteristics of MDA-MB-231 cell-derived organoids (Fig. 5d–f). MDA-MB-231 carcinomas represent cells displaying at least partial EMT [31], whereby the protein abundance of E-cadherin in these cells can very low to undetectable compared to other mammary epithelial and carcinoma cells [32]. Interestingly, MDA-MB-231 multicellular structures show a relatively higher protein abundance of E-cadherin as compared to 2D-cultured cells [32]. We found that expression of exogenous PIAS1 or TIF1γ suppressed E-cadherin loss by TGFβ, which was reversed by expression of the SUMO E3 ligase inactive TIF1γ and PIAS1, respectively (Fig. 5g) [18]. Together, these data suggest that the TIF1γ–PIAS1 SUMO E3 ligase complex suppresses TGFβ-induced EMT, including invasive growth and E-cadherin loss, in the MDA-MB-231 cell-derived organoids.
Next, we assessed the role of SnoN in mediating TIF1γ–PIAS1-suppression of EMT in the MDA-MB-231 cell-derived organoids. We found that expression of TIPtide promoted EMT as displayed by disruptive/deformed behaviour and loss of E-cadherin in the MDA-MB-231 cell-derived organoids even in the absence of TGFβ as compared to the control (Fig. 6a–d). Importantly, TIPtide suppressed the ability of TIF1γ and PIAS1 to reverse TGFβ-induced EMT in the MDA-MB-231 cell-derived organoids (Fig. 6a–d). Collectively, these data suggest that that the TIF1γ–PIAS1 complex acts via SnoN SUMOylation to suppress TGFβ-induced EMT-associated phenotypes in the TNBC carcinoma cell-derived organoids.
The protein abundance of PIAS1 correlated positively with survival in women with breast cancer according to a recent breast cancer tissue microarray study that included samples from luminal breast cancer subtypes [17], raising the key question whether the PIAS1–TIF1γ SUMO E3 complex regulates EMT in luminal breast cancer-derived cells. We used a three-dimensional model of the luminal breast cancer MCF7 cells, that express estrogen and progesterone receptors and display an epithelial phenotype, to address this question [33]. We found that Matrigel-grown MCF7 cells formed spherical-acinar type of organoids characterized by junctional localization of E-cadherin (Fig. 7b–d) [34]. TGFβ induced EMT in the MCF7 cell-derived organoids characterized by filling and disruption of the organoids accompanied by reduction of the protein abundance and mislocalization of E-cadherin (Fig. 7b–d). Knockdown of endogenous SnoN induced EMT in MCF7-derived organoids even in the absence of TGFβ (Fig. 7a–d), which was reversed by expression of the SnoNi-resistant SnoNres indicating that endogenous SnoN promotes an epithelial phenotype in the luminal breast cancer MCF7 cell-derived organoids. We also found that the SUMO loss of function SnoNKdR promoted, while the SUMO gain of function SUMO–SnoN suppressed EMT induction suggesting that SUMOylation is important for SnoN to inhibit TGFβ-induced EMT in the MCF7 cell-derived organoids (Figs. 7e–h, S4). Consistently, the SUMO E3 ligases PIAS1 and TIF1γ suppressed TGFβ-induced EMT, which were reversed by SnoNKdR expression (Fig. 7e–h). Conversely, reduction of the protein abundance or SUMO E3 ligase activity of PIAS1 or TIF1γ promoted EMT in MCF7 breast cancer cell-derived organoids even in the absence of TGFβ, which was rescued by SUMO–SnoN expression (Fig S4A–D). Similar to NMuMG and MDA-MB231 cell-derived organoids, we found that the SUMO E3 ligase inactive TIF1γΔC and PIAS1CS inhibited the ability of PIAS1 and TIF1γ, respectively, to suppress TGFβ induced EMT of MCF7-derived organoids (Fig. 8a–d). Together, these data reveal that the PIAS1–TIF1γ SUMO E3 ligase complex acts via SUMOylated SnoN to suppress EMT in luminal breast cancer cell-derived organoids.
a SnoN and actin immunoblots of lysates of MCF7 cells expressing a control vector (−) or a SnoN specific shRNA (SnoNi), with or without a SnoNi-resistant SnoN expression construct (SnoNres). b Representative DIC light microscopy micrographs of untreated or 100pM TGFβ-treated 8-day-old organoids derived from MCF7 cells transfected and assessed as in a. Green and red arrows indicate hollow center and filled acinis, respectively. c Bar graph depicts mean ± SEM proportion of hollow center acini expressed as a percentage of total colonies counted for each experimental condition from three biological replicates including the one with data shown in b. d Representative fluorescence microscopy scans of E-cadherin-(red) and nuclei-(blue) stained fixed 8-day-old three-dimensional organoids derived from MCF7 cells transfected and assessed in b, c. e PIAS1, TIF1γ, SnoN, and actin immunoblots of lysates of MCF7 cells expressing PIAS1, TIF1γ, SnoNKdR, alone or together. f Representative DIC light microscopy micrographs of untreated or 100pM TGFβ-treated 8-day-old organoids derived from MCF7 cells transfected and assessed as in e. Green and red arrows indicate acinar and filled organoids, respectively. g Bar graph depicts mean ± SEM proportion of acinar organoids expressed as a percentage of total colonies counted for each experimental condition from three biological replicates including the one with data shown in f. h Representative fluorescence microscopy scans of E-cadherin-(red) and nuclei-(blue) stained fixed 8-day-old three-dimensional organoids derived from MCF7 cells transfected and assessed in f, g. Statistical difference, ANOVA: *P ≤ 0.05; ***P ≤ 0.001. Scale bar indicates 50 μm.
a PIAS1, FLAG, and actin immunoblotting of lysates of MCF7 cells expressing PIAS1, FLAG-TIF1γ, PIAS1CS, and FLAG-TIF1γΔC alone or together. b Representative DIC light microscopy micrographs of untreated or 100pM TGFβ-treated 8-day-old organoids derived from MCF7 cells transfected and assessed as in a. Green and red arrows indicate acinar and filled organoids, respectively. c Bar graph depicts mean ± SEM proportion of acinar organoids expressed as a percentage of total colonies counted for each experimental condition from three biological replicates including the one with data shown in b. d Representative fluorescence microscopy scans of E-cadherin-(red) and nuclei-(blue) stained fixed 8-day-old three-dimensional organoids derived from MCF7 cells transfected and assessed in a, b. Statistical difference, ANOVA: ***P ≤ 0.001. Scale bar indicates 50 μm. The scale bar indicated in the top left image (first image) in d reflects the relative 50 μm ruler of that image and that of other images that do not contain a scale bar. Any other images that display a different length of the 50 μm scale bar reflects images that are scaled down or up relative to the first image.
Collectively, our study reveals that the transcriptional regulator SnoN promotes the assembly of a PIAS1–TIF1γ SUMO E3 ligase complex that promotes its SUMOylation and ability to suppress TGFβ-induced induction in organoids derived from breast epithelial and carcinoma cells. These data add insights into mechanisms by which the SUMO pathway specifies SUMO substrates for SUMOylation with important consequences for biological responses in normal and pathological conditions.
Discussion
In this study, we have discovered a novel link between the major transcriptional regulator SnoN and the SUMO pathway with implications for SnoN SUMOylation and ability to regulate the fundamental process of EMT and epithelial and carcinoma-derived tissue phenotypes. Our data reveal that SnoN promotes a protein complex containing the SUMO E3 ligases PIAS1 and TIF1γ that enhances SnoN SUMOylation. Functionally, our novel findings point to a co-dependency between the SUMO E3 ligases TIF1γ and PIAS1 in suppression of EMT induction by TGFβ and associated morphological changes of breast epithelial cells and carcinomas-tissue organoids. The idea that a SUMO E3 ligase complex can promote SUMOylation of a SUMO substrate with functional significance should advance our understanding of mechanisms that regulate protein targeting by the SUMO pathway and biological implications.
The findings revealed in this study that as a SUMO substrate, SnoN helps in the assembly of a multiprotein complex containing two SUMO E3 ligases to promote its modification by the SUMO pathway begins to address an outstanding question in the SUMO field regarding how substrate specificity and hence modification is determined given the limited number of SUMO E3 ligases as opposed to SUMO substrates identified in the literature [5]. Indeed, several SUMO substrates including p53 [9, 10, 35, 36] and Smad4 [37,38,39] have been described to be targeted by distinct SUMO E3 ligases raising the question of the nature of interplay between different SUMO E3 ligases targeting a selective substrate.
SUMOylation is a ubiquitous protein modification found in multicellular organism with over 6000 protein substrates identified in human cells [40]. Different extracellular and intracellular stimuli have been suggested to alter the abundance of SUMOylated species of specific substrates with importance for normal cellular functions in development and homeostasis [5, 41]. Members of the PIAS family of SUMO E3 ligases have been reported to target multiple substrates raising the question of the implications to biological responses [6, 7]. Our findings that PIAS1 and TIF1γ coordinately enhance SnoN SUMOylation and hence suppression of EMT and associated phenotypic responses of cell-derived organoids, illuminates a regulatory mechanism by which PIAS1 controls EMT via the SUMO substrate SnoN. Our data also lead to the question of whether other members of the PIAS SUMO E3 ligase family act collaboratively with other distinct SUMO E3 ligases to promote the SUMOylation of a given substrate to regulate specific biological responses.
The protein TIF1γ has been suggested to be a SUMO substrate of PIAS1, whereby SUMOylation may regulate TIF1γ suppression of TGFβ-induced EMT in mammary cells [42]. Thus, our novel findings described in this study raise the intriguing question of whether SnoN by promoting a TIF1γ–PIAS1 association may play a role in PIAS1’s ability to promote TIF1γ SUMOylation. In turn, SUMOylation may promote TIF1γ to act as a SUMO E3 ligase for SnoN and hence EMT regulation. Addressing this potential multifaceted function of the TIF1γ–SnoN–PIAS1 complex would be interesting to pursue in future studies.
Our findings that SnoN promotes a TIF1γ–PIAS1 SUMO E3 ligase complex regulating its SUMOylation and hence EMT in mammary epithelial cells and breast carcinomas-derived tissue organoids have implications for regulation of tissue development and tumorigenesis. In future studies, it will be important to determine the effect of alteration in the abundance or activity of PIAS1 or TIF1γ on developmental processes including mammary gland maturation, and tumorigenesis and progression including invasion and metastasis. Our findings also raise the key question whether the TIF1γ–PIAS1–SnoN SUMOylation axis plays a role in epithelial cells and carcinomas derived from other types of organs including gastrointestinal tract and pancreas.
Changes in the abundance of elements of the SUMO modification machinery have been associated with pathological conditions including cancer. In addition, variations in the abundance of SUMO modified substrates have been linked to increased cancer metastasis, resistance to therapy and relapse [1, 27, 41, 43]. Thus, it will be important to determine in future studies whether perturbations in the TIF1γ–PIAS1–SUMOylated SnoN axis identified in this study dictate cancer responses of cancer cells to chemotherapeutic and targeted therapies.
In conclusion, the findings presented in the current study uncovers novel mechanisms by which the SUMO pathway specifies and promotes the SUMOylation of protein substrates. These findings add important insights into mechanisms that control fundamental cellular processes in normal and disease conditions.
Change history
18 August 2020
A Correction to this paper has been published: https://doi.org/10.1038/s41418-020-00611-z
References
Bettermann K, Benesch M, Weis S, Haybaeck J. SUMOylation in carcinogenesis. Cancer Lett. 2012;316:113–25.
Palvimo JJ. PIAS proteins as regulators of small ubiquitin-related modifier (SUMO) modifications and transcription. Biochemical Soc Trans. 2007;35:1405–8.
Johnson ES. Protein modification by SUMO. Annu Rev Biochem. 2004;73:355–82.
Netherton SJ, Bonni S. Suppression of TGFbeta-induced epithelial-mesenchymal transition like phenotype by a PIAS1 regulated sumoylation pathway in NMuMG epithelial cells. PLoS ONE. 2010;5:e13971.
Pichler A, Fatouros C, Lee H, Eisenhardt N. SUMO conjugation—a mechanistic view. Biomol Concepts. 2017;8:13–36.
Rytinki MM, Kaikkonen S, Pehkonen P, Jaaskelainen T, Palvimo JJ. PIAS proteins: pleiotropic interactors associated with SUMO. Cell Mol Life Sci. 2009;66:3029–41.
Rabellino A, Andreani C, Scaglioni PP. The role of PIAS SUMO E3-ligases in cancer. Cancer Res. 2017;77:1542–7.
Ikeuchi Y, Dadakhujaev S, Chandhoke AS, Huynh MA, Oldenborg A, Ikeuchi M, et al. TIF1gamma protein regulates epithelial-mesenchymal transition by operating as a small ubiquitin-like modifier (SUMO) E3 ligase for the transcriptional regulator SnoN1. J Biol Chem. 2014;289:25067–78.
Kahyo T, Nishida T, Yasuda H. Involvement of PIAS1 in the sumoylation of tumor suppressor p53. Mol Cell. 2001;8:713–8.
Wu SY, Chiang CM. p53 sumoylation: mechanistic insights from reconstitution studies. Epigenetics. 2009;4:445–51.
Bonni S, Bonni A. SnoN signaling in proliferating cells and postmitotic neurons. FEBS Lett. 2012;586:1977–83.
Deheuninck J, Luo K. Ski and SnoN, potent negative regulators of TGF-beta signaling. Cell Res. 2009;19:47–57.
Hsu YH, Sarker KP, Pot I, Chan A, Netherton SJ, Bonni S. Sumoylated SnoN represses transcription in a promoter-specific manner. J Biol Chem. 2006;281:33008–18.
Ikeuchi Y, Stegmuller J, Netherton S, Huynh MA, Masu M, Frank D, et al. A SnoN-Ccd1 pathway promotes axonal morphogenesis in the mammalian brain. J Neurosci. 2009;29:4312–21.
Sarker KP, Wilson SM, Bonni S. SnoN is a cell type-specific mediator of transforming growth factor-beta responses. J Biol Chem. 2005;280:13037–46.
Yin X, Xu C, Zheng X, Yuan H, Liu M, Qiu Y, et al. SnoN suppresses TGF-beta-induced epithelial-mesenchymal transition and invasion of bladder cancer in a TIF1gamma-dependent manner. Oncol Rep. 2016;36:1535–41.
Chanda A, Chan A, Deng L, Kornaga EN, Enwere EK, Morris DG, et al. Identification of the SUMO E3 ligase PIAS1 as a potential survival biomarker in breast cancer. PLoS ONE. 2017;12:e0177639.
Dadakhujaev S, Salazar-Arcila C, Netherton SJ, Chandhoke AS, Singla AK, Jirik FR, et al. A novel role for the SUMO E3 ligase PIAS1 in cancer metastasis. Oncoscience. 2014;1:229–40.
Pot I, Patel S, Deng L, Chandhoke AS, Zhang C, Bonni A, et al. Identification of a novel link between the protein kinase NDR1 and TGFbeta signaling in epithelial cells. PLoS ONE. 2013;8:e67178.
Kingston RE, Chen CA, Rose JK. Calcium phosphate transfection. Curr Protoc Mol Biol. 2003;Chapter 9:Unit 91.
Chandhoke AS, Karve K, Dadakhujaev S, Netherton S, Deng L, Bonni S. The ubiquitin ligase Smurf2 suppresses TGFbeta-induced epithelial-mesenchymal transition in a sumoylation-regulated manner. Cell Death Differ. 2016;23:876–88.
Bonni S, Wang HR, Causing CG, Kavsak P, Stroschein SL, Luo K, et al. TGF-beta induces assembly of a Smad2-Smurf2 ubiquitin ligase complex that targets SnoN for degradation. Nat Cell Biol. 2001;3:587–95.
Blainey P, Krzywinski M, Altman N. Points of significance: replication. Nat Methods. 2014;11:879–80.
Barrios-Rodiles M, Brown KR, Ozdamar B, Bose R, Liu Z, Donovan RS, et al. High-throughput mapping of a dynamic signaling network in mammalian cells. Science. 2005;307:1621–5.
De Craene B, Berx G. Regulatory networks defining EMT during cancer initiation and progression. Nat Rev Cancer. 2013;13:97–110.
Ye X, Weinberg RA. Epithelial-mesenchymal plasticity: a central regulator of cancer progression. Trends Cell Biol. 2015;25:675–86.
Chanda A, Sarkar A, Bonni S. The SUMO System and TGFbeta signaling interplay in regulation of epithelial–mesenchymal transition: implications for cancer progression. Cancers. 2018;10:264.
Xu J, Lamouille S, Derynck R. TGF-beta-induced epithelial to mesenchymal transition. Cell Res. 2009;19:156–72.
Debnath J, Brugge JS. Modelling glandular epithelial cancers in three-dimensional cultures. Nat Rev Cancer. 2005;5:675–88.
Chandhoke AS, Chanda A, Karve K, Deng L, Bonni S. The PIAS3-Smurf2 sumoylation pathway suppresses breast cancer organoid invasiveness. Oncotarget. 2017;8:21001–14.
Wei Z, Shan Z, Shaikh ZA. Epithelial-mesenchymal transition in breast epithelial cells treated with cadmium and the role of Snail. Toxicol Appl Pharmacol. 2018;344:46–55.
Aiello NM, Maddipati R, Norgard RJ, Balli D, Li J, Yuan S, et al. EMT subtype influences epithelial plasticity and mode of cell migration. Dev Cell. 2018;45:681–95.e4.
Comsa S, Cimpean AM, Raica M. The story of MCF-7 breast cancer cell line: 40 years of experience in research. Anticancer Res. 2015;35:3147–54.
Krause S, Maffini MV, Soto AM, Sonnenschein C. The microenvironment determines the breast cancer cells’ phenotype: organization of MCF7 cells in 3D cultures. BMC Cancer. 2010;10:263.
Kung CP, Khaku S, Jennis M, Zhou Y, Murphy ME. Identification of TRIML2, a novel p53 target, that enhances p53 SUMOylation and regulates the transactivation of proapoptotic genes. Mol Cancer Res. 2015;13:250–62.
Weger S, Hammer E, Heilbronn R. Topors acts as a SUMO-1 E3 ligase for p53 in vitro and in vivo. FEBS Lett. 2005;579:5007–12.
Lee PS, Chang C, Liu D, Derynck R. Sumoylation of Smad4, the common Smad mediator of transforming growth factor-beta family signaling. J Biol Chem. 2003;278:27853–63.
Long J, Wang G, He D, Liu F. Repression of Smad4 transcriptional activity by SUMO modification. Biochem. J. 2004;379:23–9.
Ohshima T, Shimotohno K. Transforming growth factor-beta-mediated signaling via the p38 MAP kinase pathway activates Smad-dependent transcription through SUMO-1 modification of Smad4. J Biol Chem. 2003;278:50833–42.
Hendriks IA, Lyon D, Young C, Jensen LJ, Vertegaal AC, Nielsen ML. Site-specific mapping of the human SUMO proteome reveals co-modification with phosphorylation. Nat Struct Mol Biol. 2017;24:325–36.
Seeler JS, Dejean A. SUMO and the robustness of cancer. Nat Rev Cancer. 2017;17:184–97.
Fattet L, Ay AS, Bonneau B, Jallades L, Mikaelian I, Treilleux I, et al. TIF1gamma requires sumoylation to exert its repressive activity on TGFbeta signaling. J Cell Sci. 2013;126:3713–23.
Anderson DB, Zanella CA, Henley JM, Cimarosti H. Sumoylation: implications for neurodegenerative diseases. Adv Exp Med Biol. 2017;963:261–81.
Acknowledgements
We would like to thank Dr. Derrick Rancourt and Dr. Tannin Schmidt for use of the Olympus IX70 and the Olympus Fluoview FV1000 fluorescence microscopes, respectively. This work was supported by grants from the Natural Sciences and Engineering Research Council of Canada (NSERC), the Canadian Cancer Society (CCS), the Breast Cancer Society of Canada (BCSC) and the Calgary Centre for Cancer Research (CCCR) to S. Bonni, and the National Institutes of Health (NIH) - NS041021 to A. Bonni. A. Chanda is a recipient of a Charbonneau Cancer Institute Director’s Award for Excellence in Research Productivity, an Eyes High International Doctoral Scholarship, a William H Davies Medical Research Scholarship, an Achievers in Medical Science (AIMS) Graduate Recruitment Scholarship, and an Eyes High Recruitment Award from the University of Calgary. K. Karve and A. Sarkar are recipients of a Doctoral Scholarship and an Eyes High Doctoral Recruitment Scholarship, respectively, from the University of Calgary.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
ASC is currently employed at Fog Pharma, USA. However, this study is neither funded nor associated in any manner with Fog Pharma. The other authors declare no conflict of interest.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Edited by M. Bianchi
Rights and permissions
About this article
Cite this article
Chanda, A., Ikeuchi, Y., Karve, K. et al. PIAS1 and TIF1γ collaborate to promote SnoN SUMOylation and suppression of epithelial–mesenchymal transition. Cell Death Differ 28, 267–282 (2021). https://doi.org/10.1038/s41418-020-0599-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41418-020-0599-8
This article is cited by
-
The emerging roles of SUMOylation in the tumor microenvironment and therapeutic implications
Experimental Hematology & Oncology (2023)
-
Sumoylated SnoN interacts with HDAC1 and p300/CBP to regulate EMT-associated phenotypes in mammary organoids
Cell Death & Disease (2023)
-
SUMOylation of Rho-associated protein kinase 2 induces goblet cell metaplasia in allergic airways
Nature Communications (2023)
-
PIAS1 potentiates the anti-EBV activity of SAMHD1 through SUMOylation
Cell & Bioscience (2021)