Abstract
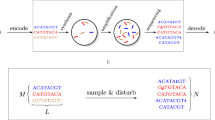
Here we report a simple and flexible method for DNA data storage based on Perl script. For this approach, the text data of the preamble of the “Universal Declaration of Human Rights” consisting of 2,046 words was encoded into the corresponding 8,148 base pairs of DNA using Perl-based encoding with a hash table. The encoded DNA sequences were then artificially synthesized for storage. The information DNA consisted of a total of 22 chemically synthesized DNA fragments with 400 nucleotides each, which were inserted into a cloning vector to multiply the plasmid DNA. The nucleotide integrity of the data-carrying DNA sequences were ensured under the accelerated aging conditions. Also, an erroneous nucleotide in the information DNA sequences was successfully corrected using the overlap extension PCR method. The stored DNA was read by sequencing, and the resulting DNA sequence information was successfully decoded to convert the DNA records back to the original document. Our results indicate that textual data can be stored in DNA using a simple, easy, and flexible Perl by running a script from the command line.
Similar content being viewed by others
References
Shrivastava, S. and R. Badlani (2014) Data storage in DNA. Int. J. Electr. Energy. 2: 119–124.
Hakami, H. A., Z. Chaczko, and A. Kale (2015) Review of big data storage based on DNA computing. Proceedings of the 2015 Asia-Pacific Conference on Computer Aided System Engineering. July 14–16. Washington, DC, USA.
Choi, Y., T. Ryu, A. C. Lee, H. Choi, H. Lee, J. Park, S. H. Song, S. Kim, H. Kim, W. Park, and S. Kwon (2019) High information capacity DNA-based data storage with augmented encoding characters using degenerate bases. Sci. Rep. 9: 6582.
Ceze, L., J. Nivala, and K. Strauss (2019) Molecular digital data storage using DNA. Nat. Rev. Genet. 20: 456–466.
Yazdi, S. M. H. T., H. M. Kiah, E. Garcia-Ruiz, J. Ma, H. Zhao, and O. Milenkovic (2015) DNA-based storage: Trends and methods. IEEE Trans. Mol. Biol. Multiscale Commun. 1: 230–248.
Panda, D., K. A. Molla, M. J. Baig, A. Swain, D. Behera, and M. Dash (2018) DNA as a digital information storage device: hope or hype? 3 Biotech. 8: 239.
Church, G. M., Y. Gao, and S. Kosuri (2012) Next-generation digital information storage in DNA. Science. 337: 1628.
Meyer, M., Q. Fu, A. Aximu-Petri, I. Glocke, B. Nickel, J. L. Arsuaga, I. Martinez, A. Gracia, J. M. B. de Castro, E. Carbonell, and S. Paabo (2014) A mitochondrial genome sequence of a hominin from Sima de los Huesos. Nature. 505: 403–406.
Meyer, M., J. L. Arsuaga, C. de Filippo, S. Nagel, A. Aximu-Petri, B. Nickel, I. Martinez, A. Gracia, J. M. B. de Castro, E. Carbonell, B. Viola, J. Kelso, K. Prüfer, and S. Paabo (2016) Nuclear DNA sequences from the Middle Pleistocene Sima de los Huesos hominins. Nature. 531: 504–507.
Wong, P. C., K. Wong, and H. Foote (2003) Organic data memory using the DNA approach. Commun. ACM. 46: 95–98.
Clelland, C. T., V. Risca, and C. Bancroft (1999) Hiding messages in DNA microdots. Nature. 399: 533–534.
Arita, M. and Y. Ohashi (2004) Secret signatures inside genomic DNA. Biotechnol. Prog. 20: 1605–1607.
Goldman, N., P. Bertone, S. Chen, C. Dessimoz, E. M. Le Proust, B. Sipos, and E. Birney (2013) Towards practical, high-capacity, low-maintenance information storage in synthesized DNA. Nature. 494: 77–80.
Grass, R. N., R. Heckel, M. Puddu, D. Paunescu, and W. J. Stark (2015) Robust chemical preservation of digital information on DNA in silica with error-correcting codes. Angew. Chem. Int. Ed. Engl. 54: 2552–2555.
Erlich, Y. and D. Zielinski (2017) DNA Fountain enables a robust and efficient storage architecture. Science. 355: 950–954.
Farzadfard, F. and T. K. Lu (2014) Genomically encoded analog memory with precise in vivo DNA writing in living cell populations. Science. 346: 1256272.
Shipman, S. L., J. Nivala, J. D. Macklis, and G. M. Church (2017) CRISPR-Cas encoding of a digital movie into the genomes of a population of living bacteria. Nature. 547: 345–349.
Nguyen, H. H., J. Park, S. J. Park, C. S. Lee, S. Hwang, Y. B. Shin, T. H. Ha, and M. Kim (2018) Long-term stability and integrity of plasmid-based DNA data storage. Polymers (Basel). 10: 28.
Nguyen, H. H., J. Park, S. Hwang, O. S. Kwon, C. S. Lee, Y. B. Shin, T. H. Ha, and M. Kim (2018) On-chip fluorescence switching system for constructing a rewritable random access data storage device. Sci. Rep. 8: 337.
Takahashi, C. N., B. H. Nguyen, K. Strauss, and L. Ceze (2019) Demonstration of end-to-end automation of DNA data storage. Sci. Rep. 9: 4998.
Fosdick, H. (2005) Programming languages for library and textual processing. Bul. Am. Soc. Info. Sci. Tech. 31: 21–26.
Rice, P. (2002) Beginning Perl for bioinformatics: An introduction to Perl for biologists. Brief Bioinform. 3: 210–212.
Baiocchi, G. (2004) Using Perl for statistics: Data processing and statistical computing. J. Stat. Softw. 11: i01.
Dung, T. T., Y. Oh, S. J. Choi, I. D. Kim, M. K. Oh, and M. Kim (2018) Applications and advances in bioelectronic noses for odour sensing. Sensors (Basel). 18: 103.
Nguyen, H. H., S. H. Lee, U. J. Lee, C. D. Fermin, and M. Kim (2019) Immobilized enzymes in biosensor applications. Materials (Basel). 12: 121.
Bryksin, A. V. and I. Matsumura (2010) Overlap extension PCR cloning: a simple and reliable way to create recombinant plasmids. Biotechniques. 48: 463–465.
Limbachiya, D., V. Dhameliya, M. Khakhar, and M. K. Gupta (2016) On optimal family of codes for archival DNA storage. arXiv. arXiv:1501.07133.
Yim, A. K. Y., A. C. S. Yu, J. W. Li, A. I. C. Wong, J. F. C. Loo, K. M. Chan, S. K. Kong, K. Y. Yip, and T. F. Chan (2014) The essential component in DNA-based information storage system: robust error-tolerating module. Front. Bioeng. Biotechnol. 2: 49.
Hebsgaard, M. B., M. J. Phillips, and E. Willerslev (2005) Geologically ancient DNA: fact or artefact? Trends Microbiol. 13: 212–220.
Acknowledgements
This work was supported by the Basic Science Research Program of the NRF funded by MSIT CNRF-2019R1A2C 1010149), and the Korea Research Institute of Bioscience and Biotechnology (KRBB) Initiative Research Program.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
The authors declare no conflict of interest.
Neither ethical approval nor informed consent was required for this study.
Additional information
Publisher’s Note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Lee, U.J., Hwang, S., Kim, K.E. et al. DNA Data Storage in Perl. Biotechnol Bioproc E 25, 607–615 (2020). https://doi.org/10.1007/s12257-020-0022-9
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12257-020-0022-9