Abstract
Key message
When Populus wutunensis seedlings are salt stressed, many genes involved in cellular metabolism, signal transduction, transport, osmotic regulation, and reactive oxygen species scavenging show elevated expression levels.
Abstract
Populus wutunensis, a natural hybrid individual resulting from a cross between Populus × xiaozhuanica W. Y. Hsu et Liang, shows salt and drought tolerance and is fast-growing. It is widely cultivated in the Three-North Shelterbelt Forest area in China. To determine the transcriptome changes in P. wutunensis under salt stress, RNA-seq was conducted in salt-treated leaves and non-salt-treated control leaves. A total of 81,188 unigenes, with an average length of 982 bp, were obtained. In salt-treated leaves, 2466 differentially expressed genes were identified that differed from those in control leaves at any time point. Among them, 391 were upregulated and 44 were downregulated during all salt stress time points. They were mainly involved in cellular and metabolic processes, signal transduction, transport, osmotic regulation, and reactive oxygen species (ROS) scavenging. In addition, 2543 transcription factor genes were identified and divided into 60 families. qRT-PCR analysis revealed that the levels of differentially expressed genes were consistent with those from sequencing of the transcriptome under salt stress. These findings will increase our understanding of gene regulation in response to salt stress in P. wutunensis and are beneficial for future research on molecular-assisted breeding.
Similar content being viewed by others
Introduction
Soil salinization has become an important challenge in managing global resources and conserving the environment as it reduces productivity (Nawar et al. 2015; Zhu et al. 2016). Areas supporting major crops, affected by salinity, alkalinity, and extreme environments worldwide, are > 50% annually. If saline-alkali lands can be improved and utilized, they can provide significantly more ecological and economic value to society. The mechanisms of plant responses to salt stress have been extensively studied with respect to reactive oxygen species (ROS) scavenging (Yan et al. 2018), hormone balance regulation (Wang et al. 2018a, b), and substance synthesis and accumulation (Guanet al. 2018). Many transcription factors are known to regulate the expression levels of various genes that may influence the level of salt tolerance in plants (Deinlein et al. 2014). However, regardless of these studies, further research is necessary to develop an integrated and comprehensive understanding of salt-stress mechanisms in plants.
Populus wutunensis is a natural hybrid poplar resulting from a cross between Populus × xiaozhuanica W. Y. Hsu et Liang. It grows fast and is highly tolerant of salinity and drought (Jia et al. 2009). After many years of breeding, comparative experiments between potted plants and field grown ones, and evaluation by experts, P.wutunensis was characterized as a species that can resist drought stress, barren stress, and salinity stress. Owing to its high productivity and fast growth, it can be widely used to create timber in farmland shelterbelts, coastal areas, and inland saline land (Jia et al. 2009; Ma et al. 2013). Gene expression profiling has been employed to try to achieve a comprehensive understanding of the genetic mechanisms that control stress responses in plants (Golldack et al. 2011). Salt-tolerant plants can be used to study the changes in gene expression under different salt treatments and to identify potential salt adaptation-related genes (Wang et al. 2015). Research on P. wutunensis is still rare, and the molecular mechanisms underlying its resistance to salt and alkali remain unclear.
In this study, we used the Illumina HiSeqXTen sequencing platform to perform deep transcriptome sequencing to compare the genome-wide expression profile between salt treatment and normal growing conditions. Validation of the current salt-tolerance genes may contribute to future salt-resistance research in woody plants. This study may provide a starting point for elucidation of the molecular mechanism underlying the salt stress response in P. wutunensis.
Materials and methods
Plant growth and salt stress treatments
Populus wutunensis plants were obtained from the Fuxin forestry bureau and cultured in plastic pots in the laboratory. For the salt treatment, 1-year-old seedlings were kept at a mean temperature of 25.0 ± 2 °C and relative humidity of 60 ± 5%. They were divided into four groups, and a set of leaves was collected every 4 days. They were watered every 4 days with 200 mmol/l of NaCl-containing Hoagland’s nutrient solution in three replicates. Three more were administered with Hoagland nutrient solution only, as controls. For RNA-Seq analysis, P. wutunensis seedlings without 200 mmol/l NaCl treatment were used as the control plants, and those treated with 200 mmol/l NaCl were considered the treatment group. The second to fifth leaves of these poplars were collected and immediately frozen in liquid nitrogen. The four groups of collected samples were CK 0d (T1), 200 mM NaCl treatment for 4d (T2), 200 mM NaCl treatment for 8d (T3), and 200 mM NaCl treatment for 12d (T4).
RNA extraction, cDNA library construction, and Illumina sequencing
Total RNA from the four leaf samples, with and without NaCl treatment (T1, T2, T3, and T4), was isolated using the RNAprep Pure Plant Kit (Sangon Biotech, Shanghai, China) according to the manufacturer's protocol. Using the Agilent 2100 Bioanalyzer (Agilent RNA 6000 Nano Kit) for the total RNA sample QC, RNA concentration, RIN value, 28S/18S, and fragment length distribution were estimated. Construction of libraries and RNA-seq assays for paired-end transcriptome sequencing were performed by BGI (Shenzhen, China). mRNA was isolated from total RNA using the oligo(dT) method and fragmented under different conditions. The first and second strands of cDNA were synthesized. cDNA fragments were purified and resolved using EB buffer for end repair and single nucleotide A (adenine) addition. Thereafter, the cDNA fragments were connected to adapters. cDNA fragments of suitable size were selected for PCR amplification. An Agilent 2100 Bioanaylzer and an ABI Step One Plus Real-Time PCR System were used for quantitative and qualitative analyses of the libraries. Finally, these libraries were sequenced using Illumina HiSeqX Ten.
De-novo transcriptome assembly and functional annotation
We defined raw reads as those of low quality, adapter-polluted, and with a high content of unknown bases (N); these noise reads were removed before further analyses. We used Trinity to perform de novo assembly with clean reads (PCR duplication was removed to improve the efficiency) and utilized TGICL to cluster transcripts in Unigene. We used Blastn, Blastx, or Diamond to align the unigenes to NT, NR, KOG, KEGG, and Swiss Prot databases to complete the annotation. We selected Blast2 GO with NR annotation for GO annotation and used Inter Pro Scan5 for Inter Pro annotation.
Analysis of differentially expressed genes (DEGs)
We used the DESeq2 algorithm to detect differential genes. Differential genes were screened by double or higher multiples, and the corrected P value was ≤ 0.05. The pheatmap function in R software was used for hierarchical cluster analysis. DEGs were further classified by the GO function and KEGG biological pathway.
Validation of RNA-Seq with quantitative real-time PCR (qRT-PCR)
To verify the accuracy of the RNA-seq data, nine unigenes were randomly selected for the qRT-PCR test. Total RNA was extracted with TRIzol (Sangon, Shanghai, China) from the same leaves used in transcriptome analysis, as per the manufacturer’s protocol. First-strand cDNA was synthesized using the PrimeScript™ II First Strand cDNA synthesis kit (Takara Bio, Dalian, China) according to the manufacturer's protocol. Primer Premier 5.0 was used to design the primers required for qPCR, and actin20 was used as the reference gene (Table S1). qRT-PCR was performed using SYBR Premix Ex Taq™ II kit (TaKaRa) and run on an ABI 7500 Real-Time PCR System (Thermo Fisher Scientific, Inc., Waltham, MA, USA). Each reaction was repeated thrice, and the expression levels were calculated using the 2−∆∆Ct method. Data were analyzed and plotted using Microsoft Excel 2007.
Results and discussion
Sequencing statistics
To compare the transcriptomes of P. wutunensis treated with 200 mmol/l NaCl, a de-novo assembly of the species was generated. In total, 66.40 Mb of raw reads was generated from the mixed samples (Table 1). After filtration, 58.87 Mb of clean reads was obtained. More than 98.85% of bases in clean reads had a Q value ≥ 20. The GC contents were 41.88% and 41.73% for raw reads and clean reads, respectively (Table 2). Clean read data were used for de-novo assembly. Trinity software generated 135,837 total transcripts with an average length of 770 bp and a N50 of 1451 bp; 81,188 sequences were identified as unigenes. The four libraries (T1, T2, T3, and T4) produced > 4 GB of raw data (Table 3).
Functional annotation
All assembled unigenes were aligned, annotated, and classified with NR, NT, Swissprot, KEGG, KOG, GO, and Interpro databases (Table 4), and the annotated results were 56,809 (NR: 69.97%), 67,620 (NT: 83.29%), 36,168 (Swissprot: 44.55%), 40,144 (KEGG: 49.45%), 41,819 (KOG: 51.51%), 18,519 (GO: 22.81%), and 44,767 (Interpro: 55.14%). In the NR database, the highest homology with P. wutunensis was 58.85%, followed by that of the European and American poplar, which was 26.59%. Triticum urartu and Aegilopstaus chli had the lowest homology, 1.2% and 1.68%, respectively, and other species accounted for 11.68%. (Fig. 1).
The unigenes that were aligned with the NR database were annotated with the GO database; we calculated the distribution in three categories: biological processes, cellular components, and molecular functions (Fig. 2). There were 23 subcategories in the biological processes category, of which the metabolic processes subcategory had the highest proportion, followed by the cellular processes and single-organism processes. The metabolic processes, cellular processes, and single-organism processes had 9901, 8941, and 6436 unigenes, respectively. There were 17 subcategories in the cellular components’ category, with the largest proportion in cellular components, all with 8874 unigenes. In the 14 subcategories of the molecular functions’ category, binding accounted for the highest proportion, followed by catalytic activity, with 9921 and 8881 unigenes, respectively.
Through the KOG database comparison and analysis, 41,879 unigenes were divided into 25 orthologous types (Fig. 3). Among them, 10,260 were general function predictive genes accounting for 24.50%; 6146 were signal transduction mechanism genes accounting for 14.68%; 4785 corresponded to post-translational modifications, protein turnover, and chaperone genes accounting for 11.43%. There were 58 cell movement genes, the proportion of which was at least 0.14%.
To identify the biological pathways of the salt stress response, we analyzed the KEGG metabolic pathway of the assembled unigenes. The 40,144 unigenes could be divided into five major metabolic pathways, namely, cell processes (1767), environmental information processing (8362), genetic information processing (132), metabolism (21,264), and biological systems (1625), and sub-divided into 21 sub-metabolic pathways, most of which were global and overview maps (8194), glycan biosynthesis and metabolism (3138), translation (3112), folding, sorting, and degradation (2675), signal transduction (2235), transport and catabolism (1767), transcription (1750), amino acid metabolism (1739), lipid metabolism (1653), environmental adaptation (1625), and other sub-categories (Fig. 4).
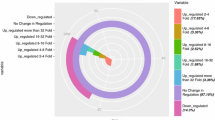
To study the differential expression of genes in P. wutunensis under salt stress, changes in gene expression in samples treated with salt stress at different treatment time (T2, T3, and T4) with respect to the control (T1) were compared. Most of the genes were found to have higher expression levels (Fig. S1 and Table S2). Differentially expressed genes were screened based on the criteria that their expression levels more than doubled and P < 0.05. Gene expression was calculated according to the FPKM value. A total of 1652 genes were found to be upregulated and 814 downregulated during salt stress. Among them, 391 genes were upregulated and 44 downregulated during salt stress (Figs. 5, 6).
Functional classification of DEGs
To further elucidate the functions of the DEGs, GO enrichment analysis was carried out. The DEGs were similarly annotated according to biological processes, cellular components, and molecular functions (Table S3). They were annotated into 166 KEGG pathways. KEGG analysis showed the differential genes in different samples were significantly enriched in seven pathways (P < 0.05), including plant hormone signal transduction, plant circadian rhythms, plant MAPK signaling pathway, plant-pathogen interactions, alpha-linolenic acid metabolism, starch and sucrose metabolism, and anthocyanin biosynthesis (Table S4).
Verification of RNA-seq data by qRT-PCR
To test the reliability of RNA-seq data, the following nine candidate genes with different transcriptional trends were randomly selected for qRT-PCR determination:E3 ubiquitin-protein (RING1), flavanol synthase/flavanone 3-hydroxylase-like (F3H), calcium-binding protein (CML3), peptidyl-prolyl cis–trans isomerase (FKBP19), leucine-rich repeat receptor-like tyrosine-protein kinase (PXC3), serine/threonine-protein kinase (CTR1), WRKY transcription factor 32 (WRKY32), WRKY transcription factor 51(WRKY51), and BTB/POZ domain-containing protein (BTBD1), etc. All the selected functional and regulatory proteins were related to the salt stress response and signaling (Yu et al. 2013). Expression of the selected genes under different salt stress durations was measured by qRT-PCR. The results were similar to those of RNA-seq (Fig. 7, Fig. S2). Although there were some differences in the multiples of unigene expression between RNA-seq data and qRT-PCR, the trend of most gene expression patterns, detected under salt stress at different times, was consistent. Therefore, expression of the selected genes measured by qRT-PCR was consistent with the results of transcriptome sequencing.
Identification of key DEGs in response to salt stress
Under abiotic stress, plants adapt to the environment by regulating a series of complex gene networks. Plants have many mechanisms to adapt to the environment under salt stress, for example, ion transport, ROS scavenging, osmotic regulation, and cell signal conduction (Wang et al. 2018a, b). In this study, the P. wutunensis transcriptome sequencing data were found to be related to the genes involved in the above pathways. By comparing the DEGs of salt-stressed and control samples (T2/T1, T3/T1, and T4/T1, respectively), those that were continuously high throughout the salt stress were identified (Table 5). Change in gene expression was different at different time points of salt stress treatment, which indicated that the mechanism of response of P. wutunensis to salt stress was complex.
Growth and development of higher plants and adversity stress processes are regulated by a variety of signal transduction systems (Liu et al. 2019). The xylan transferase gene regulates cell wall elasticity in response to adversity stress (Dong et al. 2011). In this study, three auxin-coding proteins and one xylan transferase gene were identified as being upregulated under salt stress. The ABA-regulated ABA-signaling pathway responded to adversity by activating SnRK2 protein kinase through the interaction of its receptor PYL protein family with PP2C (Zhao et al. 2018). In this study, we identified one continuously downregulated ABA receptor PYL gene and two continuously upregulated PP2C genes.
Through the synergistic action of ion translocators on the surface of membrane systems, plants accomplish the excretion and intracellular segregation of excess salt ions and reconstruct and maintain ion homeostasis under salt stress. Transporters, embedded in the plasma membrane, cytoplasm, or nucleus, are associated with various signaling pathways (Julkowska and Testerink, 2015). In this study, six translocator genes were identified: five were upregulated and one was downregulated, including the encoding translocator, calcium ion translocator, and potassium ion translocator genes.
When plants grow in a saline environment, excessive salt ions reduce the osmotic potential of soil, thus challenging the plants to either absorb water or discard water, further causing osmotic stress (Yan et al. 2013). Osmotic regulation is an important response of plants to stress. Plants can regulate intra- and extracellular ion concentrations through ion pumps on cell membranes and accumulate small molecules such as sugars for osmotic pressure regulation (Liet al. 2018). To combat the osmotic stress caused by higher concentrations of Na+ in the vacuoles, plants accumulate compatible solutes in the cytoplasm, such as betaine, proline, free sugar, and polyalcohol. In this study, five consecutively expressed genes were identified, including two involved in ion transport and three involved in glycometabolism. Among them, two glycosylhydrolase genes participate in the glucose synthesis pathway in response to salt stress through the up- and downregulation of two different expression modes.
High concentrations of salt can not only induce osmotic stress in plants, but can also accumulate excessive ROS and generate oxidative stress, which can further inhibit the normal metabolism of cells, eventually leading to oxidative damage and even death (Suzuki et al. 2012; Gill and Tuteja 2010). The oxidative stress response genes are used to ensure the timely removal of intracellular ROS, thereby reducing oxidative damage under salt stress (Li et al. 2018). In this study, two ROS-scavenging genes that were continuously expressed under salt stress were identified.
Identification of TFs
Transcription factors are a group of important regulatory proteins that bind to specific nucleotide sequences upstream of genes and activate or inhibit the expression of target genes. They play a very important role in the process of adversity signal transduction (Li et al. 2015). In this study, 2543 transcription factor genes were identified and divided into 60 families (Table S5), all of which responded to salt stress in different ways. Members of the transcription factor family such as bZIP, WRKY, AP2/EREBP, C2H2, bHLH, MYB, NAC, and C2H2-Dofandthese were involved in the plant response to salt stress (Yu et al. 2013). In this study, transcription factors from AP2-EREBP, bZIP, WRKY, NAC, and MYB families were continuously expressed under salt stress (Table 6). Among them, WRKY was continuously upregulated under salt stress with the highest expression, suggesting that the WRKY transcription factor may play a major regulatory role in response to salt stress in P. wutunensis.
Key gene regulatory networks under salt-stress conditions
Salt stress usually induces ionic, osmotic, and oxidative stress in plants. The high Na+ levels under salt stress generate cytoplasmic Ca2+ signals, and the CBL-CIPK system of calcium ion receptors plays an important role in regulating the opening and closing of potassium channels (Li et al. 2006). In this study, we found that Ca2+ signals activate the CBL-CIPK complex under salt-stress conditions, and the phosphorylated CBL-CIPK complex regulates K+ channels to achieve ionic homeostasis. The CBL-CIPK interaction activates RBOHF, which acts on intracellular H2O2 (Drerup et al. 2013; Sun et al. 2015), thereby upregulating ROS-scavenging enzymes, which scavenge ROS. The PYL-mediated ABA signaling pathway plays an important role in plant responses to abiotic stress. Salt stress leads to increases in ABA content, which activates SnRK2 by inhibiting PP2C via its receptor (PYL) (Zhu et al. 2016). SnRK2 activates the mitogen-activated protein kinase (MAPK) to regulate transcription factors and induce the expression of relevant salt tolerance genes that confer resistance to salt stress in plants. Previous studies have found that direct binding between the PYL8 receptor and MYB77 regulates the growth of lateral roots via the auxin signaling pathway (Zhao et al. 2014), which may be responsible for the high expression of auxin-related genes after being exposed to salt stress (Fig. 8).
Conclusion
This study has reported for the first time the transcriptome data of P. wutunensis in response to salt stress. We identified a large number of DEGs that may play an important role in salt stress. Sustained high expression of genes under salt stress mainly involved signal transduction, transport, osmotic regulation, and ROS scavenging. Our results will facilitate the analysis of gene expression profiles related to salt tolerance in plants and serve as a valuable resource for future functional genomics and studies of molecular breeding.
Author contributions
JH designed the experiments. ZJ performed the experiments, data analysis, and drafted the manuscript. All authors read and approved the final manuscript.
Abbreviations
- ROS:
-
Reactive oxygen species
- qRT-PCR:
-
Quantitative reverse transcription polymerase chain reaction
- DEGs:
-
Differentially expressed genes
References
Deinlein U, Stephan AB, Horie T, Luo W, Xu G, Schroeder JI (2014) Plant salt-tolerance mechanisms. Trends Plant Sci 19(6):371–379
Dong J, Jiang YY, Chen RJ, Xu ZJ, Gao XL (2011) Isolation of a novel xyloglucan endotransgucosylase (OsXET9) gene from rice and analysis of the response of this gene to abiotic stresses. Afr J Biotechnol 10(76):17424–17434
Drerup MM, Schlücking K, Hashimoto K, Manishankar P, Steinhorst L, Kuchitsu K, Kudla J (2013) The cal-cineurin B-like calcium sensors CBL1 and CBL9 together with their interacting protein kinase CIPK26 regulate the Arabidopsis NADPH oxidase RBOHIJ. Mol Plant 6(2):559–569
Gill SS, Tuteja N (2010) Reactive oxygen species and antioxidant machinery in abiotic stress tolerance in crop plants. Plant Physiol Biochem 48(12):909–930
Golldack D, Luking I, Yang O (2011) Plant tolerance to drought and salinity: stress regulating transcription factors and their functional significance in the cellular transcriptional network. Plant Cell Rep 30(8):1383–1391
Guan PY, Wang J, Li H, Xie C, Zhang SZ, Wu CG, Yang GD, Yan K, Huang JG, Zheng CC (2018) Sensitive to SALT1, an endoplasmic reticulum-localized chaperone positively regulates salt resistance. Plant Physiol 178:1390–1405
Jia B, Wang W, Zhao Y, Lu X, Jiang G (2009) Growth of fast-growing Populus wutunensis with high saline-alkali tolerance. J Northeast For Univ 37(6):4–6
Julkowska MM, Testerink C (2015) Tuning plant signaling and growth to survive salt. Trends Plant Sci 20:586–594
Li L, Kim BG, Cheong YH, PandeyGK LS (2006) Ca2+ signaling pathway regulated a K+ channel for low-K response in Arabidopsis. Proc Natl Acad Sci USA 103:12625–12630
Li JB, Luan YS, Liu Z (2015) Overexpression of SpWRKY1promotes resistance to Phytophthoranicotianae and tolerance to salt and drought stress in transgenic tobacco. Physiol Plant 155(3):248–266
Li F, Zhang H, Zhao H, Gao T, Song A, Jiang J, Chen F, Chen S (2018) Chrysanthemum CmHSFA4 gene positively regulates salt stress tolerance in transgenic chrysanthemum. Plant Biotech J16(7):1311–1321
Liu A, Xiao Z, Li MW, Wong FL, Yung WS, Ku YS, Wang Q, Wang X, Xie M, Yim AK, Chan TF, Lam HM (2019) Transcriptomic reprogramming in soybean seedlings under salt stress. Plant Cell Environ 42(1):98–114
Ma JL, Jiang GB, Yao SJ, Jin H (2013) Effects of apoplastic ion contents in the tender stems of two populus cultivars on gas exchange parameters under salt stress. Sci Silv 49(7):40–47
Nawar S, Buddenbaum H, Hill J (2015) Estimation of soil salinity using three quantitative methods based on visible and near-infrared reflectance spectroscopy: a casestudy from Egypt. Arab J Geosci 8(7):5127–5140
Sun T, Wang Y, Wang M, Li TT, Zhou Y, Wang XT, Wei SY, He GY, Yang GX (2015) Identification and comprehensive analyses of the CBL and CIPK gene families in wheat (Triticum aestivum L.). BMC Plant Biol 15:260–269
Suzuki N, Koussevitzky S, Mittler R, Mittler G (2012) ROS and redox signalling in the response of plants to abiotic stress. Plant Cell Environ 35(2):259–270
Wang JC, Li BC, Meng YX, Ma XL, Lai Y, Si EJ, Ren PR, Shang XW, Wang JH (2015) Transcriptomic profiling of the salt-stress response in the halophyte Halogeton glomeratus. BMC genomics 16(1):169–183
Wang L, Du HY, Li TZ, Wuyun T (2018a) De novo transcriptome sequencing and identification of genes related to salt stress in Eucommiaulmoides oliver. Trees 32:151–163
Wang Z, Wang FX, Hong YC, Yao JJ, Ren ZZ, Shi HZ, Zhu JK (2018b) The flowering repressor SVP confers drought resistance in Arabidopsis by regulating abscisic acid catabolism. Mol Plant 11:1184–1197
Yan K, Shao H, Shao C, Chen P, Zhao SJ, Brestic M, Chen XB (2013) Physiological adaptive mechanisms of plants grown in saline soil and implications for sustainable saline agriculture in coastal zone. Acta Physiologiae Plant 35(10):2867–2878
Yan YY, Wang SS, Wei M, Gong H (2018) Effect of different rootstocks on the salt stress tolerance in watermelon seedlings. Horticu Plant J 4(6):239–249
Yu CG, Xu S, Yin YL (2013) Transcriptome analysis of the Taxodium ‘Zhongshanshan 405’ roots in response to salinity stress. Plant Physiol Biochem 100:156–165
Zhao Y, Xing L, Wang X, Hou YJ, Gao J, Wang P, Duan CG, Zhu X, Zhu JK (2014) The ABA receptor PYL8 promotes lateral root growth by enhancing MYB77-dependent transcription of auxin-responsive genes. Sci Signal 7(328):ra53
Zhao Y, Zhang ZJ,Gao JH, Wang PC, Hu T, Wang ZG, Hou Y J, Wan YZ, Liu WS, XieSJ, Lu TJ, Xue L,Liu YJ,Macho AP, Tao WA, Bressan RA, Zhu JK (2018) Arabidopsis duodecuple mutant of PYL ABA receptors reveals PYL repression of ABA-independent SnRK2 activity. Cell Rep 23(11):3340–3351
Zhu JK (2016) Abiotic stress signaling and responses in plants. Cell 167(2):313–324
Acknowledgements
This work was supported by the Central Finance Forestry Science and Technology Promotion Demonstration Fund Project (Liao [2019] TG01) and the Fundamental Research Funds for the Central Universities(DC201501070103).
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by B. Wang.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Zou, J., Jin, H. Transcriptome sequencing and gene expression profiling of Populus wutunensis, a natural hybrid, during salinity stress. Trees 34, 1427–1438 (2020). https://doi.org/10.1007/s00468-020-02014-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00468-020-02014-6