Abstract
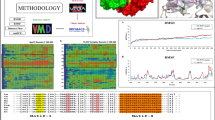
Leishmaniasis is a tropical neglected disease that imposes major health concerns in many endemic countries worldwide and requires urgent attention to the identification of new drug targets as well as drug candidates. In the current study, we propose homoserine kinase (HSK) inhibition as a strategy to induce pathogen mortality via generating threonine deficiency. We introduce a homology-based molecular model of leishmanial HSK that appears to possess all conserved structural as well as functional features in the GHMP kinase family. Furthermore, 200 ns molecular dynamics data of the enzyme in open and closed state attempts to provide the mechanistic details involved in the substrate as well as phosphate binding to this enzyme. We discuss the structural and functional significance of movements involved in various loops (motif 1, 2, 3) and lips (upper and lower) in the transition of leishmanial HSK from closed to open state. Virtual screening data of more than 40,000 compounds from the present investigation tries to identify a few potential HSK inhibitors that possess important features to act as efficient HSK inhibitors. These compounds can be considered an effective starting point for the identification of novel drug-like scaffolds. We hope the structural wealth that is offered in this report will be utilized in designing competent experimental and therapeutic interventions for leishmaniasis management.

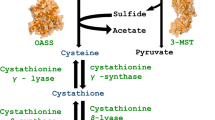
Graphical abstract








Similar content being viewed by others
References
Akhoundi M, Kuhls K, Cannet A, Votýpka J, Marty P, Delaunay P &Sereno D. (2016) A historical overview of the classification, evolution, and dispersion of Leishmania parasites and sandflies. PLoS Negl Trop Dis 10(3):e0004349. https://doi.org/10.1371/journal.pntd.0004349
Sosa N, Pascale JM, Jiménez AI, Norwood JA, Kreishman-Detrick M, Weina PJ, Lawrence K, McCarthy WF, Adams RC, Scott C, Ransom J, Tang D, Grogl M (2019) Topical paromomycin for New World cutaneous leishmaniasis. PLoS Negl Trop Dis 13(5):e0007253
Alcântara LM, Ferreira TCS, Gadelha FR, Miguel DC (2018) Challenges in drug discovery targeting TriTryp diseases with an emphasis on leishmaniasis. Int J Parasitol Drugs Drug Resist 8(3):430–439
Montoya A, López MC, Vélez ID, Robledo SM (2019) Label-free quantitative proteomic analysis reveals potential biomarkers for early healing in cutaneous leishmaniasis. PeerJ 6:e6228
Borba JVB, Silva AC, Ramos PIP, Grazzia N, Miguel DC, Muratov EN, Furnham N, Andrade CH (2019) Unveiling the kinomes of Leishmaniainfantum and L. braziliensis empowers the discovery of new kinase targets and antileishmanial compounds. Comput Struct Biotechnol J 17:352–361
Basher A, Rashid MM, Habibullah AM, Nath R, Akter D, Chowdhury IH, Azim A, Nath P, Faiz MA (2019) Miltefosine induced reduced male fertility capacity after treatment of post kala-azar dermal leishmaniasis, Bangladesh. Mymensingh Med J 28(2):328–332
Burza S, Croft SL, Boelaert M (2018) Leishmaniasis. Lancet 392(10151):951–970
Tiwari N, Gedda MR, Tiwari VK, Singh SP, Singh RK (2018) Limitations of current therapeutic options, possible drug targets and scope of natural products in control of leishmaniasis. Mini-Rev Med Chem 18(1):26–41
Chakravarty J, Sundar S (2010) Drug resistance in leishmaniasis. J Global Infect Dis 2(2):167–176
Veronica J, Chandrasekaran S, Dayakar A, Devender M, Prajapati VK, Sundar S, Maurya R (2019) Iron superoxide dismutase contributes to miltefosine resistance in Leishmaniadonovani. FEBS J. https://doi.org/10.1111/febs.14923
Chakravarty J, Sundar S (2019) Current and emerging medications for the treatment of leishmaniasis. Expert Opin Pharmacother 7:1–15
Alexandrino-Junior F, Silva KGHE, Freire MCLC, Lione VOF, Cardoso EA, Marcelino HR, Genre J, Oliveira AG, Egito ESTD (2019) A functional wound dressing as a potential treatment for cutaneous leishmaniasis. Pharmaceutics 11(5):E200
Bekhit AA, El-Agroudy E, Helmy A, Ibrahim TM, Shavandi A, Bekhit AEA (2018) Leishmania treatment and prevention: natural and synthesized drugs. Eur J Med Chem 160:229–244
Ong YC, Kedzierski L, Andrews PC (2018) Do bismuth complexes hold promise as antileishmanial drugs? Future Med Chem 10(14):1721–1733
Jain V, Jain K (2018) Molecular targets and pathways for the treatment of visceral leishmaniasis. Drug Discov Today 23(1):161–170. https://doi.org/10.1016/j.drudis.2017.09.006
Meshram RJ, Goundge MB, Kolte BS, Gacche RN (2019) An in-silico approach in identification of drug targets in Leishmania: a subtractive genomic and metabolic simulation analysis. Parasitol Int 69:59–70
Ong HB, Lee WS, Patterson S, Wyllie S, Fairlamb AH (2015) Homoserine and quorum-sensing acyl homoserine lactones as alternative sources of threonine: a potential role for homoserine kinase in insect-stage Trypanosoma brucei. Mol Microbiol 95(1):143–156
Zhou T, Daugherty M, Grishin NV, Osterman AL, Zhang H (2000) Structure and mechanism of homoserine kinase: prototype for the GHMP kinase superfamily. Structure 8(12):1247–1257
Hospital A, Goñi JR, Orozco M, Gelpí JL (2015) Molecular dynamics simulations: advances and applications. Adv Appl Bioinforma Chem 8:37–47
Karplus M, Kuriyan (2005) Molecular dynamics and protein function. Proc Natl Acad Sci U S A 102(19):6679–6685
Boeckmann B, Bairoch A, Apweiler R, Blatter MC, Estreicher A, Gasteiger E, Martin MJ, Michoud K, O’Donovan C, Phan I, Pilbout S, Schneider M (2003) The SWISS-PROT protein knowledgebase and its supplement TrEMBL. Nucleic Acids Res 31:365–370
Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ (1990) Basic local alignment search tool. J Mol Biol 215(3):403–410
Meshram RJ, Gavhane A, Gaikar R, Bansode TS, Maskar A, Gupta A, Sohni S, Patidar M, Pandey T, Jangle S (2010) Sequence analysis and homology modeling of laccase from Pycnoporus cinnabarinus. Bioinformation 5(4):150–154
Sali A, Blundell TL (1993) Comparative protein modelling by satisfaction of spatial restraints. J Mol Biol 234(3):779–815
Needleman SB, Wunsch CD (1970) A general method applicable to the search for similarities in the amino acid sequence of two proteins. J Mol Biol 48(3):443–453
Guex N, Peitsch MC (1997) SWISS-MODEL and the Swiss-PdbViewer: an environment for comparative protein modeling. Electrophoresis 18(15):2714–2723
vanGunsteren WF, Brunne RM, Gros P, van Schaik RC, Schiffer CA, Torda AE (1994) Accounting for molecular mobility in structure determination based on nuclear magnetic resonance spectroscopic and X-ray diffraction data. Methods Enzymol 239:619–654
Hooft RW, Vriend G, Sander C, Abola EE (1996) Errors in protein structures. Nature 381(6580):272
Laskowski RA, MacAurther MW, Moss DS, Thornton JM (1993) PROCHECK-a program to check the stereochemical quality of proteins. J Appl Crystallogr 26:47–60
Castrignanò T, De Meo PD, Cozzetto D, Talamo IG, Tramontano A (2006) The PMDB protein model database. Nucleic Acids Res 34(Database issue):D306–D309
Phillips JC, Braun R, Wang W, Gumbart J, Tajkhorshid E, Villa E, Chipot C, Skeel RD, Kalé L, Schulten K (2005) Scalable molecular dynamics with NAMD. J Comput Chem 26(16):1781–1802
Meshram RJ, Bagul KT, Pawnikar SP, Barage SH, Kolte BS, Gacche RN (2019) Known compounds and new lessons: structural and electronic basis of flavonoid-based bioactivities. J Biomol Struct Dyn 21:1–17
Yesselman JD, Price DJ, Knight JL, Brooks CL (2012) MATCH: an atom-typing toolset for molecular mechanics force fields. J Comput Chem 33(2):189–202
Pettersen EF, Goddard TD, Huang CC, Couch GS, Greenblatt DM, Meng EC, Ferrin TE (2004) UCSF Chimera--a visualization system for exploratory research and analysis. J Comput Chem 25(13):1605–1612
Ryckaert JP, Ciccotti G, Berendsen JCH (1977) Numerical integration of the cartesian equations of motion of a system with constraints: molecular dynamics of n-alkanes. J Comput Phys 23(3):327–341
Darden T, York D, Pedersen L (1993) Particle mesh Ewald: an Nlog·(N) method for Ewald sums in large systems. J Chem Phys 98(12):10089–10092
Feller SE, Zhang Y, Pastor RW (1995) Constant pressure molecular dynamics simulation: the Langevin piston method. J Chem Phys 103(11):4613–4621
Martyna GJ, Tobias DJ, Klein ML (1994) Constant pressure molecular dynamics algorithms. J Chem Phys 101(5):4177–4189
Hoover WG (1985) Canonical dynamics: equilibrium phase-space distributions. Phys Rev A Gen Phys 31(3):1695–1697
Pronk S, Páll S, Schulz R, Larsson P, Bjelkmar P, Apostolov R, Shirts MR, Smith JC, Kasson PM, Van der Spoel D, Hess B, Lindahl E (2013) GROMACS 4.5: a highthroughput and highly parallel open source molecular simulation toolkit. Bioinformatics 29:845–854
Daura X, Gademann K, Jaun B, Seebach D, VanGunsteren WF, Mark AE (1999) Peptide folding: when simulation meets experiment. Angew Chem Int Ed 38(1–2):236–240
Knapp B, Lederer N, Omasits U, Schreiner W (2010) vmdICE: a plug-in for rapid evaluation of molecular dynamics simulations using VMD. J Comput Chem 31(16):2868–2873
Huey R, Morris GM, Olson AJ, Goodsell DS (2007) A semiempirical free energy force field with charge-based desolvation. J Comput Chem 28(6):1145–1152
Morris GM, Huey R, Lindstrom W, Sanner MF, Belew RK, Goodsell DS, Olson AJ (2009) AutoDock4 and AutoDockTools4: automated docking with selective receptor flexibility. J Comput Chem 30(16):2785–2791. https://doi.org/10.1002/jcc.21256
Ambhore AN, Kamble SS, Kadam SN, Kamble RD, Hebade MJ, Hese SV, Gaikwad MV, Meshram RJ, Gacche RN, Dawane BS (2019) Design, synthesis and in silico study of pyridine based 1,3,4-oxadiazole embedded hydrazinecarbothioamide derivatives as potent anti-tubercular agent. Comput Biol Chem 80:54–65
Patil KK, Meshram RJ, Barage SH, Gacche RN (2019) Dietary flavonoids inhibit the glycation of lens proteins: implications in the management of diabetic cataract. 3 Biotech 9(2):47
Kolte BS, Londhe SR, Solanki BR, Gacche RN, Meshram RJ (2018) FilTerBaSe: a web accessible chemical database for small compound libraries. J Mol Graph Model 80:95–103
Azizian H, Bahrami H, Pasalar P, Amanlou M (2010) Molecular modeling of Helicobacter pylori arginase and the inhibitor coordination interactions. J Mol Graph Model 28(7):626–635
Laskowski RA, Swindells MB (2011) LigPlot+: multiple ligand-protein interaction diagrams for drug discovery. J Chem Inf Model 51(10):2778–2786
Wallace AC, Laskowski RA, Thornton JM (1995) LIGPLOT: a program to generate schematic diagrams of protein-ligand interactions. Protein Eng 8(2):127–134
Willard L, Ranjan A, Zhang H, Monzavi H, Boyko RF, Sykes BD, Wishart DS (2003) VADAR: a web server for quantitative evaluation of protein structure quality. Nucleic Acids Res 31(13):3316–3319
Bork P, Sander C, Valencia A (1993) Convergent evolution of similar enzymatic function on different protein folds: the hexokinase, ribokinase, and galactokinase families of sugar kinases. Protein Sci 2(1):31–40
Krishna SS, Zhou T, Daugherty M, Osterman A, Zhang H (2002) Structural basis for the catalysis and substrate specificity of homoserine kinase. Biochemistry 40(36):10810–10818
Ban C, Yang W (1998) Crystal structure and ATPase activity of MutL: implications for DNA repair and mutagenesis. Cell 95(4):541–552
Ban C, Junop M, Yang W (1999) Transformation of MutL by ATP binding and hydrolysis: a switch in DNA mismatch repair. Cell 97(1):85–97
Murzin AG (1995) A ribosomal protein module in EF-G and DNA gyrase. Nat Struct Biol 2(1):25–26
Stams T, Niranjanakumari S, Fierke CA, Christianson DW (1998) Ribonuclease P protein structure: evolutionary origins in the translational apparatus. Science 280(5364):752–755
Shames SL, Wedler FC (1984) Homoserine kinase of Escherichia coli: kinetic mechanism and inhibition by L-aspartate semialdehyde. Arch Biochem Biophys 235(2):359–370
Huo X, Viola RE (1996) Substrate specificity and identification of functional groups of homoserine kinase from Escherichia coli. Biochemistry 35(50):16180–16185
Potter D &Miziorko HM. (1997) Identification of catalytic residues in human mevalonate kinase. J Biol Chem 272(41):25449–25454
Cox S, Radzio-Andzelm E, Taylor SS (1994) Domain movements in protein kinases. Curr Opin Struct Biol 4(6):893–901
Gerstein M, Lesk AM, Chothia C (1994) Structural mechanisms for domain movements in proteins. Biochemistry 33(22):6739–6749
Andreassi JL, Leyh TS (2004) Molecular functions of conserved aspects of the GHMP kinase family. Biochemistry 43(46):14594–14601
Aleshin AE, Kirby C, Liu X, Bourenkov GP, Bartunik HD, Fromm HJ, Honzatko RB (2000) Crystal structures of mutant monomeric hexokinase I reveal multiple ADP binding sites and conformational changes relevant to allosteric regulation. J Mol Biol 296(4):1001–1015
Romanowski MJ, Bonanno JB, Burley SK (2002) Crystal structure of the Streptococcus pneumoniae phosphomevalonate kinase, a member of the GHMP kinase superfamily. Proteins 47(4):568–571
Houten SM, Romeijn GJ, Koster J, Gray RG, Darbyshire P, Smit GP, de Klerk JB, Duran M, Gibson KM, Wanders RJ, Waterham HR (1999) Identification and characterization of three novel missense mutations in mevalonate kinase cDNA causing mevalonic aciduria, a disorder of isoprene biosynthesis. Hum Mol Genet 8(8):1523–1528
Potter D, Wojnar JM, Narasimhan C, Miziorko HM (1997) Identification and functional characterization of an active-site lysine in mevalonate kinase. J Biol Chem 272(9):5741–5746
Fu Z, Wang M, Potter D, Miziorko HM, Kim JJ (2002) The structure of a binary complex between a mammalian mevalonate kinase and ATP: insights into the reaction mechanism and human inherited disease. J Biol Chem 277(20):18134–18142
Luz JG, Hassig CA, Pickle C, Godzik A, Meyer BJ, Wilson IA (2003) XOL-1, primary determinant of sexual fate in C. elegans, is a GHMP kinase family member and a structural prototype for a class of developmental regulators. Genes Dev 17(8):977–990
Wada T, Kuzuyama T, Satoh S, Kuramitsu S, Yokoyama S, Unzai S, Tame JR, Park SY (2003) Crystal structure of 4-(cytidine 5′-diphospho)-2-C-methyl-D-erythritol kinase, an enzyme in the non-mevalonate pathway of isoprenoid synthesis. J Biol Chem 278(32):30022–30027
Cheek S, Zhang H, Grishin NV (2002) Sequence and structure classification of kinases. J Mol Biol 320(4):855–881
Marchenko GN, Marchenko ND, Tsygankov YD, Chistoserdov AY (1999) Organization of threonine biosynthesis genes from the obligate methylotroph Methylobacillus flagellatus. Microbiology 145(Pt 11):3273–3282
Patte JC, Clepet C, Bally M, Borne F, Méjean V, Foglino M (1999) ThrH, a homoserine kinase isozyme with in vivo phosphoserine phosphatase activity in Pseudomonas aeruginosa. Microbiology 145(Pt 4):845–853
Singh SK, Yang K, Karthikeyan S, Huynh T, Zhang X, Phillips MA, Zhang H (2004) The thrH gene product of Pseudomonas aeruginosa is a dual activity enzyme with a novel phosphoserine:homoserine phosphotransferase activity. J Biol Chem 279(13):13166–13173
Prati F, Goldman-Pinkovich A, Lizzi F, Belluti F, Koren R, Zilberstein D, Bolognesi ML (2014) Quinone-amino acid conjugates targeting Leishmania amino acid transporters. PLoS One 9(9):e107994. https://doi.org/10.1371/journal.pone.0107994
Marchese L, Nascimento JF, Damasceno FS, Bringaud F, Michels PAM, Silber AM (2018) The uptake and metabolism of amino acids, and their unique role in the biology of pathogenic trypanosomatids. Pathogens 7(2)
Acknowledgments
Authors are grateful to Dr. Sangeeta Sawant, Director, Bioinformatics Centre, for providing cluster machines for performing MD simulations and continuous support throughout this project. We also acknowledge Mr. Dattatraya Desai, Information Scientist, Bioinformatics Centre, for his critical suggestions during writing this manuscript.
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Meshram, R.J., Shirsath, A., Aouti, S. et al. Molecular modeling and simulation study of homoserine kinase as an effective leishmanial drug target. J Mol Model 26, 218 (2020). https://doi.org/10.1007/s00894-020-04473-7
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00894-020-04473-7