Abstract
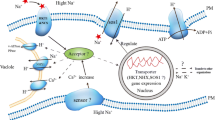
Salt stress in the rice field is one of the most common abiotic stresses, reducing crop productivity, especially at reproductive stage, which is very sensitive to salt stress. The aim of this investigation was to study mRNA-related Na+ uptake/translocation and Na+ enrichment in the cellular level, leading to physiological changes, growth characteristics, and yield attributes in FL530 [salt-tolerant genotype; carrying SKC1 (in relation to high-affinity potassium transporters controlling Na+ and K+ translocation) and qSt1b (linking to salt injury score) QTLs] and KDML105 (salt-sensitive cultivar; lacking both QTLs) parental lines and 221–48 (carrying SKC1 and qSt1b QTLs) derived from BILs (backcross introgression lines) at 50% flowering of rice, under 150-mM NaCl until harvesting process. The upregulation of OsHKT1;5 (mediating Na+ exclusion into xylem parenchyma cells) and OsNHX1 (Na+/H+ exchanger to secrete Na+ into vacuole) and downregulation of OsHKT2;1 and OsHKT2;2 (mediating Na+ restriction in the roots, leaf sheath and older leaves) in cvs. FL530 and 221–48 (+ SKC1; + qSt1b) under salt stress were observed. It restricted Na+ level in flag leaf, thereby preventing salt toxicity, as indicated by maintenance of photon yield of PSII (ΦPSII), net photosynthetic rate (Pn), transpiration rate (E) and overall growth performances. In contrast, Na+ enrichment in flag leaf of cv. KDML105 (−SKC1;−qSt1b) caused the reduction in ΦPSII by 30.5% over the control, leading to the reduction in Pn by 62.3%, in seed sterility by 88.2%, and yield loss by 85.1%. Moreover, the negative relationships between Na+ enrichment in flag leaf, physiological changes, and yield traits in rice crop grown under salt stress were demonstrated. Based on this investigation, rice genotype 221–48 was found to possess salt-tolerant traits at reproductive stage and thus could prove to be a potential candidate for future breeding programs.






Similar content being viewed by others
Abbreviations
- BILs:
-
backcross introgression lines
- OsHKT1;5:
-
plasma membrane-localized Na+ transporter
- OsHKT2;1 :
-
Na+ transporter gene
- OsHKT2;2 :
-
Na+ transporter gene
- KDML 105:
-
Khao Dawk Mali 105
- NaCl:
-
sodium chloride
- OsNHX1 :
-
vacuolar Na+/H+ exchanger gene
- Φ PSII :
-
photon yield of PSII
- P n :
-
net photosynthetic rate qSt1b SaltQTLchr1b
- QTLs:
-
quantitative trait loci
- SKC1:
-
sodium ion transporter
- E :
-
transpiration rate
References
Aguilar M, Fernández-Ramírez JL, Aguilar-Blanes M, Ortiz-Romero C (2017) Rice sensitivity to saline irrigation in southern Spain. Agric Water Manag 188:21–28. https://doi.org/10.1016/j.agwat.2017.03.027
Ahmadizadeh M, Vispo NA, Calapit-Palao CDO, Pangaan ID, Viña CD, Singh RK (2016) Reproductive stage salinity tolerance in rice: a complex trait to phenotype. Indian J Plant Physiol 21:528–536. https://doi.org/10.1007/s40502-016-0268-6
Apse MP, Blumwald E (2007) Na+ transport in plants. FEBS Lett 581:2247–2254. https://doi.org/10.1016/j.febslet.2007.04.01
Blumwald E (2000) Sodium transport and salt tolerance in plants. Curr Opin Cell Biol 12:431–434. https://doi.org/10.1016/S0955-0674(00)00112-5
Britto DT, Kronzucker HJ (2015) Sodium efflux in plant roots: what do we really know? J Plant Physiol 186:1–12. https://doi.org/10.1016/j.jplph.2015.08.002
Chakraborty K, Chattaopadhyay K, Nayak L, Ray S, Yeasmin L, Jena P, Gupta S, Mohanty SK, Swain P, Sarkar RK (2019) Ionic selectivity and coordinated transport of Na+ and K+ in flag leaves render differential salt tolerance in rice at the reproductive stage. Planta 250:1637–1653. https://doi.org/10.1007/s00425-019-03253-9
Chang J, Cheong BE, Natera S, Roessner U (2019) Morphological and metabolic responses to salt stress of rice (Oryza sativa L.) cultivars which differ in salinity tolerance. Plant Physiol Biochem 144:427–-435. https://doi.org/10.1016/j.plaphy.2019.10.017
Cha-um S, Supaibulwatana K, Kirdmanee C (2006) Water relation, photosynthetic ability and growth of Thai jasmine rice (Oryza sativa L. ssp. indica cv. KDML 105) to salt stress by application of exogenous glycinebetaine and choline. J Agron Crop Sci 192:25–36. https://doi.org/10.1111/j.1439-037X.2006.00186.x
Chen ZC, Yamaji N, Horie T, Che J, Li J, An G, Ma JF (2017) A magnesium transporter OsMGT1 plays a critical role in salt tolerance in rice. Plant Physiol 174:1837–1849. https://doi.org/10.1104/pp.17.00532
Chuamnakthong S, Nampei M, Ueda A (2019) Characterization of Na+ exclusion mechanism in rice under saline-alkaline stress conditions. Plant Sci 287:11071. https://doi.org/10.1016/j.plantsci.2019.110171
Cotsaftis O, Plett D, Shirley N, Tester M, Hrmova M (2012) A two-staged model of Na+ exclusion in rice explained by 3D model of HKT transporters and alternative splicing. PLoS ONE 7e39865. https://doi.org/10.1371/journal.pone.0039865
Fahad S, Hussain S, Matloob A, Khan FA, Khaliq A, Saud S, Hassan S, Shan D, Khan F, Ullah N, Faiq M, Khan MA, Tareen AK, Khan A, Ullah A, Ullah N, Huang J (2015) Phytohormones and plant responses to salinity stress: a review. Plant Growth Regul 75:391. https://doi.org/10.1007/s10725-014-0013-y
Flowers TJ, Hajibagheri MA (2001) Salt tolerance in Hordeum vulgare: ion concentrations in root cells of cultivars differing in salt tolerance. Plant Soil 231:1–9. https://doi.org/10.1023/A:1010372213938
Formentin E (2017) Salt tolerance in crops: not only a matter of gene regulation. Plant Physiol 174:1287–1288. https://doi.org/10.1104/pp.17.00747
Fukuda A, Nakamura A, Hara N, Toki S, Tanaka Y (2011) Molecular and functional analyses of rice NHX-type Na+ /H+ antiporter genes. Planta 233:175–188. https://doi.org/10.1007/s00425-010-1289-4
Garciadeblas B, Senn ME, Banuelos MA, Rodriguez-Navarro A (2003) Sodium transport and HKT transporters: the rice model. Plant J 34:788–801. https://doi.org/10.1046/j.1365-313X.2003.01764.x
Gay F, Maraval I, Roques S, Gunata Z, Boulanger R, Audebert A, Mestres C (2010) Effect of salinity on yield and 2-acetyl-1-pyrroline content in grains of three fragrant rice cultivars (Oryza sativa L.) in Camargue (France). Field Crop Res 117:154–160. https://doi.org/10.1016/j.fcr.2010.02.008
Gerona MEB, Deocampo MP, Egdane JA, Ismail AM, Dionisio-Sese ML (2019) Physiological responses of contrasting rice genotypes to salt stress at reproductive stage. Rice Sci 26:207–219. https://doi.org/10.1016/j.rsci.2019.05.001
Gorji T, Tanik A, Sertel E (2015) Soil salinity prediction, monitoring and mapping using modern technologies. Proc Earth Planet Sci 15:507–512. https://doi.org/10.1016/j.proeps.2015.08.062
Grattan S, Zeng L, Shannon M, Roberts S (2002) Rice is more sensitive to salinity than previously thought. Calif Agric 56:189–198. https://doi.org/10.3733/ca.v056n06p189
Hauser F, Horie T (2010) A conserved primary salt tolerance mechanism mediated by HKT transporters: a mechanism for sodium exclusion and maintenance of high K+/Na+ ratio in leaves during salinity stress. Plant Cell Environ 33:552–565. https://doi.org/10.1111/j.1365-3040.2009.02056.x
Horie T, Schroeder JI (2004) Sodium transporters in plants. Diverse genes and physiological functions. Plant Physiol 136:2457–2462. https://doi.org/10.1104/pp.104.046664
Horie T, Costa A, Kim TH, Han MJ, Horie R, Leung HY, Miyao A, Hirochika H, An G, Schroeder JI (2007) Rice OsHKT2;1 transporter mediates large Na+ influx component into K +-starved roots for growth. EMBO J 26:3003–3014. https://doi.org/10.1038/sj.emboj.7601732
Hossain MS (2019) Present scenario of global salt affected soils, its management and importance of salinity research. Int Res J Biol Sci 1:1–3
Hossain GS, Waditee R, Hibino T, Tanaka Y, Takabe T (2006) Root specific expression of Na+/H+ antiporter gene from Synechocystis sp. PCC6803 confers salt tolerance of tobacco plant. Plant Biotechnol 23:275–281. https://doi.org/10.5511/plantbiotechnology.23.275
IRRI (2002) Standard evaluation system for rice, 4th edition. International Rice Research Institute, P.O. Box 933, Manila, Philippines
Ishikawa T, Shabala S (2019) Control of xylem Na+ loading and transport to the shoot in rice and barley as a determinant of differential salinity stress tolerance. Physiol Plant 165:619–631. https://doi.org/10.1111/ppl.12758
Jain M, Nijhawan A, Tyagi AK, Khurana JP (2006) Validation of housekeeping genes as internal control for studying gene expression in rice by quantitative real-time PCR. Biochem Biophys Res Commun 345:646–651. https://doi.org/10.1016/j.bbrc.2006.04.140
Kader MA, Seidel T, Golldack D, Lindberg S (2006) Expressions of OsHKT1, OsHKT2, and OsVHA are differentially regulated under NaCl stress in salt-sensitive and salt-tolerant rice (Oryza sativa L.) cultivars. J Exp Bot 57:4257–4268. https://doi.org/10.1093/jxb/erl199
Kanpawee N, Sanitchon J, Srihaban P, Theerakulpisut P (2012) Genetic diversity analysis of rice cultivars (Oryza sativa L.) differing in salinity tolerance based on RAPD and SSR markers. Electron J Biotechnol 14:2–2. https://doi.org/10.2225/vol14-issue6-fulltext-4
Keisham M, Mukherjee S, Bhatla S (2018) Mechanisms of sodium transport in plants—progresses and challenges. Int J Mol Sci 19:647. https://doi.org/10.3390/ijms19030647
Khan MH, Panda SK (2008) Alterations in root lipid peroxidation and antioxidative responses in two rice cultivars under NaCl-salinity stress. Acta Physiol Plant 30:81–89. https://doi.org/10.1007/s11738-007-0093-7
Khush GS (2005) What it will take to feed 5.0 billion rice consumers in 2030. Plant Mol Biol 59:1–6. https://doi.org/10.1007/s11103-005-2159-5
Kobayashi NI, Yamaji N, Yamamoto H, Okubo K, Ueno H, Costa A, Tanoi K, Matsumura H, Fujii-Kashino M, Horiuchi T, Nayef MA, Shabala S, An G, Ma JF, Horie T (2017) OsHKT1;5 mediates Na+ exclusion in the vasculature to protect leaf blades and reproductive tissues from salt toxicity in rice. Plant J 91:657–670. https://doi.org/10.1111/tpj.13595
Kopittke PM (2012) Interactions between Ca, mg, Na and K: alleviation of toxicity in saline solutions. Plant Soil 352:353–362. https://doi.org/10.1007/s11104-011-1001-x
Kumar V, Khare T (2016) Differential growth and yield responses of salt-tolerant and susceptible rice cultivars to individual (Na+ and Cl−) and additive stress effects of NaCl. Acta Physiol Plant 38:170. https://doi.org/10.1007/s11738-016-2191-x
Lin HX, Zhu MZZ, Yano M, Gao JP, Liang ZW, Su WA, Hu XH, Ren ZH, Chao DY (2004) QTLs for Na+ and K+ uptake of shoot and root controlling rice salt tolerance. Theor Appl Genet 108:253–260. https://doi.org/10.1007/s00122-003-1421-y
Liu S, Zheng L, Xue Y, Zhang Q, Wang L, Shou H (2010) Overexpression of OsVP1 and OsNHX1 increases tolerance to drought and salinity in rice. J Plant Biol 53:444–452. https://doi.org/10.1007/s12374-010-9135-6
Liu M, Song X, Jiang Y (2018) Growth, ionic response, and gene expression of shoots and roots of perennial ryegrass under salinity stress. Acta Physiol Plant 40:112. https://doi.org/10.1007/s11738-018-2687-7
Liu J, Shabala S, Shabala L, Zhou M, Meinke H, Venkataraman G, Chen Z, Zeng F, Zhao Q (2019) Tissue-specific regulation of Na+ and K+ transporters explains genotypic differences in salinity stress tolerance in rice. Front Plant Sci 10:1361. https://doi.org/10.3389/fpls.2019.01361
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2-ΔΔCT method. Methods 25:402–408. https://doi.org/10.1006/meth.2001.1262
Loggini B, Scartazza A, Brugnoli E, Navari-Izzo F (1999) Antioxidant defense system, pigment composition, and photosynthetic efficiency in two wheat cultivars subjected to drought. Plant Physiol 119:1091–1099. https://doi.org/10.1104/pp.119.3.1091
Maxwell K, Johnson GN (2000) Chlorophyll fluorescence – a practical guide. J Exp Bot 51:659–668. https://doi.org/10.1093/jexbot/51.345.659
Mekawy AMM, Assaha DV, Yahagi H, Tada Y, Ueda A, Saneoka H (2015) Growth, physiological adaptation, and gene expression analysis of two Egyptian rice cultivars under salt stress. Plant Physiol Biochem 87:17–25. https://doi.org/10.1016/j.plaphy.2014.12.007
Mishra P, Mishra V, Singh NK, Rai V (2017) Gene expression dynamics of HKT family genes in salt-tolerant and salt-sensitive indica rice cultivars. Indian. J Genet 77:364–370. https://doi.org/10.5958/0975-6906.2017.00049.9
Mitsuya S, Murakami N, Sato T, Kazama T, Toriyama K, Skoulding NS, Kano-Nakata M, Yamauchi A (2019) Evaluation of rice grain yield and yield components of Nona Bokra chromosome segment substitution lines with the genetic background of Koshihikari, in a saline paddy field. AoB plants 11:plz040. https://doi.org/10.1093/aobpla/plz040
Munns R, Gilliham M (2015) Salinity tolerance of crops–what is the cost? New Phytol 208:668–673. https://doi.org/10.1111/nph.13519
Munns R, Tester M (2008) Mechanisms of salinity tolerance. Annu Rev Plant Biol 59:651–681. https://doi.org/10.1146/annurev.arplant.59.032607.092911
Munns R, James RA, Gilliham M, Flowers TJ, Colmer TD (2016) Tissue tolerance: an essential but elusive trait for salt-tolerant crops. Funct Plant Biol 43:1103–1113. https://doi.org/10.1071/FP16187
Negrão S, Schmöckel SM, Tester M (2017) Evaluating physiological responses of plants to salinity stress. Ann Bot 119:1–11. https://doi.org/10.1093/aob/mcw191
Nishimura T, Cha-um S, Takagaki M, Ohyama K, Kirdmanee C (2011) Survival percentage, photosynthetic abilities and growth characters of two indica rice (Oryza sativa L. spp. indica) cultivars in response to iso-osmotic stress. Span J Agric Res 9:262–270. https://doi.org/10.5424/sjar/20110901-162-10
Parihar P, Singh S, Singh R, Singh V, Prasad SM (2015) Effect of salinity stress on plants and its tolerance strategies: a review. Environ Sci Pollut Res 22:4056–4075. https://doi.org/10.1007/s11356-014-3739-1
Punyawaew K, Suriya-arunroj D, Siangliw M, Thida M, Lanceras-Siangliw J, Fukai S, Toojinda T (2016) Thai jasmine rice cultivar KDML105 carrying Saltol QTL exhibiting salinity tolerance at seedling stage. Mol Breed 36:150. https://doi.org/10.1007/s11032-016-0574-8
Puteh AB, Monjural M, Mondal A (2016) Salinity stress effect on ion uptake and yield attributes in rice. Life Sci J 13. https://doi.org/10.7537/marslsj131116.10
Qiu QS, Guo Y, Dietrich MA, Schumaker KS, Zhu JK (2002) Regulation of SOS1, a plasma membrane Na+/H+ exchanger in Arabidopsis thaliana, by SOS2 and SOS3. Proc Natl Acad Sci U S A 99:8436–8441. https://doi.org/10.1073/pnas.122224699
Ren Z-H, Gao J-P, Li L-G, Cai X-L, Huang W, Chao D-Y, Zhu M-Z, Wang Z-Y, Luan S, Lin H-X (2005) A rice quantitative trait locus for salt tolerance encodes a sodium transporter. Nat Genet 37:1141–1146. https://doi.org/10.1038/ng1643
Rengasamy P (2006) World salinization with emphasis on Australia. J Exp Bot 57:1017–1023. https://doi.org/10.1093/jxb/erj108
Riedelsberger J, Vergara-Jaque A, Piñeros M, Dreyer I, González W (2019) An extracellular cation coordination site influences ion conduction of OsHKT2;2. BMC Plant Biol 19:316. https://doi.org/10.1186/s12870-019-1909-5
Rodríguez-Navarro A, Rubio F (2006) High-affinity potassium and sodium transport systems in plants. J Exp Bot 57:1149–1160. https://doi.org/10.1093/jxb/erj068
Rojas RV, Achouri M, Maroulis J, Caon L (2016) Healthy soils: a prerequisite for sustainable food security. Environ Earth Sci 75(3):180. https://doi.org/10.1007/s12665-015-5099-7
Roy SJ, Negrão S, Tester M (2014) Salt resistant crop plants. Curr Opin Biotechnol 26:115–124. https://doi.org/10.1016/j.copbio.2013.12.004
Shabala S, Pottosin I (2014) Regulation of potassium transport in plants under hostile conditions: implications for abiotic and biotic stress tolerance. Physiol Plant 151:257–279. https://doi.org/10.1111/ppl.12165
Shi H, Quintero FJ, Pardo JM, Zhu JK (2002) The putative plasma membrane Na+/H+ antiporter SOS1 controls long-distance Na+ transport in plants. Plant Cell 14:465–477. https://doi.org/10.1105/tpc.010371
Shohan MU, Sinha S, Nabila FH, Dastidar SG, Seraj ZI (2019) HKT1;5 transporter gene expression and association of amino acid substitutions with salt tolerance across rice genotypes. Front Plant Sci 10:1420. https://doi.org/10.3389/fpls.2019.01420
Siangliw M, Phatanathara A, Toojinda T, Theerakulpisut P, Fukai S, Vanavichit A (2014) QTL and candidate gene identification for salt tolerance and Na/K ratio at seedling stage under modified soil and hydroponic conditions in rice (Oryza sativa L.). The 4th International Rice Congress (IRC2014), 27 October – 1 November 2014, Bangkok international trade and exhibition Centre (BITEC), Bangkok, Thailand
Singh V, Singh AP, Bhadoria J, Giri J, Singh J, Vineeth TV, Sharma PC (2018) Differential expression of salt-responsive genes to salinity stress in salt-tolerant and salt-sensitive rice (Oryza sativa L.) at seedling stage. Protoplasma 255:1667–1681. https://doi.org/10.1007/s00709-018-1257-6
Sultana N, Ikeda T, Itoh R (1999) Effect of NaCl salinity on photosynthesis and dry matter accumulation in developing rice grains. Environ Exp Bot 42:211–220. https://doi.org/10.1016/S0098-8472(99)00035-0
Suriya-arunroj D, Supapoj N, Toojinda T, Vanavichit A (2004) Relative leaf water content as an efficient method for evaluating rice cultivars for tolerance to salt stress. Sci Asia 30:411–415. https://doi.org/10.2306/scienceasia1513-1874.2004.30.411
Suriya-arunroj D, Supapoj N, Vanavichit A, Toojinda T (2005) Screening and selection for physiological characters contributing to salinity tolerance in rice. Nat Sci 39:174–185
Tanaka K, Ohta K, Haddad PR, Fritz JS, Lee KP, Hasebe K, Ieuji A, Miyanaga A (1999) Acid-rain monitoring in East Asia with a portable-type ion-exclusion-cation-exchange chromatographic analyzer. J Chromatogr 850:311–317. https://doi.org/10.1016/S0021-9673(99)00286-1
Theerawitaya C, Tisarum R, Samphumphuang T, Singh HP, Takabe T, Cha-um S (2019) Expression levels of vacuolar ion homeostasis-related genes, Na+ enrichment, and their physiological responses to salt stress in sugarcane genotypes. Protoplasma 257:525–536. https://doi.org/10.1007/s00709-019-01450-7
Theerawitaya C, Tisarum R, Samphumphuang T, Takabe T, Cha-um S (2020a) Expression levels of the Na+/K+ transporter OsHKT2;1 and vacuolar Na+/H+ exchanger OsNHX1, Na enrichment, maintaining the photosynthetic abilities and growth performances of indica rice seedlings under salt stress. Physiol Mol Biol Plants 26:513–523. https://doi.org/10.1007/s12298-020-00769-3
Theerawitaya C, Samphumphuang T, Tisarum R, Siangliw M, Cha-um S, Takabe T, Toojinda T (2020b) Transcriptional expression of Na homeostasis-related genes and physiological responses of rice seedlings under salt stress. J Plant Biochem Biotechnol. https://doi.org/10.1007/s13562-020-00573-w
Thomson MJ, de Ocampo M, Egdane J, Rahman MA, Sajise AG, Adorada DL, Tumimbang-Raiz E, Blumwald E, Seraj ZI, Singh RK, Gregorio GB, Gregorio GB (2010) Characterizing the Saltol quantitative trait locus for salinity tolerance in rice. Rice 3:148. https://doi.org/10.1007/s12284-010-9053-8
Thu TTP, Yasui H, Yamakawa T (2017) Effects of salt stress on plant growth characteristics and mineral content in diverse rice genotypes. Soil Sci Plant Nutr 63:264–273. https://doi.org/10.1080/00380768.2017.1323672
Ueda A, Yahagi H, Fujikawa Y, Nagaoka T, Esaka M, Calcaño M, González MM, Hernández-Martich JD, Saneoka H (2013) Comparative physiological analysis of salinity tolerance in rice. Soil Sci Plant Nutr 59:896–903. https://doi.org/10.1080/00380768.2013.842883
Vanavichit A, Kamolsukyeunyoung W, Siangliw M, Siangliw JL, Traprab S, Ruengphayak S, Chaichoompu E, Saensuk C, Phuvanartnarubal E, Toojinda T, Trakoonrung S (2018) Thai Hom Mali Rice: origin and breeding for subsistence rainfed lowland rice system. Rice 11:20. https://doi.org/10.1186/s12284-018-0212-7
Wankhade SD, Sanz A (2013) Chronic mild salinity affects source leaves physiology and productivity parameters of rice plants (Oryza sativa L., cv. Taipei 309). Plant Soil 367:663–672. https://doi.org/10.1007/s11104-012-1503-1
Wu H (2018) Plant salt tolerance and Na+ sensing and transport. Crop J 6:215–225. https://doi.org/10.1016/j.cj.2018.01.003
Wu GQ, Wang SM (2009) Calcium regulates K+/Na+ homeostasis is rice (Oryza sativa L.) under saline conditions. Plant Soil Environ 58:121–127
Wu Y, Hu Y, Xu G (2009) Interactive effects of potassium and sodium on root growth and expression of K/Na transporter genes in rice. Plant Growth Regul 57:271–280. https://doi.org/10.1007/s10725-008-9345-9
Yang Y, Guo Y (2018a) Elucidating the molecular mechanisms mediating plant salt-stress responses. New Phytol 217:523–539. https://doi.org/10.1111/nph.14920
Yang Y, Guo Y (2018b) Unraveling salt stress signaling in plants. J Integr Plant Biol 60:796–804. https://doi.org/10.1111/jipb.12689
Yao X, Horie T, Xue S, Leung HY, Katsuhara M, Brodsky DE, Wu Y, Schroeder JI (2010) Differential sodium and potassium transport selectivities of the rice OsHKT2;1 and OsHKT2;2 transporters in plant cells. Plant Physiol 152:341–355. https://doi.org/10.1104/pp.109.145722
Zhu JK (2016) Abiotic stress signaling and responses in plants. Cell 167:313–324. https://doi.org/10.1016/j.cell.2016.08.029
Acknowledgments
The authors would like to sincerely thank National Science and Technology Development Agency (NSTDA) for funding support (Grant number P-17-50079) and post-doctoral fellowship for CT.
Funding
This work was financially supported by National Science and Technology Development Agency (NSTDA) (Grant number P-17-50,079) and post-doctoral fellowship for CT.
Author information
Authors and Affiliations
Contributions
Suriyan Cha-um, Teruhiro Takabe, and Theerayut Toojinda conceived and designed the experiments. Cattarin Theerawitaya, Thapanee Samphumphuang, Rujira Tisarum, and Meechai Siangliw performed the experiments. Teruhiro Takabe provided advice on the analysis and interpretation of the data. Cattarin Theerawitaya and Suriyan Cha-um wrote the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Handling Editor: Handling Editor: Peter Nick
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Theerawitaya, C., Samphumphuang, T., Tisarum, R. et al. Expression level of Na+ homeostasis-related genes and salt-tolerant abilities in backcross introgression lines of rice crop under salt stress at reproductive stage. Protoplasma 257, 1595–1606 (2020). https://doi.org/10.1007/s00709-020-01533-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00709-020-01533-w