Abstract
In Saccharomyces cerevisiae, the mitoribosomal RNA of the minor subunit, 15S rRNA, is transcribed as a bicistronic transcript along with tRNAW. 5′ and 3′ sequences flanking the mature transcript must be removed by cleavage at the respective junctions before incorporating it into the mitoribosome. An in vivo dose–response triphasic system was created to elucidate the role of Ccm1p in the processing of 15S rRNA: Ccm1p supply (“On”), deprivation (“Off”), and resupply (“Back on”). After 72 h under “Off” status, the cells started to exhibit a complete mutant phenotype as assessed by their lack of growth in glycerol medium, while keeping their mitochondrial DNA integrity (ρ+). Full functionality of mitochondria was reacquired upon “Back on.” 15S rRNA levels and phenotype followed the Ccm1p intramitochondrial concentrations throughout the “On–Off–Back on” course. Under “Off” status, cells gradually accumulated unprocessed 5′ and 3′ junctions, which reached significant levels at 72–96 h, probably due to a saturation of the mitochondrial degradosome (mtEXO). The Ccm1p/mtEXO mutant (Δccm1/Δdss1) showed a copious accumulation of 15S rRNA primary transcript forms, which were cleaved upon Ccm1p resupply. The gene that codes for the RNA component of RNase P was conserved in wild-type and mutant strains. Our results indicate that Ccm1p is crucial in processing the 15S rRNA primary transcript and does not stabilize the already mature 15S rRNA. Consequently, failure of this function in Δccm1 cells results, as it happens to any other unprocessed primary transcripts, in total degradation of 15S rRNA by mtEXO, whose mechanism of action is discussed.




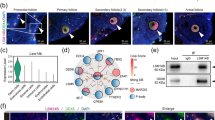
taken from a master plate of YEPG-geneticin, grown in SDGal for 48 h (“On”; t = 0 h), and cultured on SSD (“Off” [filled downward pointing arrows]) and SDGal (“On” [unfilled upward pointing arrows]). b From the 72-h SDD culture (“Off”), cells were either grown in SDD (“Off” [filled downward pointing arrows] or SDGal (“Back on” [half filled upward pointing arrows]), as shown by the small horizontal arrow. White-dashed rectangles indicate the acquired transient mutant and wild-type phenotype during the entire experimental course




Similar content being viewed by others
References
Amberg DC, Burke DJ, Strathern JN (2005) Methods in yeast genetics: a Cold Spring Harbor Laboratory Course Manual. Cold Spring Harbor Laboratory Press, Cold Spring Harbor
Anderson S, Bankier AT, Barrell BG, de Bruijn MH, Coulson AR, Drouin J, Eperon IC, Nierlich DP, Roe BA, Sanger F, Schreier PH, Smith AJ, Staden R, Young IG (1981) Sequence and organization of the human mitochondrial genome. Nature 290:457–465
Becker DM, Fikes JD, Guarente L (1991) A cDNA encoding a human CCAAT-binding protein cloned by functional complementation in yeast. Proc Natl Acad Sci USA 88:1968–1972
Bindoff LA, Howell N, Poulton J, McCullough DA, Morten KJ, Lightowlers RN, Turnbull DM, Weber K (1993) Abnormal RNA processing associated with a novel tRNA mutation in mitochondrial DNA. A potential disease mechanism. J Biol Chem 268:19559–19564
Biswas TK, Getz GS (1999) The single amino acid changes in the yeast mitochondrial S4 ribosomal protein cause temperature-sensitive defect in the accumulation of mitochondrial 15S rRNA. Biochemistry 38:13042–13054
Björkholm P, Harish A, Hagström E, Ernst AM, Andersson SGE (2015) Mitochondrial genomes are retained by selective constraints on protein targeting. Proc Natl Acad Sci USA 112:10154–10161
Boniecki MT, Rho SB, Tukalo M, Hsu JL, Romero EP, Martinis SA (2009) Leucyl-tRNA synthetase-dependent and-independent activation of a group I intron. J Biol Chem 284:26243–26250
Daoud R, Forget L, Lang BF (2012) Yeast mitochondrial RNase P, RNase Z and the RNA degradosome are part of a stable supercomplex. Nucleic Acids Res 4:1728–1736
de la Cruz J, Gómez-Herreros F, Rodríguez-Galán O, Begley V, de la Cruz M-C, Chávez S (2018) Feedback regulation of ribosome assembly. Curr Genet 64:393–404
De Silva D, Tu YT, Amunts A, Fontanesi F, Barrientos A (2015) Mitochondrial ribosome assembly in health and disease. Cell Cycle 14:2226–2250
Defontaine A, Lecocq FM, Hallet JN (1991) A rapid miniprep method for the preparation of yeast mitochondrial DNA. Nucleic Acids Res 19:185
Deutschmann AJ, Amberger A, Zavadil C, Steinbeisser H, Mayr JA, Feichtinger RG, Oerum S, Yue WW, Zschocke J (2014) Mutation or knock-down of 17β-hydroxysteroid dehydrogenase type 10 cause loss of MRPP1 and impaired processing of mitochondrial heavy strand transcripts. Hum Mol Genet 23:3618–3628
Dziembowski A, Malewicz M, Minczuk M, Golik P, Dmochowska A, Stepien PP (1998) The yeast nuclear gene DSS1, which codes for a putative RNase II, is necessary for the function of the mitochondrial degradosome in processing and turnover of RNA. Mol Gen Genet 260:108–114
Dziembowski AJ, Piwowarski R, Hoser M, Minczuk A, Dmochowska M, Siep H, van der Spek LG, Stepien PP (2003) The yeast mitochondrial degradosome. Its composition, interplay between RNA helicase and RNase activities and the role in mitochondrial RNA metabolism. J Biol Chem 278:1603–1611
Ellis TP, Helfenbein KG, Tzagoloff A, Dieckmann CL (2004) Aep3p stabilizes the mitochondrial bicistronic mRNA encoding subunits 6 and 8 of the H+-translocating ATP synthase of Saccharomyces cerevisiae. J Biol Chem 16:15728–15733
Falk MJ, Gai X, Shigematsu M, Vilardo E, Takase R, McCormick E, Christian T, Place E, Pierce EA, Consugar M, Gamper HB, Rossmanith W, Hou YM (2016) A novel HSD17B10 mutation impairing the activities of the mitochondrial RNase P complex causes X-linked intractable epilepsy and neurodevelopmental regression. RNA Biol 13:477–485
Fang F, Phillips S, Butler JS (2005) Rat1p and Rai1p function with the nuclear exosome in the processing and degradation of rRNA precursors. RNA 11:1571–1578
Fekete Z, Ellis TP, Schonauer MS, Dieckmann CL (2008) Pet127 governs a 5′ 3′-exonuclease important in maturation of apocytochrome b mRNA in Saccharomyces cerevisiae. J Biol Chem 7:3767–3772
Foury F, Roganti T, Lecrenier N, Purnelle B (1998) The complete sequence of the mitochondrial genome of Saccharomyces cerevisiae. FEBS Lett 440:325–331
Fox TD (2012) Mitochondrial protein synthesis, import, and assembly. Genetics 192:1203–1234
Fritsch ES, Chabbert CD, Klaus B, Steinmetz LM (2014) A genome-wide map of mitochondrial DNA recombination in yeast. Genetics 198:755–771
Gan X, Kitakawa M, Yoshino KI, Oshiro N, Yonezawa K, Isono K (2002) Tag-mediated isolation of yeast mitochondrial ribosome and mass spectrometric identification of its new components. Eur J Biochem 269:5203–5214
Gavin AC et al (2002) Functional organization of the yeast proteome by systematic analysis of protein complexes. Nature 415:141–147
Gobert A, Gutmann B, Taschner A, Gössringer M, Holzmann J, Hartmann RK, Rossmanith W, Giegé P (2010) A single Arabidopsis organellar protein has RNase P activity. Nat Struct Mol Biol 17:740–744
Gray MW (2012) Mitochondrial evolution. Cold Spring Harb Perspect Biol 4:a011403
Guo XE, Chen CF, Wang DD, Modrek AS, Phan VH, Lee WH, Chen PL (2011) Uncoupling the roles of the SUV3 helicase in maintenance of mitochondrial genome stability and RNA degradation. J Biol Chem 286:38783–38794
Haack TB, Kopajtich R, Freisinger P, Wieland T, Rorbach J, Nicholls TJ, Baruffini E, Walther A, Danhauser K, Zimmermann FA, Husain RA, Schum J, Mundy H, Ferrero I, Strom TM, Meitinger T, Taylor RW, Minczuk M, Mayr JA, Prokisch H (2013) ELAC2 mutations cause a mitochondrial RNA processing defect associated with hypertrophic cardiomyopathy. Am J Hum Genet 93:211–223
Heude M, Fukuhara H, Moustacchi E (1979) Spontaneous and induced rho mutants of Saccharomyces cerevisiae: patterns of loss of mitochondrial genetic markers. J Bacteriol 139:460–467
Hollingsworth MJ, Martin NC (1986) RNase P activity in the mitochondria of Saccharomyces cerevisiae depends on both mitochondrion and nucleus-encoded components. Mol Cell Biol 6:1058–1064
Holzmann J, Frank P, Löffler E, Bennett KL, Gerner C, Rossmanith W (2008) RNase P without RNA: identification and functional reconstitution of the human mitochondrial tRNA processing enzyme. Cell 135:462–474
Jhuang HY, Lee HY, Leu JY (2017) Mitochondrial-nuclear co-evolution leads to hybrid incompatibility through pentatricopeptide repeat proteins. EMBO Rep 1:87–101
Jourdain AA, Koppen M, Rodley CD, Maundrell K, Gueguen N, Reynier P, Guaras AM, Enriquez JA, Anderson P, Simarro M, Martinou JC (2015) A mitochondria-specific isoform of FASTK is present in mitochondrial RNA granules and regulates gene expression and function. Cell Rep 10:1110–1121
Keeling PJ, Palmer JD (2008) Horizontal gene transfer in eukaryotic evolution. Nat Rev Genet 9:605–618
Kobayashi K, Kawabata M, Hisano K, Kazama T, Matsuoka K, Sugita M, Nakamura T (2012) Identification and characterization of the RNA binding surface of the pentatricopeptide repeat protein. Nucleic Acids Res 40:2712–2723
Kritsiligkou P, Chatzi A, Charalampous G, Mironov A Jr, Grant CM, Tokatlidis K (2017) Unconventional targeting of a thiol peroxidase to the mitochondrial intermembrane space facilitates oxidative protein folding. Cell Rep 18:2729–2741
Manna S (2015) An overview of pentatricopeptide repeat proteins and their applications. Biochimie 113:93–99
Manthey GM, McEwen JE (1995) The product of the nuclear gene PET309 is required for translation of mature mRNA and stability or production of intron-containing RNAs derived from the mitochondrial COX1 locus of Saccharomyces cerevisiae. EMBO J 14:4031–4043
Margossian SP, Li H, Zassenhaus HP, Butow RA (1996) The DExH box protein Suv3p is a component of a yeast mitochondrial 3′-to-5′ exoribonuclease that suppresses group I intron toxicity. Cell 84:199–209
Meisinger C, Pfanner N, Truscott KN (2006) Isolation of yeast mitochondria. Methods Mol Biol 313:33–39
Merz S, Westermann B (2009) Genome-wide deletion mutant analysis reveals genes required for respiratory growth, mitochondrial genome maintenance and mitochondrial protein synthesis in Saccharomyces cerevisiae. Genome Biol 10:R95
Metodiev MD, Thompson K, Alston CL, Morris AA, He L, Assouline Z, Rio M, Bahi-Buisson N, Pyle A, Griffin H, Siira S, Filipovska A, Munnich A, Chinnery PF, McFarland R, Rötig A, Taylor RW (2016) Recessive mutations in TRMT10C cause defects in mitochondrial RNA processing and multiple respiratory chain deficiencies. Am J Hum Genet 98:993–1000
Mishra P, Chan DC (2014) Mitochondrial dynamics and inheritance during cell division, development and disease. Nat Rev Mol Cell Biol 15:634–646
Möller-Hergt BV, Carlström A, Stephan K, Imhof A, Ott M (2018) The ribosome receptors Mrx15 and Mba1 jointly organize cotranslational insertion and protein biogenesis in mitochondria. Mol Biol Cell 29:2386–2396
Montoya J, Christianson T, Levens D, Rabinowitz M, Attardi G (1982) Identification of initiation sites for heavy-strand and light-strand transcription in human mitochondrial DNA. Proc Natl Acad Sci USA 79:7195–7199
Moreno JI (1996) A Trypanosoma cruzi polyantigen obtained by gene fusion: its expression in Staphylococcus aureus and rapid purification. Protein Expr Purif 8:332–340
Moreno JI, Buie KS, Price RE, Piva MA (2009) Ccm1p/Ygr150cp, a pentatricopeptide repeat protein, is essential to remove the fourth intron of both COB and COX1 pre-mRNAs in Saccharomyces cerevisiae. Curr Genet 55:475–484
Moreno JI, Patlolla B, Belton KR, Jenkins BC, Radchenkova PV, Piva MA (2012) Two independent activities define Ccm1p as a moonlighting protein in Saccharomyces cerevisiae. Biosci Rep 32:549–557
Mörl M, Marchfelder A (2001) The final cut. The importance of tRNA 3′-processing. EMBO Rep 2:17–20
Nandakumar MP, Shen J, Raman B, Marten MR (2003) Solubilization of trichloroacetic acid (TCA) precipitated microbial proteins via NaOH for two-dimensional electrophoresis. J Proteome Res 2:89–93
Naquin D, Panozzo C, Dujardin G, van Dijk E, d'Aubenton-Carafa Y, Thermes C (2018) Complete sequence of the intronless mitochondrial genome of the Saccharomyces cerevisiae strain CW252. Genome Announc 6:e00219–e318
Osinga KA, Evers RF, Van der Laan JC, Tabak HF (1981) A putative precursor for the small ribosomal RNA from mitochondria of Saccharomyces cerevisiae. Nucleic Acids Res 9:1351–1364
Perks KL, Rossetti G, Kuznetsova I, Hughes LA, Ermer JA, Ferreira N, Busch JD, Rudler DL, Spahr H, Schöndorf T, Shearwood AM (2018) PTCD1 is required for 16S rRNA maturation complex stability and mitochondrial ribosome assembly. Cell Rep 23:127–142
Pillon MC, Stanley RE (2018) Nuclease integrated kinase super assemblies (NiKs) and their role in RNA processing. Curr Genet 64:183–190
Pinker F, Bonnard G, Gobert A, Gutmann B, Hammani K, Sauter C, Gegenheimer PA, Giegé P (2013) PPR proteins shed a new light on RNase P biology. RNA Biol 10:1457–1468
Rackham O, Davies SM, Shearwood AM, Hamilton KL, Whelan J, Filipovska A (2009) Pentatricopeptide repeat domain protein 1 lowers the levels of mitochondrial leucine tRNAs in cells. Nucleic Acids Res 17:5859–5867
Sánchez MI, Mercer TR, Davies SM, Shearwood AM, Nygard KK, Richman TR, Mattick JS, Rackham O, Filipovska A (2011) RNA processing in human mitochondria. Cell Cycle 10:2904–2916
Schülke N, Sepuri NB, Pain D (1997) In vivo zippering of inner and outer mitochondrial membranes by a stable translocation intermediate. Proc Natl Acad Sci USA 94:7314–7319
Séraphin B, Boulet A, Simon M, Faye G (1987) Construction of a yeast strain devoid of mitochondrial introns and its use to screen nuclear genes involved in mitochondrial splicing. Proc Natl Acad Sci USA 84:6810–6814
Shajani Z, Sykes MT, Williamson JR (2011) Assembly of bacterial ribosomes. Annu Rev Biochem 80:501–526
Slomovic S, Laufer D, Geiger D, Schuster G (2005) Polyadenylation and degradation of human mitochondrial RNA: the prokaryotic past leaves its mark. Mol Cell Biol 25:6427–6435
Small ID, Peeters N (2000) The PPR motif a TPR-related motif prevalent in plant organellar proteins. Trends Biochem Sci 25:46–47
Stribinskis V, Gao GJ, Sulo P, Dang YL, Martin NC (1996) Yeast mitochondrial RNase P RNA synthesis is altered in an RNase P protein subunit mutant: insights into the biogenesis of a mitochondrial RNA-processing enzyme. Mol Cell Biol 16:3429–3436
Szczesny RJ, Borowski LS, Brzezniak LK, Dmochowska A, Gewartowski K, Bartnik E, Stepien PP (2010) Human mitochondrial RNA turnover caught in flagranti: involvement of hSuv3p helicase in RNA surveillance. Nucleic Acids Res 38:279–298
Taschner A, Weber C, Buzet A, Hartmann RK, Hartig A, Rossmanith W (2012) Nuclear RNase P of Trypanosoma brucei: a single protein in place of the multicomponent RNA-protein complex. Cell Rep 2:19–25
Teste MA, Duquenne M, François JM, Parrou JL (2009) Validation of reference genes for quantitative expression analysis by real-time RT-PCR in Saccharomyces cerevisiae. BMC Mol Biol 10:99
Turk EM, Das V, Seibert RD, Andrulis ED (2013) The mitochondrial RNA landscape of Saccharomyces cerevisiae. PLoS ONE 8:e78105
Ulery TL, Jang SH, Jaehning JA (1994) Glucose repression of yeast mitochondrial transcription: kinetics of derepression and role of nuclear genes. Mol Cell Biol 14:1160–1170
Van Haute L, Pearce SF, Powell CA, D'Souza AR, Nicholls TJ, Minczuk M (2015) Mitochondrial transcript maturation and its disorders. J Inherit Metab Dis 38:655–680
Wang Z, Sun X, Wee J, Gu Z (2019) Novel insights into global translational regulation through Pumilio family RNA-binding protein Puf3p revealed by ribosomal profiling. Curr Genet 65:201–212
Wiesenberger G, Fox TD (1997) Pet127p, a membrane-associated protein involved in stability and processing of Saccharomyces cerevisiae mitochondrial RNAs. Mol Cell Biol 17:2816–2824
Xu F, Ackerley C, Maj MC, Addis JB, Levandovskiy V, Lee J, Mackay N, Cameron JM, Robinson BH (2008) Disruption of a mitochondrial RNA-binding protein gene results in decreased cytochrome b expression and a marked reduction in ubiquinol-cytochrome c reductase activity in mouse heart mitochondria. Biochem J 416:15–26
Acknowledgements
This work was supported by the Mississippi INBRE funded by grants from the National Center for Research Resources [5P20RR016476] and the National Institute of General Medical Sciences (NIGMS) [8P20GM103476] from the National Institutes of Health (NIH), and by grants number [5SC3GM087169] and [W911NF-13-1-0174] from NIGMS, NIH, and DoD, respectively. We thank Mrs. Kimberley S. Buie for her contributions. The support from Dr. Sandra Barnes, Department of Chemistry and Physics, and the office of the Dean of the School of Art and Sciences are honestly appreciated. Finally, we acknowledge the valuable input from the reviewers.
Author information
Authors and Affiliations
Corresponding authors
Additional information
Communicated by M. Kupiec.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Moreno, J.I., Coleman, I.S., Johnson, C.L. et al. Ccm1p is a 15S rRNA primary transcript processing factor as elucidated by a novel in vivo system in Saccharomyces cerevisiae. Curr Genet 66, 775–789 (2020). https://doi.org/10.1007/s00294-020-01064-0
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00294-020-01064-0