Abstract
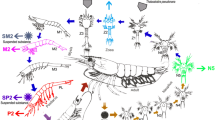
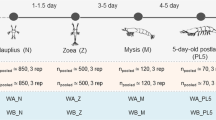
This study aimed to evaluate the structural changes in the microbiota of Penaeus monodon during its ontogenetic development and evaluate functional shifts in metabolic pathways. To address these questions, shrimp were maintained under isolated conditions to avoid cross-contamination with other environmental bacteria and reared in twelve 50 L plastic tanks connected to a recirculation system. The replicates of pooled intestine specimens from 15 shrimp were analyzed at postlarvae, juvenile, and adult phases for nucleic acid extraction. We tested the V3 region of the 16S rRNA gene from the postlarvae, juvenile, and adult stages. Resulting amplicon sequence variants (ASVs) were used to evaluate both structural and functional shifts during tiger shrimp development. Although the richness and diversity of microbiota were only slightly affected, the phylum Proteobacteria (genus Vibrio) was significantly predominant in the earliest stages of life, while Actinobacteria (genus Actinotalea) significantly increased in adult shrimp. Moreover, functional predictions by PICRUSt suggested that microbiota is primarily involved in metabolic activities during each developmental stage. Overall, these results suggest that each ontogenetic stage may provide specific conditions for the proliferation of certain bacterial taxa. Nerveless, alternative microbiota maintains similar ecological activities; thus, we posit that “stage-specific” changes in microbiota may be regulated to guarantee pivotal host functions.





Similar content being viewed by others
References
Anderson MJ (2001) A new method for non-parametric multivariate analysis of variance. Austral Ecol 26:32–46. https://doi.org/10.1080/13645700903062353
Bokulich NA, Subramanian S, Faith JJ, Gevers D, Gordon I, Knight R, Mills DA, Caporaso JG (2013) Quality-filtering vastly improves diversity estimates from Illumina amplicon sequencing. Nat Methods 10:57–59. https://doi.org/10.1038/nmeth.2276.Quality-filtering
Bolyen E, Rideout JR, Dillon MR, Bokulich NA, Abnet CC, Al-Ghalith GA, Bai Y (2019) Reproducible, interactive, scalable and extensible microbiome data science using QIIME 2. Nat Biotechnol 37:852–857. https://doi.org/10.1038/s41587-019-0209-9
Callahan BJ, McMurdie PJ, Rosen MJ, Han AW, Johnson AJA, Holmes SP (2016) DADA2: high-resolution sample inference from Illumina amplicon data. Nat Methods 13:581–583. https://doi.org/10.1038/nmeth.3869
Camacho C, Coulouris G, Avagyan V, Ma N, Papadopoulos J, Bealer K, Madden TL (2009) BLAST+: architecture and applications. BMC Bioinformatics 10:1–9. https://doi.org/10.1186/1471-2105-10-421
Chaiyapechara S, Rungrassamee W, Suriyachay I, Kuncharin Y, Klanchui A, Karoonuthaisiri N, Jiravanichpaisal P (2012) Bacterial community associated with the intestinal tract of P. monodon in commercial farms. Microb Ecol 63:938–953. https://doi.org/10.1007/s00248-011-9936-2
Chen WY, Ng TH, Wu JH, Chen JW, Wang HC (2017) Microbiome dynamics in a shrimp grow-out pond with possible outbreak of acute hepatopancreatic necrosis disease. Sci Rep 7:1–12. https://doi.org/10.1038/s41598-017-09923-6
Clarke KR, Warwick RM (2001) A further biodiversity index applicable to species lists: variation in taxonomic distinctness. Mar Ecol Prog Ser 216:265–278. https://doi.org/10.3354/meps216265
Cornejo-Granados F, Gallardo-Becerra L, Leonardo-Reza M, Ochoa-Romo JP, Ochoa-Leyva A (2018) A meta-analysis reveals the environmental and host factors shaping the structure and function of the shrimp microbiota. PeerJ. https://doi.org/10.7717/peerj.5382
de Voogd NJ, Cleary DFR, Polónia ARM, Gomes NCM (2015) Bacterial community composition and predicted functional ecology of sponges, sediment and seawater from the thousand islands reef complex, West Java, Indonesia. FEMS Microbiol Ecol 91:1–12. https://doi.org/10.1093/femsec/fiv019
Douglas GM, Maffei VJ, Zaneveld J, Yurgel SN, Brown JR, Taylor CM, Huttenhower C, Langille MGI (2019) PICRUSt2: an improved and extensible approach for metagenome inference. BioRxiv:1–42. https://doi.org/10.3997/2214-4609.201404048
Duan Y, Liu Q, Wang Y, Zhang J, Xiong D (2018) Impairment of the intestine barrier function in Litopenaeus vannamei exposed to ammonia and nitrite stress. Fish Shellfish Immunol 78:279–288. https://doi.org/10.1016/j.fsi.2018.04.050
Erasmus JH, Cook P, Coyne VE (1997) The role of bacteria in the digestion of seaweed by the abalone Haliotis midae. Aquaculture 155:377–386. https://doi.org/10.1016/S0044-8486(97)00112-9
Garibay-Valdez E, Martínez-Córdova LR, Vargas-Albores F, Gollas-Galván T, Lago-Leston A, Calderón K, Martínez-Porchas M (2019) Biofilm consumption shapes the intestinal microbiota of shrimp (Penaeus vannamei). Aquac Nutr 25:427–435. https://doi.org/10.1111/anu.12868
Gollas-Galvan T, Cabanillas-Gámez M, Hernández-López J, Coronado-Molina D, Martínez-Porchas M (2017) Transcriptional expression of immune system genes in Litopenaeus vannamei during ontogenetic development. Aquac Res 48:1110–1118. https://doi.org/10.1111/are.12953
Heberle H, Meirelles VG, da Silva FR, Telles GP, Minghim R (2015) InteractiVenn: a web-based tool for the analysis of sets through Venn diagrams. BMC Bioinformatics 16:1–7. https://doi.org/10.1186/s12859-015-0611-3
Huse SM, Dethlefsen L, Huber JA, Welch DM, Relman DA, Sogin ML (2008) Exploring microbial diversity and taxonomy using SSU rRNA hypervariable tag sequencing. PLoS Genet 4. https://doi.org/10.1371/journal.pgen.1000255
Katoh K, Standley DM (2013) MAFFT multiple sequence alignment software version 7: improvements in performance and usability. Mol Biol Evol 30:772–780. https://doi.org/10.1093/molbev/mst010
Langille M, Zaneveld J, Caporaso JG, McDonald D, Knights D, Reyes J, Clemente J, Burkepile D, Vega Thurber R, Knight R, Beiko R, Huttenhower C (2013) Predictive functional profiling of microbial communities using 16S rRNA marker gene sequences. Nat Biotechnol 31:814–821. https://doi.org/10.1038/nbt.2676
Lovett D, Felder D (1989) Ontogeny of gut morphology in the white shrimp Penaeus setiferus (Decapoda, Penaeidae). J Morphol 272:253–272
Martínez-Córdova, LR (1993) Camaronicultura: Bases técnicas y científicas para el cultivo de camarones peneidos. Universidad de Sonora. AGT Editor. México, D.F. p 233
McNally CP, Eng A, Noecker C, Gagne-Maynard WC, Borenstein E (2018) BURRITO: an interactive multi-omic tool for visualizing taxa-function relationships in microbiome data. Front Microbiol 9:1–11. https://doi.org/10.3389/fmicb.2018.00365
Moya A, Ferrer M (2016) Functional redundancy-induced stability of gut microbiota subjected to disturbance. Trends Microbiol 24:402–413
Nicholson JK, Holmes E, Kinross J, Burcelin R, Gibson G, Jia W, Pettersson S (2012) Host-gut microbiota metabolic interactions. Science 336:1262–1268
Parks D, Tyson GW, Hugenholtz P, Beiko RG (2014) STAMP: statistical analysis of taxonomic and functional profiles. Bioinformatics 30:3123–3124. https://doi.org/10.1093/bioinformatics/btu494
Price MN, Dehal PS, Arkin AP (2010) FastTree 2-approximately maximum-likelihood trees for large alignments. PLoS One 5. https://doi.org/10.1371/journal.pone.0009490
Quast C, Pruesse E, Yilmaz P, Gerken J, Schweer T, Yarza P, Peplies J, Glöckner FO (2013) The SILVA ribosomal RNA gene database project: improved data processing and web-based tools. Nucleic Acids Res 41:590–596. https://doi.org/10.1093/nar/gks1219
Ramesh S, Mathivanan N (2009) Screening of marine actinomycetes isolated from the Bay of Bengal, India for antimicrobial activity and industrial enzymes. World J Microbiol Biotechnol 25:2103–2111. https://doi.org/10.1007/s11274-009-0113-4
Rungrassamee W, Klanchui A, Maibunkaew S, Chaiyapechara S, Jiravanichpaisal P, Karoonuthaisiri N (2014) Characterization of intestinal bacteria in wild and domesticated adult black tiger shrimp (Penaeus monodon). PLoS One 9:e91853. https://doi.org/10.1371/journal.pone.0091853
Saulnier D, Haffner P, Goarant C, Levy P, Ansquer D (2000) Experimental infection models for shrimp vibriosis studies: a review. Aquaculture 191:133–144. https://doi.org/10.1016/S0044-8486(00)00423-3
Segata N, Izard J, Waldron L, Gevers D, Miropolsky L, Garrett WSW, Huttenhower C, Golub T, Petricoin E, Ardekani A et al (2011) Metagenomic biomarker discovery and explanation. Genome Biol 12:1–18. https://doi.org/10.1186/gb-2011-12-6-r60
Soriano EL, Ramírez DT, Araujo DR, Gómez-Gil B, Castro LI, Sánchez CG (2018) Effect of temperature and dietary lipid proportion on gut microbiota in yellowtail kingfish Seriola lalandi juveniles. Aquaculture 497:269–277. https://doi.org/10.1016/j.aquaculture.2018.07.065
Subramani R, Aalbersberg W (2013) Culturable rare Actinomycetes: diversity, isolation and marine natural product discovery. Appl Microbiol Biotechnol 97:9291–9321. https://doi.org/10.1007/s00253-013-5229-7
Tacon AGJ (1990) Standard methods for the nutrition and feeding of farmed fish and shrimp. Argent Laboratories Press, Redmond, p 208
Tarazona J, León Q, Galindo C, Vallejo M, Marguet E (2018) Caracterización de actinomicetos de sedimento marino y su actividad antagonista frente a Vibrio sp aislados de «langostino blanco» Litopenaeus vannamei (Boone, 1931). Rev Investig Vet del Perú 29:676. https://doi.org/10.15381/rivep.v29i2.14499
Vargas-Albores F, Porchas-Cornejo MA, Martínez-Porchas M, Villalpando-Canchola E, Gollas-Galván T, Martínez-Córdova LR (2017) Bacterial biota of shrimp intestine is significantly modified by the use of a probiotic mixture: a high throughput sequencing approach. Helgol Mar Res:71. https://doi.org/10.1186/s10152-017-0485-z
Vázquez-Baeza Y, Pirrung M, Gonzalez A, Knight R (2013) EMPeror: a tool for visualizing high-throughput microbial community data. Gigascience 2:1–4
Venema K (2010) Role of gut microbiota in the control of energy and carbohydrate metabolism. Curr Op Clin Nutr Metab Care 13:432–438
Wang J, Huang Y, Xu K, Zhang X, Sun H, Fan L, Yan M (2019) White spot syndrome virus (WSSV) infection impacts intestinal microbiota composition and function in Litopenaeus vannamei. Fish Shellfish Immunol 84:130–137. https://doi.org/10.1016/j.fsi.2018.09.076
Wigglesworth JM, Griffith DRW (1994) Carbohydrate digestion in Penaeus monodon. Mar Biol 120:571–578. https://doi.org/10.1007/BF00350077
Xiong J, Wang K, Wu J, Qiuqian L, Yang K, Qian Y, Zhang D (2015) Changes in intestinal bacterial communities are closely associated with shrimp disease severity. Appl Microbiol Biotechnol 99:6911–6919. https://doi.org/10.1007/s00253-015-6632-z
Xiong J, Dai W, Zhu J, Liu K, Dong C, Qiu Q (2017) The underlying ecological processes of gut microbiota among cohabitating retarded, overgrown and normal shrimp. Microb Ecol 73:988–999. https://doi.org/10.1007/s00248-016-0910-x
Yan Q, Li J, Yu Y, Wang J, He Z, Van Nostrand JD, Kempher ML, Wu L, Wang Y, Liao L, Li X, Wu S, Ni J, Wang C, Zhou J (2016) Environmental filtering decreases with fish development for the assembly of gut microbiota. Environ Microbiol 18:4739–4754. https://doi.org/10.1111/1462-2920.13365
Acknowledgments
We are grateful to Dr. W. Kelley Thomas who gives us the possibility to run bioinformatics pipelines using the server of the Research Computing Center at the University of New Hampshire.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical approval
All applicable international, national, and institutional guidelines for the care and use of animals were followed by the authors.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Cicala, F., Lago-Lestón, A., Gomez-Gil, B. et al. Gut microbiota shifts in the giant tiger shrimp, Penaeus monodon, during the postlarvae, juvenile, and adult stages. Aquacult Int 28, 1421–1433 (2020). https://doi.org/10.1007/s10499-020-00532-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10499-020-00532-1