Abstract
Background
Drought is the major abiotic stress factor that negatively influences growth and yield in cereal grain crops such as maize (Zea mays L.). A multitude of genes and pathways tightly modulate plant growth, development and responses to environmental stresses including drought. Therefore, crop breeding efforts for enhanced drought resistance require improved knowledge of plant drought responses.
Objective
Here, we sought to elucidate the molecular and physiological mechanisms underpinning maize drought stress tolerance.
Methods
We therefore applied a 12-day water-deficit stress treatment to maize plants of two contrasting (drought tolerant ND476 and drought sensitive ZX978) hybrid cultivars at the late vegetative (V12) growth stage and performed a large-scale RNA sequencing (RNA-seq) transcriptome analysis of the leaf tissues.
Results
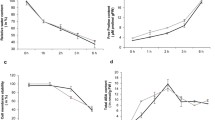
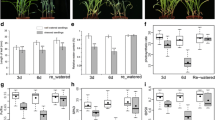
A comparative analysis of the two genotypes leaf transcriptomes and physiological parameters revealed the key differentially expressed genes (DEGs) and metabolic pathways that respond to drought in a genotype-specific manner. A total of 3114 DEGs were identified, with 21 DEGs being specifically expressed in tolerant genotype ND476 in response to drought stress. Of these, genes involved in secondary metabolites biosynthesis, transcription factor regulation, detoxification and stress defense were highly expressed in ND476. Physiological analysis results substantiated our RNA-seq data, with ND476 exhibiting better cell water retention, higher soluble protein content and guaiacol peroxidase activity, along with low lipid peroxidation extent than the sensitive cultivar ZX978 under drought conditions.
Conclusion
Our findings enrich the maize genetic resources and enhance our further understanding of the molecular mechanisms regulating drought stress tolerance in maize. Additionally, the DEGs screened in this study may provide a foundational basis for our future targeted cloning studies.






Similar content being viewed by others
References
Ahmad N, Malagoli M, Wirtz M, Hell R (2016) Drought stress in maize causes differential acclimation responses of glutathione and sulfur metabolism in leaves and roots. BMC Plant Biol 16:247. https://doi.org/10.1186/s12870-016-0940-z
Ahmadi A, Emam Y, Pessarakli M (2010) Biochemical changes in maize seedlings exposed to drought stress conditions at different nitrogen levels. J Plant Nutr 33:541–556. https://doi.org/10.1080/01904160903506274
Altschul SF, Madden TL, Schäffer AA, Zhang J, Zhang Z, Miller W, Lipman DJ (1997) Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucl Acids Res 25(17):3389–3402. https://doi.org/10.1093/nar/25.17.3389
Al-Whaibi MH (2011) Plant heat-shock proteins: A mini review. J King Saud Univ Sci 23:139–150. https://doi.org/10.1016/j.jksus.2010.06.022
Anjum SA, Xie XY, Wang LC, Saleem MF, Man C, Lei W (2011) Morphological, physiological and biochemical responses of plants to drought stress. Afr J Agric Res 6:2026–2032
Aslam M, Maqbool MA, Cengiz R (2015) Drought stress in maize (Zea mays L.): Effects, resistance mechanisms, global achievements and biological strategies for improvement. Springer: Cham, ISBN 978-3-319-25440-1.
Banerjee A, Roychoudhury A (2017) Abscisic-acid-dependent basic leucine zipper (bZIP) transcription factors in plant abiotic stress. Protoplasma 254(1):1–14. https://doi.org/10.1007/s00709-015-0920-4
Basu S, Ramegowda V, Kumar A, Pereira A (2016) Plant adaptation to drought stress. F1000 Res 5:1554. https://doi.org/10.12688/f1000research.7678.1
Benjamini Y, Hochberg Y (1995) Controlling the false discovery rate: a practical and powerful approach to multiple testing. J R Statist Soc B 57:289–300
Bhanu BB, Ulaganathan K, Shanker AK, Desai S (2016) RNA-seq analysis of irrigated vs water stressed transcriptomes of Zea mays cultivar Z59. Front Plant Sci 7:239. https://doi.org/10.3389/fpls.2016.00239
Bhargava S, Sawant K (2013) Drought stress adaptation: Metabolic adjustment and regulation of gene expression. Plant Breed 132:21–32. https://doi.org/10.1111/pbr.12004
Bianchi VJ, Rubio M, Trainotti L, Verde I, Bonghi C, Martínez-Gómez P (2015) Prunus transcription factors: breeding perspectives. Front Plant Sci 6:443. https://doi.org/10.3389/fpls.2015.00443
Bradford MM (1976) A rapid and sensitive method for the quantification of microgram quantities of protein utilizing the principle of protein-dye binding. Anal Biochem 72:248–254. https://doi.org/10.1016/0003-2697(76)90527-3
Danilevskaya ON, Yu G, Meng X, Xu J, Stephenson E, Estrada S, Chilakamarri S, Hayes ZG, Thatcher S (2019) Developmental and transcriptional responses of maize to drought stress under field conditions(Article). Plant Direct 3(5):1–20. https://doi.org/10.1002/pld3.129
Darby H, Lauer J (2006) Plant physiology: critical stages in the life of a corn plant. Department of Agronomy, Winsconsin University
Dhindsa RS, Plumb-Dhindsa P, Thorpe TA (1981) Leaf senescence: correlated with increased leaves of membrane permeability and lipid peroxidation, and decreased levels of superoxide dismutase and catalase. J Exp Bot 32:93–101. https://doi.org/10.1093/jxb/32.1.93
Dudhate A, Shinde H, Tsugama D, Liu S, Takano T (2018) Transcriptomic analysis reveals the differentially expressed genes and pathways involved in drought tolerance in pearl millet [Pennisetum glaucum (L.) R. Br]. PLoS One 13:e0195908. https://doi.org/10.1371/journal.pone.0195908
Edmeades GO (2013) Progress in achieving and delivering drought tolerance in maize—an update. ISAA, Ithaca, pp 1–39
Edreva AM, Velikova VB, Tsonev TD (2007) Phenylamides in plants. Russ J Plant Physl 54(3):287–301. https://doi.org/10.1134/S1021443707030016
Fahad S, Bajwa AA, Nazir U, Anjum SA, Farooq A, Zohaib A, Sadia S, Nasim W, Adkins S, Saud S (2017) Crop production under drought and heat stress: plant responses and management options. Front Plant Sci 8:1147. https://doi.org/10.3389/fpls.2017.01147
Fang Y, Xiong L (2015) General mechanisms of drought response and their application in drought resistance improvement in plants. Cell Mol Life Sci 72:673–689. https://doi.org/10.1007/s00018-014-1767-0
Farooq M, Wahid A, Kobayashi N, Fujita D, Basra SMA (2009) Plant drought stress: effects, mechanisms and management. Agron Sustain Dev 29(1):185–212. (Springer Verlag/EDP Sciences/INRA)
Galmés J, Flexas J, Savé R, Medrano H (2007) Water relations and stomatal characteristics of Mediterranean plants with different growth forms and leaf habits: responses to water stress and recovery. Plant Soil 290:139–155. https://doi.org/10.1007/s11104-006-9148-6
Ghannoum O (2009) C4 photosynthesis and water stress. Ann Bot 103:635–644. https://doi.org/10.1093/aob/mcn093
Glawischnig E, Grun S, Frey M, Gierl A (1999) Cytochrome P450 monooxygenases of DIBOA biosynthesis: specificity and conservation among grasses. Phytochemistry 50:925–930. https://doi.org/10.1016/s0031-9422(98)00318-5
Han LB, Song GL, Zhang X (2008) Preliminary observation of physiological responses of three turfgrass species to traffic stress. Hort Technol 18(1):139–143. https://doi.org/10.21273/HORTTECH.18.1.139
Harb A (2016) Identification of candidate genes for drought stress tolerance. In: Hossain MA (ed) Drought stress tolerance in plants 2. Springer International Publishing, Berlin
Hsiao T (1973) Plant responses to water stress. Annu Rev Plant Physiol 24:519–570
Jan AU, Hadi F, Midrarullah AA, Rahman K (2017) Role of CBF/DREB gene expression in abiotic stress tolerance. Int J Hort Agric 2(1):1–12
Jin HY, Liu ST, Zenda T, Wang X, Liu G, Duan HJ (2019) Maize leaves drought-responsive genes revealed by comparative transcriptome of two cultivars during the filling stage. PLoS One 14(10):e0223786. https://doi.org/10.1371/journal.pone.0223786
Jogaiah S, Govind SR, Tran LS (2013) Systems biology-based approaches toward understanding drought tolerance in food crops. Crit Rev Biotechnol 33:23–39. https://doi.org/10.3109/07388551.2012.659174
Joshi R, Wani SH, Singh B, Bohra A, Dar ZA, Lone AA, Pareek A, Pareek SLS (2016) Transcription factors and plants response to drought stress: current understanding and future directions. Front Plant Sci 7:1029. https://doi.org/10.3389/fpls.2016.01029
Kakumanu A, Ambavaram MMR, Klumas C, Krishnan A, Batlang U, Myers E, Grene R, Pereira A (2012) Effects of drought on gene expression in maize reproductive and leaf meristem tissue revealed by RNA-seq. Plant Physiol 160:846–867. https://doi.org/10.1104/pp.112.200444
Kimotho RN, Baillo EH, Zhang ZB (2019) Transcription factors involved in abiotic stress responses in maize (Zea mays L.) and their roles in enhanced productivity in the post genomics era. Peer J 7:e7211. https://doi.org/10.7717/peerj.7211
Kumar S, Trivedi PK (2018) Glutathione S-transferases: role in combating abiotic stresses including arsenic detoxification in plants. Front Plant Sci 9:751. https://doi.org/10.3389/fpls.2018.00751
Li P, Cao W, Fang H, Xu S, Yin S, Zhang Y, Lin D, Wang J, Chen Y, Xu C, Yang Z (2017) Transcriptomic profiling of the maize (Zea mays L.) leaf response to abiotic stresses at the seedling stage. Front Plant Sci 8:290. https://doi.org/10.3389/fpls.2017.00290
Liu ST, Zenda T, Dong AY, Yang YT, Liu XY, Wang YF, Li J, Tao YS, Duan HJ (2019) Comparative proteomic and morpho-physiological analyses of maize wild-type Vp16 and mutant vp16 germinating seed responses to PEG-induced drought stress. Int J Mol Sci. https://doi.org/10.3390/ijms20225586
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2−ΔΔCT method. Methods 25:402–408. https://doi.org/10.1006/meth.2001.1262
Love MI, Huber W, Anders S (2014) Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol 15:550. https://doi.org/10.1186/s13059-014-0550-8
Lu X, Zhou X, Cao Y, Zhou M, McNeil D, Liang S, Yang C (2017) RNA-seq analysis of cold and drought responsive transcriptomes of Zea mays ssp. mexicana L. Front Plant Sci 8:136. https://doi.org/10.3389/fpls.2017.00136
Luo M, Zhao Y, Wang Y, Shi Z, Zhang P, Zhang Y, Song W, Zhao J (2018) Comparative proteomics of contrasting maize genotypes provides insights into salt-stress tolerance mechanisms. J Proteome Res 17:141–153. https://doi.org/10.1021/acs.jproteome.7b00455
Ma D, Sun D, Wang C, Li Y, Guo T (2014) Expression of flavonoid biosynthesis genes and accumulation of flavonoid in wheat leaves in response to drought stress. Plant Physiol Biochem 80:60–66. https://doi.org/10.1016/j.plaphy.2014.03.024
Maurel C, Boursiac Y, Luu DT, Santoni V, Shahzad Z, Verdoucq L (2015) Aquaporins in plants. Physiol Rev 95(4):1321–1358. https://doi.org/10.1152/physrev.00008.2015
Min H, Chen C, Wei S, Shang X, Sun M, Xia R, Liu XG, Hao DY, Chen HB, Xie Q (2016) Identification of drought tolerant mechanisms in maize seedlings based on transcriptome analysis of recombination inbred lines. Front Plant Sci 7:1080. https://doi.org/10.3389/fpls.2016.01080
Mittal S, Arora K, Ramakrishna RA, Mallikarjuna MG, Gupta HS, Nepolean T (2017) Genomic selection for drought tolerance using genome-wide SNPs in maize. Front Plant Sci 8:550. https://doi.org/10.3389/fpls.2017.00550
Mittal S, Banduni P, Mallikarjuna MG, Rao AR, Jain PA, Dash PK, Thirunavukkarasu N (2018) Structural, functional, and evolutionary characterization of major drought transcription factors families in maize. Front Chem 6:177. https://doi.org/10.3389/fchem.2018.00177
Mortazavi A, Williams BA, McCue K, Schaeffer L, Wold B (2008) Mapping and quantifying mammalian transcriptomes by RNA-Seq. Nat Meth 5:621–628. https://doi.org/10.1038/nmeth.1226
Moussa HR, Aziz SMA (2008) Comparative response of drought tolerant and drought sensitive maize genotypes to water stress. Aust J Crop Sci 1(1):31–36
Murata N, Allakhverdiev SI, Nishiyama Y (2012) The mechanism of photoinhibition in vivo: re-evaluation of the roles of catalase, α-tocopherol, non-photochemical quenching, and electron transport. Biochim Biophys 1817:1127–1133. https://doi.org/10.1016/j.bbabio.2012.02.020
Oliver SN, Dennis ES, Dolferus R (2007) ABA regulates apoplastic sugar transport and is a potential signal for cold-induced pollen sterility in rice. Plant Cell Physiol 48:1319–1330. https://doi.org/10.1093/pcp/pcm100
Opitz N, Paschohold A, Marcon C, Malik WA, Lanz C, Piepho HP, Hochholdinger F (2014) Transcriptomic complexity in young maize primary roots in response to low water potentials. BMC Genom 15:1471–2164. https://doi.org/10.1186/1471-2164-15-741
Opitz N, Marcon C, Paschold A, Malik WA, Lithio A, Brandt R, Piepho H, Nettleton D, Hochholdinger F (2016) Extensive tissue-specific transcriptomic plasticity in maize primary roots upon water deficit. J Ex Bot 67(4):1095–1107. https://doi.org/10.1093/jxb/erv453
Peterbauer T, Mucha J, Mayer U, Popp M, Glossl J, Richter A (1999) Stachyose synthesis in seeds of adzuki bean (Vigna angularis): molecular cloning and functional expression of stachyose synthase. Plant J Cell Mol Biol 20:509–518. https://doi.org/10.1046/j.1365-313X.1999.00618.x
Prasad PVV, Staggenborg SA, Ristic Z (2008) Impacts of drought and/or heat stress on physiological, developmental, growth, and yield processes of crop plants. In: Ahuja LR, Reddy VR, Saseendran SA, Qiang Yu (eds) Response of crops to limited water: understanding and modeling water stress effects on plant growth processes. Advances in Agricultural Systems Modeling Series 1 (2008), Agronomy Society of America (ASA), Crop Science Society of America (CSSA), Soil Science Society of America (SSSA), Madison, USA. https://doi.org/10.2134/advagricsystmodel1.c11
Priya M, Dhanker OP, Siddique KHM, Rao BH, Nair RM, Pandey S, Singh S, Varshney RK, Prasad PVV, Nayyar H (2019) Drought and heat stress-related proteins: an update about their functional relevance in imparting stress tolerance in agricultural crops. Theor Appl Genet Springer Verlag GmbH Germany. https://doi.org/10.1007/s00122-019-03331-2
Schils R, Olesen JE, Kersebaum K, Rijk B, Oberforster M, Kalyada V, Khitrykau M, Gobin A, Kirchev H, Manolova V, Manolov I, Trnka M, Hlavinka P, Palosuo T, Peltonen-Sainio P, Jauhiainen L, Lorgeou J, Marrou H, Danalatos N, Archontoulis S, Fodor N, Spink J, Roggero PP, Bassu S, Pulina A, Seehusen T, Uhlen AK, Żyłowska K, Nieróbca A, Si VJ (2018) Cereal yield gaps across Europe. Eur J Agron 101:109–120. https://doi.org/10.1016/j.eja.2018.09.003
Shan X, Li Y, Jiang Y, Jiang Z, Hao W, Yuan Y (2013) Transcriptome profile analysis of maize seedlings in response to high-salinity, drought and cold stresses by deep sequencing. Plant Mol Biol Rep 31:1485–1491. https://doi.org/10.1007/s11105-013-0622-z
Sharma P, Jha AB, Dubey RS, Pessarakli M (2012) Reactive oxygen species, oxidative damage, and antioxidative defense mechanism in plants under stressful conditions. J Bot 10:26. https://doi.org/10.1155/2012/217037
Shiferaw B, Prasanna BM, Hellin J, Bänziger M (2011) Crops that feed the world 6. Past successes and future challenges to the role played by maize in global food security. Food Sec 3(3):307–327. https://doi.org/10.1007/s12571-011-0140-5
Shinde H, Tanaka K, Dudhate A, Tsugama D, Mine Y, Kamiya T, Gupta SK, Liu S, Takano T (2018) Comparative de novo transcriptomic profiling of the salinity stress responsiveness in contrasting pearl millet lines. Environ Exp Bot 155:619–627. https://doi.org/10.1016/j.envexpbot.2018.07.008
Singh D, Laxmi A (2015) Transcriptional regulation of drought response: a tortuous network of transcriptional factors. Front Plant Sci 6:895. https://doi.org/10.3389/fpls.2015.00895
Singh S, Kumar V, Kapoor D, Kumar S, Singh J (2019) Revealing on hydrogen sulfide and nitric oxide signals cordination for plant growth under stress conditions. Physiol Plant. https://doi.org/10.1111/ppl.13002
Tai F, Yuan Z, Li S, Wang Q, Liu F (2016) ZmCIPK8, a CBL-interacting protein kinase, regulates maize response to drought stress. Plant Cell Tiss Org 124:59–469. https://doi.org/10.1007/s11240-015-0906-0
Thirunavukkarasu N, Sharma R, Singh N, Shiriga K, Mohan S, Mittal S, Mittal S, Mallikarjuna MG, Rao AR, Dash PK, Hossain F, Gupta HS (2017) Genomewide expression and functional interactions of genes under drought stress in maize. Int J Genet. https://doi.org/10.1155/2017/2568706. (Article ID 2568706)
Trapnell C, Williams BA, Pertea G, Mortazavi A, Kwan G, van Baren MJ, Salzberg SL, Wold BJ, Pachter L (2010) Transcript assembly and quantification by RNA-Seq reveals unannotated transcripts and isoform switching during cell differentiation. Nat Biotechnol 28:511–515. https://doi.org/10.1038/nbt.1621
Trapnell C, Roberts A, Goff L, Pertea G, Kim D, Kelley DR, Pimentel H, Salzberg SL, Rinn JL, Pachter L (2012) Differential gene and transcript expression analysis of RNA-seq experiments with TopHat and Cufflinks. Nat Protoc 7:562–578. https://doi.org/10.1038/nprot.2012.016
Wang L, Feng Z, Wang X, Wang X, Zhang X (2010) DEGseq: an R package for identifying differentially expressed genes from RNA-seq data. Bioinformatics 26:136–138. https://doi.org/10.1093/bioinformatics/btp612
Wang H, Wang H, Shao H, Tang X (2016) Recent advances in utilizing transcription factors to improve plant abiotic stress tolerance by transgenic technology. Front Plant Sci 7:67. https://doi.org/10.3389/fpls.2016.00067
Wang CT, Ru JN, Liu YW, Yang JF, Li M, Xu ZS, Fu JD (2018) The maize WRKY transcription factor ZmWRKY40 confers drought resistance in transgenic Arabidopsis. Int J Mol Sci 19:2580. https://doi.org/10.3390/ijms19092580
Wang BM, Liu C, Zhang DF, He CM, Zhang JR, Li ZX (2019a) Effects of maize organ-specific drought stress response on yields from transcriptome analysis. BMC Plant Biol 19:335. https://doi.org/10.1186/s12870-019-1941-5
Wang X, Zenda T, Liu ST, Liu G, Jin HY, Dai L, Dong AY, Yang YT, Duan HJ (2019b) Comparative proteomics and physiological analyses reveal important maize filling-kernel drought-responsive genes and metabolic pathways. Int J Mol Sci 20:3743. https://doi.org/10.3390/ijms20153743
Xie C, Mao X, Huang J, Ding Y, Wu J, Dong S, Kong L, Gao G, Li CY, Wei L (2011) KOBAS2.0: a web server for annotation and identification of enriched pathways and diseases. Nucleic Acids Res 39:W316–W322. https://doi.org/10.1093/nar/gkr483
Yang M, Geng MY, Shen PF, Chen XH, Li YJ, Wen XX (2019) Effect of post-silking drought stress on the expression profiles of genes involved in carbon and nitrogen metabolism during leaf senescence in maize (Zea mays L.). Plant Physiol Biochem 135:304–309
Zenda T, Liu ST, Wang X, Jin HY, Liu G, Duan HJ (2018) Comparative proteomic and physiological analyses of two divergent maize inbred lines provide more insights into drought-stress tolerance mechanisms. Int J Mol Sci 19:3225. https://doi.org/10.3390/ijms19103225
Zenda T, Liu ST, Wang X, Liu G, Jin HY, Dong AY, Yang YT, Duan HJ (2019) key maize drought-responsive genes and pathways revealed by comparative transcriptome and physiological analyses of contrasting inbred lines. Int J Mol Sci 20(6):1268. https://doi.org/10.3390/ijms20061268
Zhang JY, Cruz de Carvalho MH, Torres-Jerez I, Kang Y, Allen SN, Huhman DV, Tang Y, Murray J, Sumner LW, Udvardi MK (2014) Global reprogramming of transcription and metabolism in Medicago truncatula during progressive drought and after rewatering. Plant Cell Environ 37:2553–2576. https://doi.org/10.1111/pce.12328
Zhang X, Lei L, Lai J, Zhao H, Song W (2018) Effects of drought stress and water recovery on physiological responses and gene expression in maize seedlings. BMC Plant Biol 18:68. https://doi.org/10.1186/s12870-018-1281-x
Zhao F, Zhang D, Zhao Y, Wang W, Yang H, Tai F, Li C, Hu X (2016a) The difference of physiological and proteomic changes in maize leaves adaptation to drought, heat, and combined both stresses. Front Plant Sci 7:1–19. https://doi.org/10.3389/fpls.2016.01471
Zhao P, Liu P, Yuan GX, Jia JT, Li XX, Qi D, Chen SY, Ma T, Liu GS, Cheng LQ (2016b) New insights on drought stress response by global investigation of gene expression changes in sheepgrass (Leymus chinensis). Front Plant Sci 7:954. https://doi.org/10.3389/fpls.2016.00954
Zheng J, Fu JJ, Gou MY, Huai JL, Liu Y, Jian M, Huang QS, Guo XY, Dong ZG, Wang HZ, Wang GY (2010) Genome-wide transcriptome analysis of two maize inbred lines under drought stress. Plant Mol Biol 72:407–421. https://doi.org/10.1007/s11103-009-9579-6
Acknowledgements
This research was supported by the National Key Research and Development Project of China (Selection and Efficient Combination Model of Wheat and Maize Water Saving, High Yield and High Quality Varieties) (Grant No. 2017YFD0300901).
Author information
Authors and Affiliations
Contributions
GL, TZ, and HD conceived and designed the experiment; GL, TZ, SL, XW, HJ, AD and YY performed the investigations and collected data; GL, TZ, SL and XW analyzed the data; GL and TZ wrote the original manuscript; All authors made the revision of the manuscript and approved this submission.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest. Furthermore, the founding sponsors had no role in the design of the study; in the collection, analyses, or interpretation of data; in the writing of the manuscript, and in the decision to publish the results.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Liu, G., Zenda, T., Liu, S. et al. Comparative transcriptomic and physiological analyses of contrasting hybrid cultivars ND476 and ZX978 identify important differentially expressed genes and pathways regulating drought stress tolerance in maize. Genes Genom 42, 937–955 (2020). https://doi.org/10.1007/s13258-020-00962-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13258-020-00962-4