Abstract
The detection of copy-number variations (CNVs) from NGS data is underexploited as chip-based or targeted techniques are still commonly used. We assessed the performances of a workflow centered on CANOES, a bioinformatics tool based on read depth information. We applied our workflow to gene panel (GP) and whole-exome sequencing (WES) data, and compared CNV calls to quantitative multiplex PCR of short fluorescent fragments (QMSPF) or array comparative genomic hybridization (aCGH) results. From GP data of 3776 samples, we reached an overall positive predictive value (PPV) of 87.8%. This dataset included a complete comprehensive QMPSF comparison of four genes (60 exons) on which we obtained 100% sensitivity and specificity. From WES data, we first compared 137 samples with aCGH and filtered comparable events (exonic CNVs encompassing enough aCGH probes) and obtained an 87.25% sensitivity. The overall PPV was 86.4% following the targeted confirmation of candidate CNVs from 1056 additional WES. In addition, our CANOES-centered workflow on WES data allowed the detection of CNVs with a resolution of single exons, allowing the detection of CNVs that were missed by aCGH. Overall, switching to an NGS-only approach should be cost-effective as it allows a reduction in overall costs together with likely stable diagnostic yields. Our bioinformatics pipeline is available at: https://gitlab.bioinfo-diag.fr/nc4gpm/canoes-centered-workflow.
Similar content being viewed by others
Introduction
Copy-number variations (CNVs) are a major cause of Mendelian disorders [1] as well as risk factors for common diseases [2]. With the advent of next-generation sequencing (NGS), a number of software tools have been developed to detect CNVs [3,4,5]. Whole-genome sequencing (WGS) is often presented as an almost universal technique allowing the assessment of almost any type of variation, including CNVs and other structural variations [6]. WGS may eventually be used as a first-tier diagnostic tool in the context of genetically highly heterogeneous disorders. However, the detection of structural variations from data generated using the technology of short read sequencing is still associated with a number of false positives. Such events can be detected using a plethora of bioinformatics tools based on different principles, including depth of coverage (DOC) information, relative position of paired reads, split reads and De Novo Assembly [7]. Besides the development of WGS, targeted sequencing of gene panel, and whole-exome sequencing (WES) remain of primary use in many diagnostic and research laboratories. They are indeed still considered as more affordable and of easier access as they can be processed using usual informatics facilities accessible to most laboratories. Moreover, the input of WGS is questioning in disorders with low genetic heterogeneity and high phenotypic specificity. Hence, gene panels and WES remain largely used.
The detection of CNVs from exonic capture-based targeted sequencing solutions primarily relies on DOC information [8, 9]. Tools based on DOC information compare one sample with a reference, and predict deletions or duplications depending on the increase or decrease of the DOC as compared with the reference (Fig. 1). As each tool was set up and trained on a specific dataset, one of the main challenges is to evaluate the specificity and sensitivity of a given software tool on large datasets. Studies evaluating the diagnostic performances of CNV detection pipelines are scarce although they appear to be critical for their use in routine procedures [10,11,12]. In order to optimize CNV detection from NGS data, a classical approach consists in running multiple tools in parallel and then aggregate the results to keep a CNV as candidate only if multiple tools called it [13]. As it is more effective to do so with tools using different types of bioinformatics methods (DOC, split reads, etc.), this combinatory approach is most adapted when working on WGS, or at least if most of the intergenic or intronic regions—where breakends are more frequently found—are captured. Here, we decided to focus on one tool using the DOC approach as it still remains the most adapted one for exonic capture. In a precision workflow approach, we developed a workflow based on the already existing software tool CANOES [14]. To select this tool, we previously compared features of each tool that are related to the definition of a reference for CNV detection. Indeed, defining the reference is critical [15], as calling candidate deletions or duplications requires the comparison of DOC data of each sample to the reference. Our main criteria concerning the definition of reference were that (i) the tool should take into account information from multiple samples and that (ii) it should associate a Hidden Markov Model (HMM) with a distribution model to represent the variability of coverage between samples and between each target. CANOES appeared as the best candidate as it adopts a pooling strategy to build its reference model and it uses an HMM associated to a binomial negative distribution. In addition, CANOES defines the reference independently for each sample, by selecting samples with the closest mean and variance.
Schematic distribution of reads among three different samples over five sequenced exons. a The absence of any CNV. b Duplication of two exons (2 and 3). c Deletion of exon 4. In order to call those CNVs, software tools have to establish a reference. Some tools compare paired data from the same patient, e.g., tumor tissue against germline, while others build their reference from a pool of samples and then compare a given sample to this reference, as the CANOES tool used in our workflow.
We performed a diagnostic performance evaluation of this workflow regarding gene panel and WES data, in two steps. First, we compared CNV calls with a reference technique, namely a comprehensive assessment by quantitative multiplex PCR of short fluorescent fragments (QMPSF) [16] or array comparative genomic hybridization (aCGH), regarding targeted gene panel and WES data, respectively. Second, we implemented our workflow in our routine procedures and performed an additional evaluation of the positive predictive value of our CANOES-centered workflow using targeted confirmation of CNVs using an independent targeted technique.
Material and methods
Gene panel sequencing
In order to evaluate our workflow, we analyzed data from three gene panels (for detailed information, see Supplementary Table 1). Patients provided informed written consent for genetic analyses in a diagnostic setting.
Panel 1 was set up to focus on genes involved in predisposition to colorectal cancer and digestive polyposis or Li-Fraumeni syndrome [17]. This panel was implemented in two successive versions. V1 was used to sequence 11 genes in 2771 samples. V2 was used to sequence 15 genes (same 11 genes plus 4) in 549 samples. In both versions and for all genes, exons, and introns outside repeated sequences were captured.
Panel 2 also has two successive versions and was designed to focus on two clinical indications: (i) hydrocephaly (3 genes) and (ii) Cornelia de Lange syndrome and differential diagnoses (24 genes in v1, 30 in v2). In total, 320 samples were sequenced using this panel (240 with v1, 80 with v2). For this panel, introns outside repeated sequences were captured only for two genes, namely L1CAM and NIPBL.
Panel 3 was designed to focus on genes involved in nonspecific intellectual disability. It has been used to analyze 220 samples and is composed of 48 genes (coding regions only). The list of genes is available upon request.
Assessment of CNV calls from gene panel data: step 1
For the comparison with a reference technique, we used data obtained from samples for which both NGS (panel 1, v1) and comprehensive QMPSF screening data were available (n = 465). This QMSPF assessment included all 60 exons of 4 genes from this panel (APC, MSH2, MSH6, and MLH1) and was applied to all 465 samples.
Assessment of CNV calls from gene panel data: step 2
Following step 1, we implemented our CANOES-centered workflow in our routine diagnostic procedures on NGS data from all three panels (n = 3311 additional samples in total). We performed confirmations of candidate CNVs using QMPSF or multiplex ligation-dependent probe amplification (MLPA) only in samples with a CANOES call. Primers used for QMPSF screening and validation are available upon request.
Whole-exome sequencing
Patients provided informed written consent for genetic analyses either in a diagnostic or in a research setting, following the approval by respective ethics committees.
Whole exomes were sequenced in the context of diverse research and diagnostic purposes (Supplementary Table 1). Exomes were captured using Agilent SureSelect Human All Exon kits (V1, V2 V4 + UTR, V5, V5 + UTR, and V6) (Agilent technologies, Santa Clara, CA, USA). Final libraries were sequenced on an Illumina Genome Analyzer GAIIX (corresponding to exomes captured with the V1, V2, or V4UTR kit, n = 10), or on an Illumina HiSeq2000, 2500, or 4000 with paired ends, 76 or 100 bp reads (Illumina, San Diego, Ca, USA). Exome sequencing was performed in three sequencing centers: Integragen (Evry, France) (n = 6), the French National Center in Human Genomics Research (CNRGH, Evry, France) (n = 1065) and the Genome Quebec Innovation Center (Montreal, Canada) (n = 128) [18]. Exomes were all processed through the same bioinformatics pipeline following the Broad Institute Best Practices recommendations [19]. Reads were mapped to the 1000 Genomes GRCh37 build using BWA 0.7.5a. [20]. Picard Tools 1.101 (http://broadinstitute.github.io/picard/) was used to flag duplicate reads. We applied GATK [21] for short insertion and deletions (indel) realignment and base quality score recalibration. All quality checks were processed as previously described [18].
Assessment of CNV calls from WES data: step 1
For the comparison with a reference technique, we analyzed data from 147 unrelated individuals with both WES and aCGH data available.
Array CGH analysis
Oligonucleotide aCGH was performed as previously described [22]. Briefly, high-resolution aCGH analysis was performed using the 1 × 1M Human High-Resolution Discovery Microarray Kit or the 4 × 180k SurePrint G3 Human CGH Microarray kit (Agilent Technologies, Santa Clara, CA, USA), using standard recommended protocols. An in-house and sex-matched genomic DNA pool of at least ten control individuals was used as reference sample. Hybridization results were analyzed with the Agilent’s DNA-Analytics software (version 4.0.81, Agilent Technologies) or the Agilent Genomic Workbench (version 7.0, Agilent Technologies). Data were processed using the ADM-2 algorithm, with threshold set at 6.0 SD or 5.0 SD.
WES/aCGH comparison
Array CGH enables the detection of genome-wide rearrangements thanks to the measurement of the deviation of the fluorescent signal of the patient as compared with a control DNA. The number of probes depends of the type of chip that is used (here, Agilent 1M or 180k). The threshold to consider a deletion or a duplication was set to the deviation of five or three consecutive probes, respectively. This restricts the detection to CNVs of 8 or 20 kb for Agilent 1M or Agilent 180k chips, respectively, on average. On the contrary, as CANOES analysis is based on WES data, it is strictly restricted to CNVs covering exonic sequences, but it can detect CNVs as small as one single exon.
In order to combine these approaches to evaluate the sensitivity of our workflow, we filtered out CNVs located in intronic and intergenic regions exclusively from the aCGH data (and on X and Y chromosomes for the samples processed without sex-chromosome CNV calling). Moreover, as CANOES analysis is based on the calculation of a mean and variance of coverage on a given genomic region, the detection of polymorphic rearrangements is very uncertain. For that reason, we also filtered out all polymorphic CNVs from aCGH data. We defined as polymorphic a CNV that overlaps at least at 70% with CNVs reported in the Gold Standard section of the Database of Genomic Variants with a frequency superior to 1% [23].
Regarding the evaluation of the positive predictive value of our workflow, we restricted our analysis to candidate non-polymorphic CNVs detected from WES data (i) that are theoretically detectable by aCGH as they encompass at least three or five probes, depending on the chip used and (ii) that overlap with segmental duplication regions <50% of the CANOES target regions. The segmental duplication regions have been extracted from the UCSC Table browser [24] (https://genome-euro.ucsc.edu/cgi-bin/hgTables).
As most aCGH data were processed using the hg18 genome as reference, we used the lift over tool from UCSC (https://genome.ucsc.edu/cgi-bin/hgLiftOver) to establish the correspondence to hg19. If there were no lift over possibility, we manually checked genes encompassing CNVs.
Assessment of CNV calls from WES data: step 2
Following step 1, we implemented our workflow in our routine procedures. From additional 1056 WES (Supplementary Table 1), we performed targeted confirmations following the detection of candidate CNVs by CANOES using QMPSF or ddPCR [25]. We focused our confirmations on a list of 350 genes that belong to the so-called Aβ network [26], as all the samples used at this step were sequenced in the context of Alzheimer disease research. This list of genes was built thanks to literature curation on Alzheimer pathophysiology, independently of any genomic information. Candidate CNVs were selected for targeted confirmation if (i) they encompassed genes belonging to this network, and (ii) they were not polymorphic i.e., with a frequency below 1% in our dataset.
Primers used for QMPSF or ddPCR validation are available upon request.
CNV calling from NGS data using CANOES
The CANOES software tool implements an algorithm dedicated to the detection of quantitative genomic variations based on DOC information. Basically, CANOES requires DOC data for each target of the capture kit used for each of the sample that are analyzed together. It also integrates the GC content information of each target to reduce the background variability observed in high-throughput sequencing data [27]. The read depth was calculated using BEDtools [28], and the GC content was determined using the GATK suite.
CANOES builds its statistical reference model from a subset of the samples included in the same analysis (at least 30 samples are recommended). To obtain the best possible fit, CANOES selects the samples that are the most correlated to the currently analyzed sample. This allows the detection of small CNVs, but also reduces the detection susceptibility of recurrent events. CANOES uses a Hidden Markov Model to represent the variability of the DOC distribution built from the selected samples. Then, it uses the Viterbi algorithm to assign deletions, duplications or normal regions. After the calling step, a “Not Applicable” (NA) score is attributed to all CNVs from samples carrying more than 50 rearrangements. Such samples are usually characterized by higher or lower average read depth and cannot be compared with the reference model. All CNVs assigned with an NA score were thus removed from further analyses. As CANOES used the capture kit definition to detect CNVs, boundaries of events were defined by the start position of the first target and the end position of the last target detected as deviated in comparison with the model.
A CANOES-centered workflow
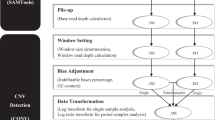
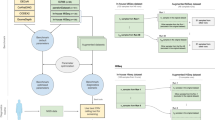
To optimize CANOES performances, we focused on two different approaches, a methodological approach in sample selection and a bioinformatics approach (Fig. 2).
File (square) with their format in parenthesis, and process (rounded) constituting the workflow. From the original capture kit definition, we merge closed target from the same exon, then do in parallel the DOC and the GC content estimation. We gather DOC individual files depending on the project, sequencing batch, unrelated samples, and remove non-informative regions. The last steps consist in CNV calling using CANOES and annotation with annotSV.
As previously described, CANOES defines a statistical model for a particular sample from a judicious selection of other samples included in the analysis. The first step of our workflow consisted in the implementation of rules to select the samples that should better be analyzed together. In order to get enough material to build an efficient statistical model and following the CANOES recommendations, we always worked with at least 30 samples. Importantly, we analyzed samples with the less technical variability from each other. Practically, this consists in analyzing samples from the same run, and not to merge multiple runs if not necessary. When merging multiple runs was inevitable (e.g., sequencing of <30 samples per run), we combined sequencing runs from the same platform and processed using the same capture kit and technical conditions, including the same number of samples per lane in order to reduce read depth variability from each sample. Of note, CANOES is not originally set up for the analysis of CNVs on sex-chromosomes, but we implemented modifications in the original script in order to include sex-chromosomes in our analyses with a modification into the output file, the copy number is replaced by a GAIN or LOSS information. Hence, we ran our workflow after gathering either N ≥ 30 males or N ≥ 30 females for the analysis of gene panels 2 and 3 that contain X-linked genes and for WES data.
Bioinformatics optimization
The first step consisted in the modification of the target definition from the capture kit information. We decided to merge close targets (<30 bp) if they covered the same exon. The only exception to this rule is if a target is larger than 1 kb. In this particular case, targets are split. Concerning gene panels that include introns, we decided to split large targets that include both intronic and exonic regions. In this case, we split targets at the intron/exon junction.
In order to gain flexibility in our analysis and to be able to add or remove samples easily, we implemented a two-step strategy consisting in (i) performing the read count step for each sample separately, and then (ii) aggregating selected samples before running CANOES. Doing so allowed, for example, intrafamilial analyses including patient–parent trio approaches, where cases can be analyzed without taking related samples into account, preventing biasing the statistical model. Finally, we removed non-informative regions from our analyses. We considered a region as non-informative if more than 90% of the samples each had <10 reads on the target. Then, we called the CNVs using CANOES, and annotated the results using AnnotSV [29] in order to get additional information about the possible effect and populations frequencies.
Nextflow integration
In order to complete our optimization of processing and analysis time, we integrated our bioinformatics pipeline into Nextflow, a data-driven workflow manager [30]. This software tool allows a quick deployment of new pipelines on different kind of computational environments, from local computers to a cloud environment. Another interest of Nextflow is to increase the performance by distributing the different steps of the workflow in regards to the computational resources available. The complete workflow, including the specific adaption of CANOES to analyze sex-chromosomes, is available on https://gitlab.bioinfo-diag.fr/nc4gpm/canoes-centered-workflow.
Interpretation of CNVs
The CNV detection workflow that is available on the above-mentioned reference finally includes the whole code required for both calling steps and annotation as a last step. Hence, the output file is a tab-delimited file that can be opened in a spreadsheet software to allow further filtration or sorting. For CNV interpretation in a Mendelian context, we prioritized CNVs based on (i) their frequencies in the DGV, potentially refined by frequencies in Exac [31], (ii) the inclusion or not of a gene from the OMIM morbid list [32] and (iii) probability of loss-of-function intolerance score (pLi) based on gnomAD database [33] and visualized CNVs in the UCSC genome browser [34].
Results
After building a workflow centered on the CANOES tool, we assessed its performances in the context of (i) gene panel NGS data and (ii) WES data, both generated following capture and Illumina short read sequencing.
Gene panel sequencing data
We first evaluated the performances of the CANOES tool using targeted sequencing data of a panel of 11 genes (panel 1, n = 465 samples). In parallel, all samples were assessed using custom comprehensive QMPSF assessing the presence or absence of a CNV encompassing any of the 60 coding exons of four of these genes. We identified 14 CNVs by QMPSF (12 deletions, 2 duplications, size range: [1,556 bp–97 kbp]). All of them were accurately detected by our CANOES-based workflow from NGS data (Table 1). In addition, no additional CNV was called by CANOES, allowing us to obtain a sensitivity and a specificity of 100% (95% CI: [73.24–100]) for those four genes. (see Supplementary Table 2).
To further assess the positive predictive value (PPV) of our workflow in the identification of CNVs from gene panels, we applied it to additional NGS data obtained from three gene panels (2222 samples from panel 1, 320 samples from panel 2, and 220 samples from panel 3). We detected 101 candidate CNVs in 98 samples and assessed their presence using either QMPSF or MLPA (Table 2). We validated 87/101 CNVs (86.13%, 95% CI: [77.50–91.94], false-positive rate: 13.9%). Overall, the PPV of our workflow applied to gene panel sequencing data was 87.83% (95% CI: [80.01–92.94]). True positive calls of our workflow were 71 deletions (size range: [391 bp–1.06 Mbp]) and 16 duplications (size range: [360 bp–39.4 kbp]) (see Supplementary Table 3). False positives were mainly deletions (10/14) and five of them were monoexonic.
Whole-exome sequencing data
We then evaluated the performances of our workflow for the detection of CNVs from WES data. We first applied our workflow to the data obtained from 147 samples with both WES (average DOC = 110×) and aCGH data available (50 samples assessed with the Agilent 1M chip and 97 samples with the Agilent 180k chip). Overall, ten samples were removed due to a high or low number of rearrangements detected by aCGH or exome, mostly due to low DNA quality or low coverage in WES.
From aCGH data, we detected 1873 CNVs over the 137 samples remaining, of which 102 were non-polymorphic exonic CNVs. Our workflow accurately detected 89 (87.2%) of them (Table 1 and Supplementary Table 4). Among the CNVs that were missed by our workflow, seven were large CNVs (from 14 to 80 kb) that encompassed only one (n = 5) or two (n = 2) targets defined by the capture kit (see Fig. 3).
A CNV (highlight region) detected by aCGH encompassing multiple CGH probes (1M probes array, in gray) but only one target from the SureSelect V5 capture kit. Of note, this deletion would have been missed by using a 180k probes array CGH (in black). View extract from UCSC genome Browser (https://genome-euro.ucsc.edu/cgi-bin/hgTracks).
In order to determine the PPV of our workflow from WES data, we selected 223 CNVs called by our workflow and (i) theoretically detectable by aCGH as encompassing at least three (180k chips) or five (1M chips) probes and (ii) which did not overlap with segmental duplication regions for more than 50% of the CANOES targets. Of them, 190 (85.2%) CNVs were confirmed as true positives following aCGH data assessment (Table 1 and Supplementary Table 5).
Of note, an additional set of 519 candidate CNVs were detected by our CANOES-based workflow that overlapped <50% of segmental duplication regions but encompassed <3 (180k chips) or 5 aCGH probes (1M chips). Hence, they were not reported by the CGH analysis tool and would then have been overlooked following classical aCGH data analysis (see Fig. 4). We did not perform targeted confirmation of all these candidate CNVs. Instead, with the aim to further assess the PPV of our workflow regarding exonic non-polymorphic CNVs of any size, we applied it to 1056 additional WES performed in the context of Alzheimer disease research (with no corresponding aCGH data). We selected non-polymorphic CNVs targeting 355 genes belonging to the Aβ network involved in the pathophysiology of Alzheimer disease [26], whatever their size. We validated 111/125 candidate CNVs (88.8%, false-positive rate: 11.2%) by QMPSF [35] or ddPCR (Table 2 and Supplementary Table 6). True positive calls of our workflow were 39 deletions (size range: [165 bp–24.2 Mbp]) and 69 duplications (size range [166 bp–5.9 Mbp]). Interestingly, among the 125 candidate CNVs obtained from our workflow, 78 were considered to be theoretically detectable by aCGH 1M, and 47 were considered as not detectable by aCGH 1M. Among the ones theoretically detectable by aCGH, 74 were true positives (94.9%). Among the theoretically not detectable ones, 37 were true positives (78.7%).
a The highlighted region represents the CNV called by the CANOES-centered workflow, encompassing one exon of RHCE. b View of the same region from DNA-Analytics (aCGH data 1M) in the same patient. This deletion was not called following aCGH data analysis as the number of deviated probes did not reach the threshold for calling. However, as three probes (in white) were deviated, this allows the confirmation of the deletion of the region. View extract from UCSC genome Browser (https://genome-euro.ucsc.edu/cgi-bin/hgTracks).
Overall, the PPV of our CANOES-based workflow was 86.49% from WES data after taking into account results from step 1 and step 2 altogether.
Discussion
Multiple tools have been developed to detect CNVs from NGS data. As long as such tools are being implemented in diagnostic laboratories, there is a critical need to evaluate their performances. Previous studies showed a large diversity of performances, while a number was performed using simulated datasets [5]. For example, from gene panel data, the DeCON tool [11] reached an overall sensitivity of 93% on a cancer gene panel sequencing of 94 genes, with a 100% sensitivity and 99% specificity on BRCA1 and 2 genes and, with a 96% PPV on the complete gene panel after validation by MLPA. Another study on 60,000 samples focusing on a panel of 48 genes reached a 100% sensitivity with a PPV of 63.2% compared with array CGH and MLPA [36]. From WES data, a study analyzing 1017 samples with XHMM obtained 67% of sensitivity and 15.76% PPV on rare CNVs compared with SNP array [37], while a comparative study of three tools [15] on 861 WES samples revealed an important diversity of sensitivity (from 20 to 75%) and PPV (from 20 to near 100%).
After having defined a CANOES-centered workflow, we applied it to three different gene panels and WES data. Overall, we reached very high detection performances following the comparison with independent techniques.
From gene panel data, we obtained a 100% sensitivity among a set of four genes, the copy number of all coding exons of which having been assessed prior to NGS in 465 samples. In addition, we obtained a 87.83% PPV among all genes with a CANOES call. Such high performances have previously been reported for other tools applied to small NGS panels [11]. Among 14 false positives, we observed recurrent events, which can be easily reported as so and be ignored in further analyses. We also observed false positive CNVs in regions homologous to pseudogenes. In that case, it is possible to reduce false positive calls by improving the design of the capture to reduce the chance that probes target the homologous regions, or by optimizing the alignment.
Of note, for all genes of Panel 1 and two genes of Panel 2, introns were captured in addition to exons. This might have increased the chances to detect CNVs that can be considered as small from an exon-only point of view but that can actually be much larger at the genomic level. An advantage of capturing introns might indeed be a gain in statistical power for the normalization process: increasing the number of targets may increase the robustness of the model. Among 101 CNVs detected from NGS data from all three panels, 75 CNVs encompassed one of these genes with intronic-plus-exonic capture. Interestingly, only 18 of these 75 CNVs encompassed a single coding exon. Such a frequency of monoexonic CNVs is not unexpected regarding mutation screens in MMR genes, in which monoexonic deletions account for 26.92–46.27% of all pathogenic deletions [38,39,40], or other rare diseases [41,42,43,44], for example. We hypothesize that all other CNVs, encompassing multiple targets, would probably have been easily detected, had the introns been excluded from the capture design. Further analyses may be required to better assess the performances of our workflow from single exon CNVs and the effect of including introns or not in the capture design. The observed higher rate of false positives in CNV calls encompassing genes without introns captured (22.22%) may also require further assessments,
We used here a precision workflow approach, focusing on the optimization of one tool based on DOC. Interestingly, as some of our genes included noncoding sequences in gene panels, these specific exonic-plus-intronic captures could provide us the possibility to apply complementary tools using different approaches, like the ones developed for WGS. This can indeed increase both detection performances of CNVs and the spectrum of structural variants that can be detectable in these data.
Of note, all our panels included multiple genes. We do not expect that a design including a single gene, even with its intronic sequences, would reach the sufficient number of targets for CANOES to build a robust model.
We also applied our workflow to multiple WES datasets and reached an overall PPV of 86.49% (95% CI: [82.34–89.81]). As for gene panel CNV detection, a confirmation by an independent technique is hence still required following the detection of a candidate CNV from WES data, although the low false-positive rate that we show here is expected to be associated with a limited number of molecular confirmations. One of the major features usually required to apply a new technique in a diagnostic workflow is a high sensitivity as compared with a reference technique. Here, we reached a sensitivity of 87.25% (95% CI: [78.84–82.77]). It should be noted that, after visualization on UCSC, five CNVs undetected by CANOES were located in polymorphic regions according to DGV or DGV gold standard tracks but not excluded from our comparison. This point can be explained by a different size between these CNVs in DGV and the one we called, so the criterion of 70% overlap required to consider them as identical was not fulfilled. Thus, this criterion led us to include some polymorphic CNVs in the comparison and hence could underestimate the sensitivity of our workflow. We still chose to keep this parameter as initially set up, because it is widely used in the literature and remains essential for a standardized analysis. Although the sensitivity was not 100%, it is important to notice that aCGH is considered as reference here although the spectrum of events that can be detected is still limited. When comparing our results with aCGH data, it appeared that we missed fewer events than the potential number of true positive CNVs that were missed by aCGH itself. Indeed, from aCGH data, we missed 13 CNVs, but our analyses called 519 candidate CNVs from corresponding WES data and which were theoretically undetectable by aCGH (i.e., either small CNVs or in regions with no aCGH probes coverage). Our PPVs suggest that the vast majority are eventually true. There is no reason to think that some of the CNVs detected by CANOES only might not be as or more deleterious than CNVs detected by both techniques or exclusively by aCGH. Knowing that aCGH misses many CNVs, even using the high-sensitivity chips such as the Agilent 1M one, and even if other chip designs might increase aCGH performances on coding regions, switching to a WES-only approach for CNV detection in a diagnostic setting should not reduce the overall diagnostic yield. Indeed, pathogenicity of CNVs cannot rely only on the size of CNVs as the deletion of a single coding exon in a gene can be sufficient to cause a Mendelian disorder. For example, we previously detected a single exon deletion that was not detectable by array CGH and was clearly pathogenic [42]. In addition, we expect that switching to a WES-only approach for CNV detection could be associated with reduced costs by skipping the CNV screen step by array technologies, although we did not perform cost-effectiveness analyses here.
As compared with aCGH, CANOES allowed the identification of CNVs of any size in regions not covered by probes but also for small CNVs including few exons. In addition, it is important to notice that the majority of CANOES false negatives were also CNVs with only few exons, which implies few targets for CANOES although noncoding probes may help detect some of them by aCGH. This decreased rate of detection of CNVs encompassing few targets has already been shown in other datasets [4, 12] and appears as a limitation inherent to DOC comparison methods.
Interestingly, CANOES allowed the detection of two mosaic rearrangements out of WES data: an SLC30A3 duplication and a 24 Mb CNV corresponding to a chromosome 20-long arm deletion (Supplementary Table 6). QMPSF data indicated that both CNVs were indeed confirmed albeit with ratios outside the ranges expected for germline events. Those examples highlight the capacity of CANOES to detect mosaic rearrangements, although the tool does not indicate such a feature, which can only be identified following the use of a targeted technique. Of note, the chromosome 20-long arm deletion was detected in a healthy control. This kind of postzygotic rearrangement is not rare in aging people (0.1% after 50 years old) [45]. Those examples highlight the capacity of CANOES to detect mosaic rearrangement.
Beyond the above-mentioned limitations of CNV detection tools from NGS data, somatic CNVs remain a challenge, both for array-based technologies and for NGS-based tools [10]. Among the CNVs detected by our workflow, at least one was considered as likely somatic, as suggested by QMPSF data. However, the sensitivity of DOC tools might remain low in this context [10].
Of note, it is possible to increase the detection of small events or events in complex regions by using the “GenotypeCNV” function of CANOES. The aim of this function is to look precisely at specific regions and call the genotype of the sample for these specific regions, however it is associated with an increase in false positive calls [42], as well as an increase in time and computational resources needed. In particular cases, when known core genes have already been identified in a given disorder, it is possible to combine our approach to call CNVs at the exome level and focus on specific genes using the GenotypeCNV function applied to every exon of these genes to increase the detection performances in core genes at the same time.
In conclusion, we performed an evaluation of the performances of a CNV detection workflow based on read depth comparison from capture-prepared NGS data, one of the most popular methods for NGS in research and diagnostic settings. We highlighted very high sensitivity and positive predictive value, for both NGS gene panel and WES. Although the sensitivity was not perfect for WES data as compared with aCGH, a number of additional true calls were not detected by the so-called reference technique. This highlights the absence of a genuine gold standard up to now. Overall, we consider that switching to an NGS-only approach is cost-effective as it allows a reduction in overall costs together with likely stable diagnostic yields.
References
Itsara A, Wu H, Smith JD, Nickerson DA, Romieu I, London SJ, et al. De novo rates and selection of large copy number variation. Genome Res. 2010;20:1469–81.
Huguet G, Schramm C, Douard E, Jiang L, Labbe A, Tihy F, et al. Measuring and estimating the effect sizes of copy number variants on general intelligence in community-based samples. JAMA Psychiatry. 2018;75:447–57.
Tan R, Wang Y, Kleinstein SE, Liu Y, Zhu X, Guo H, et al. An evaluation of copy number variation detection tools from whole-exome sequencing data. Hum Mutat. 2014;35:899–907.
Samarakoon PS, Sorte HS, Kristiansen BE, Skodje T, Sheng Y, Tjønnfjord GE, et al. Identification of copy number variants from exome sequence data. BMC Genomics. 2014;15:661.
Roca I, González-Castro L, Fernández H, Couce ML, Fernández-Marmiesse A. Free-access copy-number variant detection tools for targeted next-generation sequencing data. Mutat Res. 2019;779:114–25.
Mahmoud M, Gobet N, Cruz-Dávalos DI, Mounier N, Dessimoz C, Sedlazeck FJ. Structural variant calling: the long and the short of it. Genome Biol. 2019;20:246.
Hehir-Kwa JY, Pfundt R, Veltman JA. Exome sequencing and whole genome sequencing for the detection of copy number variation. Expert Rev Mol Diagn. 2015;15:1023–32.
Boeva V, Popova T, Bleakley K, Chiche P, Cappo J, Schleiermacher G, et al. Control-FREEC: a tool for assessing copy number and allelic content using next-generation sequencing data. Bioinformatics. 2012;28:423–5.
Krumm N, Sudmant PH, Ko A, O’Roak BJ, Malig M, Coe BP, et al. Copy number variation detection and genotyping from exome sequence data. Genome Res. 2012;22:1525–32.
Zare F, Dow M, Monteleone N, Hosny A, Nabavi S. An evaluation of copy number variation detection tools for cancer using whole exome sequencing data. BMC Bioinformatics. 2017;18:286.
Fowler A, Mahamdallie S, Ruark E, Seal S, Ramsay E, Clarke M, et al. Accurate clinical detection of exon copy number variants in a targeted NGS panel using DECoN. Wellcome Open Res. 2016;1:20.
Miyatake S, Koshimizu E, Fujita A, Fukai R, Imagawa E, Ohba C, et al. Detecting copy-number variations in whole-exome sequencing data using the eXome Hidden Markov Model: an ‘exome-first’ approach. J Hum Genet. 2015;60:175–82.
Collins RL, Brand H, Karczewski KJ, Zhao X, Alföldi J, Khera AV, et al. An open resource of structural variation for medical and population genetics. Genomics. 2019. https://doi.org/10.1101/578674.
Backenroth D, Homsy J, Murillo LR, Glessner J, Lin E, Brueckner M, et al. CANOES: detecting rare copy number variants from whole exome sequencing data. Nucleic Acids Res. 2014;42:e97.
Kuśmirek W, Szmurło A, Wiewiórka M, Nowak R, Gambin T. Comparison of kNN and k-means optimization methods of reference set selection for improved CNV callers performance. BMC Bioinformatics. 2019;20:266.
Charbonnier F, Raux G, Wang Q, Drouot N, Cordier F, Limacher JM, et al. Detection of exon deletions and duplications of the mismatch repair genes in hereditary nonpolyposis colorectal cancer families using multiplex polymerase chain reaction of short fluorescent fragments. Cancer Res. 2000;60:2760–3.
Baert-Desurmont S, Coutant S, Charbonnier F, Macquere P, Lecoquierre F, Schwartz M, et al. Optimization of the diagnosis of inherited colorectal cancer using NGS and capture of exonic and intronic sequences of panel genes. Eur J Hum Genet EJHG. 2018;26:1597–602.
Le Guennec K, Nicolas G, Quenez O, Charbonnier C, Wallon D, Bellenguez C, et al. ABCA7 rare variants and Alzheimer disease risk. Neurology. 2016;86:2134–7.
DePristo MA, Banks E, Poplin R, Garimella KV, Maguire JR, Hartl C, et al. A framework for variation discovery and genotyping using next-generation DNA sequencing data. Nat Genet. 2011;43:491–8.
Li H, Durbin R. Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics. 2009;25:1754–60.
McKenna A, Hanna M, Banks E, Sivachenko A, Cibulskis K, Kernytsky A, et al. The Genome Analysis Toolkit: a MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res. 2010;20:1297–303.
Rovelet-Lecrux A, Deramecourt V, Legallic S, Maurage C-A, Le Ber I, Brice A, et al. Deletion of the progranulin gene in patients with frontotemporal lobar degeneration or Parkinson disease. Neurobiol Dis. 2008;31:41–5.
MacDonald JR, Ziman R, Yuen RKC, Feuk L, Scherer SW. The Database of Genomic Variants: a curated collection of structural variation in the human genome. Nucleic Acids Res. 2014;42:D986–92.
Karolchik D, Hinrichs AS, Furey TS, Roskin KM, Sugnet CW, Haussler D, et al. The UCSC Table Browser data retrieval tool. Nucleic Acids Res. 2004;32:D493–6.
Cassinari K, Quenez O, Joly-Hélas G, Beaussire L, Le Meur N, Castelain M, et al. A simple, universal, and cost-efficient digital PCR method for the targeted analysis of copy number variations. Clin Chem. 2019;65:1153–60.
Campion D, Pottier C, Nicolas G, Le Guennec K, Rovelet-Lecrux A. Alzheimer disease: modeling an Aβ-centered biological network. Mol Psychiatry. 2016;21:861–71.
Benjamini Y, Speed TP. Summarizing and correcting the GC content bias in high-throughput sequencing. Nucleic Acids Res. 2012;40:e72.
Quinlan AR, Hall IM. BEDTools: a flexible suite of utilities for comparing genomic features. Bioinformatics. 2010;26:841–2.
Geoffroy V, Herenger Y, Kress A, Stoetzel C, Piton A, Dollfus H, et al. AnnotSV: an integrated tool for structural variations annotation. Berger B, editor. Bioinformatics. 2018. https://doi.org/10.1093/bioinformatics/bty304/4970516.
Di Tommaso P, Chatzou M, Floden EW, Barja PP, Palumbo E, Notredame C. Nextflow enables reproducible computational workflows. Nat Biotechnol. 2017;35:316–9.
Exome Aggregation Consortium, Ruderfer DM, Hamamsy T, Lek M, Karczewski KJ, Kavanagh D, et al. Patterns of genic intolerance of rare copy number variation in 59,898 human exomes. Nat Genet. 2016;48:1107–11.
Amberger JS, Bocchini CA, Schiettecatte F, Scott AF, Hamosh A. OMIM.org: Online Mendelian Inheritance in Man (OMIM®), an online catalog of human genes and genetic disorders. Nucleic Acids Res. 2015;43:D789–98.
Karczewski KJ, Francioli LC, Tiao G, Cummings BB, Alföldi J, Wang Q, et al. Variation across 141,456 human exomes and genomes reveals the spectrum of loss-of-function intolerance across human protein-coding genes. Genomics. 2019. https://doi.org/10.1101/531210.
Kent WJ, Sugnet CW, Furey TS, Roskin KM, Pringle TH, Zahler AM, et al. The human genome browser at UCSC. Genome Res. 2002;12:996–1006.
Le Guennec K, Quenez O, Nicolas G, Wallon D, Rousseau S, Richard A-C, et al. 17q21.31 duplication causes prominent tau-related dementia with increased MAPT expression. Mol Psychiatry. 2017;22:1119–25.
Mu W, Li B, Wu S, Chen J, Sain D, Xu D, et al. Detection of structural variation using target captured next-generation sequencing data for genetic diagnostic testing. Genet Med Off J Am Coll Med Genet. 2019;21:1603–10.
Fromer M, Moran JL, Chambert K, Banks E, Bergen SE, Ruderfer DM, et al. Discovery and statistical genotyping of copy-number variation from whole-exome sequencing depth. Am J Hum Genet. 2012;91:597–607.
Di Fiore F, Charbonnier F, Martin C, Frerot S, Olschwang S, Wang Q, et al. Screening for genomic rearrangements of the MMR genes must be included in the routine diagnosis of HNPCC. J Med Genet. 2004;41:18–20.
Taylor CF, Charlton RS, Burn J, Sheridan E, Taylor GR. Genomic deletions in MSH2 or MLH1 are a frequent cause of hereditary non-polyposis colorectal cancer: identification of novel and recurrent deletions by MLPA. Hum Mutat. 2003;22:428–33.
van der Klift H, Wijnen J, Wagner A, Verkuilen P, Tops C, Otway R, et al. Molecular characterization of the spectrum of genomic deletions in the mismatch repair genes MSH2, MLH1, MSH6, and PMS2 responsible for hereditary nonpolyposis colorectal cancer (HNPCC). Genes Chromosomes Cancer. 2005;44:123–38.
Baker M, Strongosky AJ, Sanchez-Contreras MY, Yang S, Ferguson W, Calne DB, et al. SLC20A2 and THAP1 deletion in familial basal ganglia calcification with dystonia. Neurogenetics. 2014;15:23–30.
David S, Ferreira J, Quenez O, Rovelet-Lecrux A, Richard A-C, Vérin M, et al. Identification of partial SLC20A2 deletions in primary brain calcification using whole-exome sequencing. Eur J Hum Genet EJHG. 2016;24:1630–4.
Guo X-X, Su H-Z, Zou X-H, Lai L-L, Lu Y-Q, Wang C, et al. Identification of SLC20A2 deletions in patients with primary familial brain calcification. Clin Genet. 2019;96:53–60.
Nicolas G, Rovelet-Lecrux A, Pottier C, Martinaud O, Wallon D, Vernier L, et al. PDGFB partial deletion: a new, rare mechanism causing brain calcification with leukoencephalopathy. J Mol Neurosci MN. 2014;53:171–5.
Machiela MJ, Zhou W, Caporaso N, Dean M, Gapstur SM, Goldin L, et al. Mosaic chromosome 20q deletions are more frequent in the aging population. Blood Adv. 2017;1:380–5.
Acknowledgements
This study received fundings from Clinical Research Hospital Program from the French Ministry of Health (GMAJ, PHRC 2008/067), the JPND PERADES and France Génomique. This study was co-supported by the Centre National de Référence Malades Alzheimer Jeunes (CNR-MAJ), European Union and Région Normandie. Europe gets involved in Normandie with the European Regional Development Fund (ERDF).
Collaborators
FREX Consortium
Principal Investigators: Emmanuelle Génin5, Dominique Campion1,4, Jean-François Dartigues6, Jean-François Deleuze3, Jean-Charles Lambert7, Richard Redon8
Bioinformatics group: Thomas Ludwig5, Benjamin Grenier-Boley7, Sébastien Letort5, Pierre Lindenbaum5, Vincent Meyer3, Olivier Quenez1
Statistical genetics group: Christian Dina8, Céline Bellenguez7, Camille Charbonnier1, Joanna Giemza8
Data collection: Stéphanie Chatel7, Claude Férec5, Hervé Le Marec7, Luc Letenneur6, Gaël Nicolas1, Karen Rouault5
Sequencing: Delphine Bacq3, Anne Boland3, Doris Lechner3
Author information
Authors and Affiliations
Consortia
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Members of the FREX Consortium are listed below Acknowledgements.
Supplementary information
supplementary table 1 : Summary of datasets used for the evaluation of the CANOES-centered workflow.
41431_2020_672_MOESM2_ESM.xls
supplementary table 2: Summary of CNVs used in step 1 (comparison to reference technique) analysis from Gene Panel Sequencing
41431_2020_672_MOESM3_ESM.xls
Supplementary table 3: Summary of candidate CNVs called by our CANOES-centered workflow from Gene Panel Sequencing Data.
41431_2020_672_MOESM4_ESM.xls
Supplementary table 4 : summary of all CNVs detected from aCGH and technically detectable by our CANOES-centered workflow
Rights and permissions
About this article
Cite this article
Quenez, O., Cassinari, K., Coutant, S. et al. Detection of copy-number variations from NGS data using read depth information: a diagnostic performance evaluation. Eur J Hum Genet 29, 99–109 (2021). https://doi.org/10.1038/s41431-020-0672-2
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41431-020-0672-2
This article is cited by
-
Novel JAG1 variants leading to Alagille syndrome in two Chinese cases
Scientific Reports (2024)
-
Exome first approach to reduce diagnostic costs and time – retrospective analysis of 111 individuals with rare neurodevelopmental disorders
European Journal of Human Genetics (2022)
-
Genetic abnormalities in biopsy-proven, adult-onset hemolytic uremic syndrome and C3 glomerulopathy
Journal of Molecular Medicine (2022)