Abstract
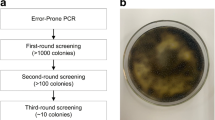
β-Glucosidase (BGL) plays a key role in cellulose hydrolysis. However, it is still a great challenge to enhance product tolerance and enzyme activity of BGL simultaneously. Here, we utilized one round error-prone PCR to engineer the Penicillium oxalicum 16 BGL (16BGL) for improving the cellulosic ethanol yield. We identified a new variant (L-6C), a triple mutant (M280T/V484L/D589E), with enhanced catalytic efficiency (\({k}_{cat}/{K}_{m}\)) for hydrolyzing pNPG and reduced strength of inhibition (\({K}_{m}^{app}/{K}_{I}\)) by glucose. To be specific, L-6C achieved a \({K}_{m}^{app}/{K}_{I}\) of 0.35 at a glucose concentration of 20 mM, which was 3.63 times lower than that attained by 16BGL. The catalytic efficiency for L-6C to hydrolyze pNPG was determined to be 983.68 mM−1 s−1, which was 22% higher than that for 16BGL. However, experiments showed that L-6C had reduced binding affinity (2.88 mM) to pNGP compared with 16BGL (1.69 mM). L-6C produced 6.15 g/L ethanol whose yield increased by about 10% than 16BGL. We performed molecular docking and molecular dynamics (MD) simulation, and binding free energy calculation using the Molecular Mechanics/Poisson Boltzmann surface area (MM/PBSA) method. MD simulation together with the MM/PBSA calculation suggested that L-6C had reduced binding free energy to pNPG, which was consistent with the experimental data.





Similar content being viewed by others
References
Abraham MJ, Murtola T, Schulz R et al (2015) GROMACS: high performance molecular simulations through multi-level parallelism from laptops to supercomputers. SoftwareX 1:19–25
Agirre J, Ariza A, Offen WA et al (2016) Three-dimensional structures of two heavily N-glycosylated Aspergillus sp. family GH3 β-d-glucosidases. Acta Crystallogr D 72:254–265
Arnold FH (1998) Design by directed evolution. Acc Chem Res 31:125–131
Basit A, Tajwar R, Sadaf S et al (2019) Improvement in activity of cellulase Cel12A of Thermotoga neapolitana by error prone PCR. J Biotechnol 306:118–124
Best RB, Zhu X, Shim J et al (2012) Optimization of the additive CHARMM all-atom protein force field targeting improved sampling of the backbone ϕ, ψ and side-chain χ1 and χ2 dihedral angles. J Chem Theory Comput 8:3257–3273
Biasini M, Bienert S, Waterhouse A et al (2014) SWISS-MODEL: modelling protein tertiary and quaternary structure using evolutionary information. Nucleic Acids Res 42:W252–W258
Bruice TC (2002) A view at the millennium: the efficiency of enzymatic catalysis. Acc Chem Res 35:139–148
Bruice TC, Benkovic SJ (2000) Chemical basis for enzyme catalysis. Biochemistry 39:6267–6274
Bussi G, Donadio D, Parrinello M (2007) Canonical sampling through velocity rescaling. J Chem Phys 126:014101
Czjzek M, Cicek M, Zamboni V et al (2000) The mechanism of substrate (aglycone) specificity in β-glucosidases is revealed by crystal structures of mutant maize β-glucosidase-DIMBOA, -DIMBOAGlc, and -dhurrin complexes. Proc Natl Acad Sci 97:13555–13560
Davies G, Henrissat B (1995) Structures and mechanisms of glycosyl hydrolases. Structure 3:853–859
Essmann U, Perera L, Berkowitz ML et al (1995) A smooth particle mesh Ewald method. J Chem Phys 103:8577–8593
Gudmundsson M, Hansson H, Karkehabadi S et al (2016) Structural and functional studies of the glycoside hydrolase family 3 β-glucosidase Cel3A from the moderately thermophilic fungus Rasamsonia emersonii. Acta Crystallogr D 72:860–870
He J, Huang X, Xue J et al (2018) Computational redesign of penicillin acylase for cephradine synthesis with high kinetic selectivity. Green Chem 20:5484–5490
Hess B, Bekker H, Berendsen HJ et al (1997) LINCS: a linear constraint solver for molecular simulations. J Comput Chem 18:1463–1472
Huang X, Han K, Zhu Y (2013a) Systematic optimization model and algorithm for binding sequence selection in computational enzyme design. Protein Sci 22:929–941
Huang X, Yang J, Zhu Y (2013b) A solvated ligand rotamer approach and its application in computational protein design. J Mol Model 19:1355–1367
Huang X, Xue J, Zhu Y (2017) Computational design of cephradine synthase in a new scaffold identified from structural databases. Chem Commun 53:7604–7607
Huang X, Xue J, Lin M et al (2016) Use of an improved matching algorithm to select scaffolds for enzyme design based on a complex active site model. PLoS ONE 11:e0156559
Jimenez-Oses G, Osuna S, Gao X et al (2014) The role of distant mutations and allosteric regulation on LovD active site dynamics. Nat Chem Biol 10:431–436
Jing X, Zhang X, Bao J (2009) Inhibition performance of lignocellulose degradation products on industrial cellulase enzymes during cellulose hydrolysis. Appl Biochem Biotechnol 159:696
Jorgensen WL, Chandrasekhar J, Madura JD et al (1983) Comparison of simple potential functions for simulating liquid water. J Chem Phys 79:926–935
Kan SJ, Lewis RD, Chen K et al (2016) Directed evolution of cytochrome c for carbon–silicon bond formation: bringing silicon to life. Science 354:1048–1051
Khalil HA, Bhat A, Yusra AI (2012) Green composites from sustainable cellulose nanofibrils: a review. Carbohydr Polym 87:963–979
Kiss G, Celebi-Olcum N, Moretti R et al (2013) Computational enzyme design. Angew Chem Int Ed Engl 52:5700–5725
Kumari R, Kumar R, Consortium OSDD et al (2014) g_mmpbsa—a GROMACS tool for high-throughput MM-PBSA calculations. J Chem Inf Model 54:1951–1962
Li Q, Huang X, Zhu Y (2014) Evaluation of active designs of cephalosporin C acylase by molecular dynamics simulation and molecular docking. J Mol Model 20:2314
Li H, Yi S, Bell EW et al (2019) Recombinant Penicillium oxalicum 16 β-glucosidase 1 displays comprehensive inhibitory resistance to several lignocellulose pretreatment products, ethanol, and salt. Appl Biochem Biotechnol. https://doi.org/10.1007/s12010-019-03183-y
Liu L, Li X, Wang J et al (2017) Two distant catalytic sites are responsible for C2c2 RNase activities. Cell 168:121–134
Mascal M, Nikitin EB (2008) Direct, high-yield conversion of cellulose into biofuel. Angew Chem Int Ed 47:7924–7926
Miller GL, Blum R, Glennon WE et al (1960) Measurement of carboxymethylcellulase activity. Anal Biochem 1:127–132
Morris GM, Huey R, Lindstrom W et al (2009) AutoDock4 and AutoDockTools4: automated docking with selective receptor flexibility. J Comput Chem 30:2785–2791
Parrinello M, Rahman A (1981) Polymorphic transitions in single crystals: a new molecular dynamics method. J Appl Phys 52:7182–7190
Qu G, Li A, Sun Z et al (2020) The crucial role of methodology development in directed evolution of selective enzymes. Angew Chem Int Ed 59:2–30
Sterling T, Irwin JJ (2015) ZINC 15—ligand discovery for everyone. J Chem Inf Model 55:2324–2337
Sterner R, Merkl R, Raushel FM (2008) Computational design of enzymes. Chem Biol 15:421–423
Tian Y, Huang X, Li Q et al (2017a) Computational design of variants for cephalosporin C acylase from Pseudomonas strain N176 with improved stability and activity. Appl Microbiol Biotechnol 101:621–632
Tian Y, Xu Z, Huang X et al (2017b) Computational design to improve catalytic activity of cephalosporin C acylase from Pseudomonas strain N176. RSC Adv 7:30370–30375
Trott O, Olson AJ (2010) AutoDock Vina: improving the speed and accuracy of docking with a new scoring function, efficient optimization, and multithreading. J Comput Chem 31:455–461
Vanommeslaeghe K, Hatcher E, Acharya C et al (2010) CHARMM general force field: a force field for drug-like molecules compatible with the CHARMM all-atom additive biological force fields. J Comput Chem 31:671–690
Wang Y, Zhou Y, Shi S et al (2020) A rational design for improving the pepsin resistance of cellulase E4 isolated from T. fusca based on the evaluation of the transition complex and molecular structure. Biochem Eng J 153:107417
Xue J, Huang X, Lin M et al (2016) A fast loop-closure algorithm to accelerate residue matching in computational enzyme design. J Mol Model 22:49
Xue J, Huang X, Zhu Y (2019) Using molecular dynamics simulations to evaluate active designs of cephradine hydrolase by molecular mechanics/Poisson–Boltzmann surface area and molecular mechanics/generalized Born surface area methods. RSC Adv 9:13868–13877
Yang J, Roy A, Zhang Y (2013) Protein–ligand binding site recognition using complementary binding-specific substructure comparison and sequence profile alignment. Bioinformatics 29:2588–2595
Yang Y, Zhang X, Yin Q et al (2015) A mechanism of glucose tolerance and stimulation of GH1 β-glucosidases. Sci Rep 5:1–12
Yi S, Zhang X, Li H et al (2018) Screening and mutation of Saccharomyces cerevisiae UV-20 with a high yield of second generation bioethanol and high tolerance of temperature, glucose and ethanol. Indian J Microbiol 58:440–447
Yin Q, Zhou G, Peng C et al (2019) The first fungal laccase with an alkaline pH optimum obtained by directed evolution and its application in indigo dye decolorization. AMB Express 9:151
Yu W, He X, Vanommeslaeghe K et al (2012) Extension of the CHARMM general force field to sulfonyl-containing compounds and its utility in biomolecular simulations. J Comput Chem 33:2451–2468
Zhang Y-HP, Himmel ME, Mielenz JR (2006) Outlook for cellulase improvement: screening and selection strategies. Biotechnol Adv 24:452–481
Zhao X, Wang W, Wang F et al (2012) A comparative study of β-1, 4-endoglucanase (possessing β-1, 4-exoglucanase activity) from Bacillus subtilis LH expressed in Pichia pastoris GS115 and Escherichia coli Rosetta (DE3). Bioresour Technol 110:539–545
Zhao X, Wang W, Tong B et al (2016) A newly isolated Penicillium oxalicum 16 cellulase with high efficient synergism and high tolerance of monosaccharide. Appl Biochem Biotechnol 178:173–183
Zhao X, Yi S, Li H (2019) The optimized co-cultivation system of Penicillium oxalicum 16 and Trichoderma reesei RUT-C30 achieved a high yield of hydrolase applied in second-generation bioethanol production. Renew Energy 136:1028–1035
Acknowledgements
We thank Dr. Xiaoqiang Huang from Department of Computational Medicine and Bioinformatics, University of Michigan, for the guidance on molecular docking, MD simulation, and binding free energy calculation. This work was supported by the National Natural Science Foundation of China (21666010, 31360217), and the Doctoral Starting up Foundation of Jiangxi Normal University (5451).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
None declared.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Wang, K., Huang, Q., Li, H. et al. Co-evolution of β-glucosidase activity and product tolerance for increasing cellulosic ethanol yield. Biotechnol Lett 42, 2239–2250 (2020). https://doi.org/10.1007/s10529-020-02935-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10529-020-02935-9