Abstract
Histone demethylation is crucial for proper chromatin structure and to ensure normal development, and requires the large family of Jumonji C (JmjC)-containing demethylases; however, the molecular mechanisms that regulate the substrate specificity of these JmjC-containing demethylases remain largely unknown. Here, we show that the substrate specificity of the Arabidopsis histone demethylase JMJ16 is broadened from Lys 4 of histone H3 (H3K4) alone in somatic cells to both H3K4 and H3K9 when it binds to the meiocyte-specific histone reader MMD1. Consistent with this, the JMJ16 catalytic domain exhibits both H3K4 and H3K9 demethylation activities. Moreover, the JMJ16 C-terminal FYR domain interacts with the JMJ16 catalytic domain and probably restricts its substrate specificity. By contrast, MMD1 can compete with the N-terminal catalytic domain of JMJ16 for binding to the FYR-C domain, thereby expanding the substrate specificity of JMJ16 by preventing the FYR domain from binding to the catalytic domain. We propose that MMD1 and JMJ16 together in male meiocytes promote gene expression in an H3K9me3-dependent manner and thereby contribute to meiotic chromosome condensation.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
All data are available in the paper and its Supplementary Information. RNA-seq data of atjmj16 and atmmd1 were deposited to the SRA under accession numbers SRR3584322 (ref. 36) and PRJNA530269. Source data are provided with this paper.
Change history
28 July 2020
A Correction to this paper has been published: https://doi.org/10.1038/s41477-020-00755-3
References
Klose, R. J. & Zhang, Y. Regulation of histone methylation by demethylimination and demethylation. Nat. Rev. Mol. Cell Biol. 8, 307–318 (2007).
Shi, Y. et al. Histone demethylation mediated by the nuclear amine oxidase homolog LSD1. Cell 119, 941–953 (2004).
Tsukada, Y. et al. Histone demethylation by a family of JmjC domain-containing proteins. Nature 439, 811–816 (2006).
Chen, X., Hu, Y. & Zhou, D. X. Epigenetic gene regulation by plant Jumonji group of histone demethylase. Biochim. Biophys. Acta 1809, 421–426 (2011).
Liu, C., Lu, F., Cui, X. & Cao, X. Histone methylation in higher plants. Annu. Rev. Plant Biol. 61, 395–420 (2010).
Kooistra, S. M. & Helin, K. Molecular mechanisms and potential functions of histone demethylases. Nat. Rev. Mol. Cell Biol. 13, 297–311 (2012).
Ko, J. H. et al. Growth habit determination by the balance of histone methylation activities in Arabidopsis. EMBO J. 29, 3208–3215 (2010).
Lu, F., Cui, X., Zhang, S., Jenuwein, T. & Cao, X. Arabidopsis REF6 is a histone H3 lysine 27 demethylase. Nat. Genet. 43, 715–719 (2011).
Yu, X. et al. Modulation of brassinosteroid-regulated gene expression by Jumonji domain-containing proteins ELF6 and REF6 in Arabidopsis. Proc. Natl Acad. Sci. USA 105, 7618–7623 (2008).
Lin, C. H. et al. Heterochromatin protein 1a stimulates histone H3 lysine 36 demethylation by the Drosophila KDM4A demethylase. Mol. Cell 32, 696–706 (2008).
Trojer, P. et al. Dynamic histone H1 isotype 4 methylation and demethylation by histone lysine methyltransferase G9a/KMT1C and the Jumonji domain-containing JMJD2/KDM4 proteins. J. Biol. Chem. 284, 8395–8405 (2009).
Whetstine, J. R. et al. Reversal of histone lysine trimethylation by the JMJD2 family of histone demethylases. Cell 125, 467–481 (2006).
Walport, L. J. et al. Mechanistic and structural studies of KDM-catalysed demethylation of histone 1 isotype 4 at lysine 26. FEBS Lett. 592, 3264–3273 (2018).
Lu, F. et al. Comparative analysis of JmjC domain-containing proteins reveals the potential histone demethylases in Arabidopsis and rice. J. Integr. Plant Biol. 50, 886–896 (2008).
Garcia-Alai, M. M., Allen, M. D., Joerger, A. C. & Bycroft, M. The structure of the FYR domain of transforming growth factor beta regulator 1. Protein Sci. 19, 1432–1438 (2010).
Zhou, X. & Ma, H. Evolutionary history of histone demethylase families: distinct evolutionary patterns suggest functional divergence. BMC Evol. Biol. 8, 294 (2008).
Lu, F., Cui, X., Zhang, S., Liu, C. & Cao, X. JMJ14 is an H3K4 demethylase regulating flowering time in Arabidopsis. Cell Res. 20, 387–390 (2010).
Shen, Y. et al. Over-expression of histone H3K4 demethylase gene JMJ15 enhances salt tolerance in Arabidopsis. Front. Plant Sci. 5, 290 (2014).
Yang, H. et al. Overexpression of a histone H3K4 demethylase, JMJ15, accelerates flowering time in Arabidopsis. Plant Cell Rep. 31, 1297–1308 (2012).
Liu, P. et al. The histone H3K4 demethylase JMJ16 represses leaf senescence in Arabidopsis. Plant Cell 31, 430–443 (2019).
Yang, H. et al. A companion cell-dominant and developmentally regulated H3K4 demethylase controls flowering time in Arabidopsis via the repression of FLC expression. PLoS Genet. 8, e1002664 (2012).
Chen, Q. et al. Structural basis of a histone H3 lysine 4 demethylase required for stem elongation in rice. PLoS Genet. 9, e1003239 (2013).
Cui, X. et al. Control of transposon activity by a histone H3K4 demethylase in rice. Proc. Natl Acad. Sci. USA 110, 1953–1958 (2013).
Yang, Z. et al. Structure of the Arabidopsis JMJ14-H3K4me3 complex provides insight into the substrate specificity of KDM5 subfamily histone demethylases. Plant Cell 30, 167–177 (2018).
Becker, J. S., Nicetto, D. & Zaret, K. S. H3K9me3-dependent heterochromatin: barrier to cell fate changes. Trends Genet. 32, 29–41 (2016).
Johnson, L. et al. Mass spectrometry analysis of Arabidopsis histone H3 reveals distinct combinations of post-translational modifications. Nucleic Acids Res. 32, 6511–6518 (2004).
Zhang, K., Sridhar, V. V., Zhu, J., Kapoor, A. & Zhu, J. K. Distinctive core histone post-translational modification patterns in Arabidopsis thaliana. PLoS ONE 2, e1210 (2007).
Turck, F. et al. Arabidopsis TFL2/LHP1 specifically associates with genes marked by trimethylation of histone H3 lysine 27. PLoS Genet. 3, e86 (2007).
Veiseth, S. V. et al. The SUVR4 histone lysine methyltransferase binds ubiquitin and converts H3K9me1 to H3K9me3 on transposon chromatin in Arabidopsis. PLoS Genet. 7, e1001325 (2011).
Zheng, S. et al. The Arabidopsis H3K27me3 demethylase JUMONJI 13 is a temperature and photoperiod dependent flowering repressor. Nat. Commun. 10, 1303 (2019).
Sun, Q. & Zhou, D. X. Rice jmjC domain-containing gene JMJ706 encodes H3K9 demethylase required for floral organ development. Proc. Natl Acad. Sci. USA 105, 13679–13684 (2008).
Lee, K., Park, O. S. & Seo, P. J. JMJ30-mediated demethylation of H3K9me3 drives tissue identity changes to promote callus formation in Arabidopsis. Plant J. 95, 961–975 (2018).
Reddy, T. V., Kaur, J., Agashe, B., Sundaresan, V. & Siddiqi, I. The DUET gene is necessary for chromosome organization and progression during male meiosis in Arabidopsis and encodes a PHD finger protein. Development 130, 5975–5987 (2003).
Yang, X., Makaroff, C. A. & Ma, H. The Arabidopsis MALE MEIOCYTE DEATH1 gene encodes a PHD-finger protein that is required for male meiosis. Plant Cell 15, 1281–1295 (2003).
Andreuzza, S., Nishal, B., Singh, A. & Siddiqi, I. The chromatin protein DUET/MMD1 controls expression of the meiotic gene TDM1 during male meiosis in Arabidopsis. PLoS Genet. 11, e1005396 (2015).
Wang, J. et al. The PHD finger protein MMD1/DUET ensures the progression of male meiotic chromosome condensation and directly regulates the expression of the condensin gene CAP-D3. Plant Cell 28, 1894–1909 (2016).
Sanchez, R. & Zhou, M. M. The PHD finger: a versatile epigenome reader. Trends Biochem. Sci. 36, 364–372 (2011).
Kelley, L. A. & Sternberg, M. J. Protein structure prediction on the web: a case study using the Phyre server. Nat. Protoc. 4, 363–371 (2009).
Ng, S. S. et al. Crystal structures of histone demethylase JMJD2A reveal basis for substrate specificity. Nature 448, 87–91 (2007).
Kruidenier, L. et al. A selective Jumonji H3K27 demethylase inhibitor modulates the proinflammatory macrophage response. Nature 488, 404–408 (2012).
Zhang, Y. & Reinberg, D. Transcription regulation by histone methylation: interplay between different covalent modifications of the core histone tails. Genes Dev. 15, 2343–2360 (2001).
Lauria, M. & Rossi, V. Epigenetic control of gene regulation in plants. Biochim. Biophys. Acta 1809, 369–378 (2011).
Mohan, M., Herz, H. M. & Shilatifard, A. SnapShot: histone lysine methylase complexes. Cell 149, 498–498 (2012).
Horton, J. R. et al. Enzymatic and structural insights for substrate specificity of a family of Jumonji histone lysine demethylases. Nat. Struct. Mol. Biol. 17, 38–43 (2010).
Yang, Y. et al. Structural insights into a dual-specificity histone demethylase ceKDM7A from Caenorhabditis elegans. Cell Res. 20, 886–898 (2010).
Li, F. et al. Lid2 is required for coordinating H3K4 and H3K9 methylation of heterochromatin and euchromatin. Cell 135, 272–283 (2008).
Okada, Y., Scott, G., Ray, M. K., Mishina, Y. & Zhang, Y. Histone demethylase JHDM2A is critical for Tnp1 and Prm1 transcription and spermatogenesis. Nature 450, 119–123 (2007).
Okada, Y., Tateishi, K. & Zhang, Y. Histone demethylase JHDM2A is involved in male infertility and obesity. J. Androl. 31, 75–78 (2010).
Luo, M., Hung, F. Y., Yang, S. G., Liu, X. C. & Wu, K. Q. Histone lysine demethylases and their functions in plants. Plant Mol. Biol. Rep. 32, 558–565 (2014).
Hojfeldt, J. W., Agger, K. & Helin, K. Histone lysine demethylases as targets for anticancer therapy. Nat. Rev. Drug Discov. 12, 917–930 (2013).
Yan, Y. et al. A MYB-domain protein EFM mediates flowering responses to environmental cues in Arabidopsis. Dev. Cell 30, 437–448 (2014).
Gan, E. S. et al. Jumonji demethylases moderate precocious flowering at elevated temperature via regulation of FLC in Arabidopsis. Nat. Commun. 5, 5098 (2014).
Loenarz, C. et al. PHF8, a gene associated with cleft lip/palate and mental retardation, encodes for an N ε-dimethyl lysine demethylase. Hum. Mol. Genet. 19, 217–222 (2010).
Qi, H. H. et al. Histone H4K20/H3K9 demethylase PHF8 regulates zebrafish brain and craniofacial development. Nature 466, 503–507 (2010).
Klose, R. J. et al. Demethylation of histone H3K36 and H3K9 by Rph1: a vestige of an H3K9 methylation system in Saccharomyces cerevisiae. Mol. Cell. Biol. 27, 3951–3961 (2007).
Crevillen, P. et al. Epigenetic reprogramming that prevents transgenerational inheritance of the vernalized state. Nature 515, 587–590 (2014).
Lalonde, M. E., Cheng, X. & Cote, J. Histone target selection within chromatin: an exemplary case of teamwork. Genes Dev. 28, 1029–1041 (2014).
Qian, S., Wang, Y., Ma, H. & Zhang, L. Expansion and functional divergence of JumonjiC-containing histone demethylases: significance of duplications in ancestral angiosperms and vertebrates. Plant Physiol. 168, 1321–1337 (2015).
Naumann, K. et al. Pivotal role of AtSUVH2 in heterochromatic histone methylation and gene silencing in Arabidopsis. The EMBO J. 24, 1418–1429 (2005).
Wang, Y., Cheng, Z., Lu, P., Timofejeva, L. & Ma, H. Molecular cell biology of male meiotic chromosomes and isolation of male meiocytes in Arabidopsis thaliana. Methods Mol. Biol. 1110, 217–230 (2014).
Hu, C. D., Chinenov, Y. & Kerppola, T. K. Visualization of interactions among bZIP and Rel family proteins in living cells using bimolecular fluorescence complementation. Mol. Cell 9, 789–798 (2002).
Yang, L., Liu, Z., Lu, F., Dong, A. & Huang, H. SERRATE is a novel nuclear regulator in primary microRNA processing in Arabidopsis. The Plant J. 47, 841–850 (2006).
Robert, S., Zouhar, J., Carter, C. & Raikhel, N. Isolation of intact vacuoles from Arabidopsis rosette leaf-derived protoplasts. Nat. Protoc. 2, 259–262 (2007).
Kelley, L. A., Mezulis, S., Yates, C. M., Wass, M. N. & Sternberg, M. J. The Phyre2 web portal for protein modeling, prediction and analysis. Nat. Protoc. 10, 845–858 (2015).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D 66, 486–501 (2010).
Gadella, T. W. Jr., van der Krogt, G. N. & Bisseling, T. GFP-based FRET microscopy in living plant cells. Trends Plant Sci. 4, 287–291 (1999).
Hwang, W. L., Deindl, S., Harada, B. T. & Zhuang, X. Histone H4 tail mediates allosteric regulation of nucleosome remodelling by linker DNA. Nature 512, 213–217 (2014).
Yang, H., Lu, P., Wang, Y. & Ma, H. The transcriptome landscape of Arabidopsis male meiocytes from high-throughput sequencing: the complexity and evolution of the meiotic process. Plant J. 65, 503–516 (2011).
Du, Z., Zhou, X., Ling, Y., Zhang, Z. & Su, Z. AgriGO: a GO analysis toolkit for the agricultural community. Nucleic Acids Res. 38, 64–70 (2010).
Wang, Y. et al. The DNA replication factor RFC1 is required for interference-sensitive meiotic crossovers in Arabidopsis thaliana. PLoS Genet. 8, e1003039 (2012).
Saleh, A., Alvarez-Venegas, R. & Avramova, Z. An efficient chromatin immunoprecipitation (ChIP) protocol for studying histone modifications in Arabidopsis plants. Nat. Protoc. 3, 1018–1025 (2008).
Acknowledgements
We thank staff at the ABRC at Ohio State University for providing the Arabidopsis mutant seeds; J. Gu and Q. Yao at Fudan University for sharing experimental technology; and A. Dong at Fudan University for discussions. This research was supported by grants from the National Natural Science Foundation of China (nos. 31925005 and 31870293) the State Key Laboratory of Genetic Engineering and Fudan University.
Author information
Authors and Affiliations
Contributions
Conceptualization: Y.W., H.M. and J.M. Methodology: J.W., C.Y., S.Z., J.Y., H.D., H.W., J.H. and X.C. Investigation: J.W., C.Y., S.Z., J.Y., H.D., H.W. and J.H. Writing, original draft: Y.W. and J.W. Review and editing, H.M., Y.W. and J.M. Supervision: H.M., Y.W., J.M. and X.C.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information: Nature Plants thanks Jose Gutierrez-Marcos and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Interaction assays of MMD1 and JMJ16.
a, Self-activation detection of full length MMD1, PHD domain (601–680aa) and MMD1ΔPHD domain (1–600aa) with the blank AD. The blue growth yeast stains in QDO/X-gal/ABA (QDO/A/g) indicate self-activation. 3 replicates were done with the similar results. b, The functional enrichment analysis of the isolated proteins by the MMD1ΔPHD as a bait. c, Detection of interaction of MMD1 with JMJ14, JMJ18 and JMJ19 in tobacco leaves using bimolecular fluorescence complementation (BiFC). DAPI staining represents the nucleus. The merged panel (right panel) is the overlay of DAPI (left, blue) and YFP (middle, green). 12 cells were observed in each experiment. d, Showing the divide fragments for JMJ16 and MMD1 for the subsequent examination. e, Y2H assay of MMD1 truncations and FYR-C domain in the QDO/ABA (QDO/A) plates. The growth yeast indicates a positive interaction. The experiment was repeated for 3 times with similar results. f, Pull-down assay shows the interactions between FYR-C domain (978–1209aa) and MMD1-9 truncations staining by Coomassie blue. g, The MMD-9 interacts with the FYR-C domain by GST-pull down staining by Coomassie blue. The interaction between full-length MMD1 and JMJ16 proteins is a positive control, and between FYR-C and GST serves as a negative control. Recombinant His tagged full-length JMJ16 was purified from insect cells. GST tagged truncated MMD-9 and full length MMD1, and His tagged FYR-C were expressed in and purified from E. coli. The experiments were repeated three times. h, The core conserved domain in the MMD1ΔPHD regions and its homologues in plants was named the MMD domain. Alignment of MMD domain in land plants shows the conserved MMD domain highlighted in the red box. i, Comparison of FYR-C domain highlighted in green box in the PKDM7 family shows the similarity and diversity of FYR-C domains.
Extended Data Fig. 2 The co-localization of MMD1 and JMJ16 in male meiocytes, and purification of JMJ16 and MMD1 proteins.
a, MMD1 and JMJ16 partially co-localize in the pachytene chromosome of meiocytes isolated from transgenic plants JMJ16 fused HA driven by a native promoter. DAPI represents chromosome, green signals represent JMJ16 and red signals represent MMD1. Merged panel in the row 4 is the overlay of MMD1 and JMJ16. The co-localization regions of JMJ16 and MMD1 show color in yellow. JMJ16 and MMD1 are marked by HA and MMD1 primary antibodies and Alex Fluor 480 or 555 secondary antibody, respectively. Scale bar = 5 μm. b, The quantification of co-localization ratio of MMD1 and JMJ16 in by Image J in a (7 and 10 biologically independent leptotene and pachytene cells were used, respectively). The data was represented by mean ± SD and P-value (Pachytene/Leptotene) = 6.71 × 10−7. c, The HA-IP-purified JMJ16 protein from the JMJ16-YFP-HA transgenic Arabidopsis leaves was verified by western blot with anti-GFP antibody. d, His-purified full-length JMJ16- from insect cells (SF9) was stained by Coomassie (left) and verified by Western blot with anti-His antibody (right). e, MMD1-GST purified from E. coli (left panel) and after its cleavage (middle panel) was stained by Coomassie blue staining and verified by western blot with anti-MMD1 antibody (right panel). The Western blotting or Coomassie blue staining in each experiment was repeated for 3 times.
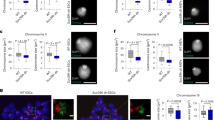
Extended Data Fig. 3 Immunostaining of specific histone modification in WT, mmd1 and jmj16 male meiocytes, and methylation assay of specific histone modification in WT, mmd1 and jmj16 inflorescences by WB.
a–g, Immunostaining of specific histone modification (CENH3, H3K4me1, H3K4me2, H3K9me1, H3K9me2, H3K27me1 and H3K9Ac) on pachytene chromosome in WT mmd1 and jmj16. DAPI was observed in 340 nm and methylated histones were recorded in 555 nm. The merged panel (bottom) is the overlay of DAPI (above, blue) and methylated histones (middle, red). The sample sizes for each experiment were shown in Fig. 3b. h, i, H3 methylation levels in inflorescences by western blot (h) and their quantification in WT, mmd1 and jmj16 (i). Scale bar = 5μm. P-value (jmj16/WT) = 0.0369, * P < 0.05, ** P < 0.01, two-tailed Student’s t test with two-sided. Data are represented as mean ± SEM from three independent experiments.
Extended Data Fig. 4 Interaction examination of JMJ16-N and -C termini, and structural modeling of JMJ16 catalytic domain with zinc-finger and helical domain.
a, Illustration of FRET receptor bleaching in tobacco. The single-direction black arrow represents energy transfer in the top figure. We hypothesize that the closed distance or interaction between the JMJ16 N- and C-termini would result in energy transfer from CFP (donor) to YFP (receptor). Therefore, in the basal state, when YFP is inactivated using photo-bleaching, the energy will remain at CFP. If MMD1 can disrupt the self-interaction of JMJ16 in the bottom part, YFP should be always undetectable if the laser is only given to CFP. CFP will remain unchanged with the photo-bleaching of YFP. b, As the control to Fig. 4i, the laser was given to both CFP and YFP. Then, the YFP was inactivated by photo-bleaching, which demonstrates the normal function of the YFP protein in the presence of MMD1, and the photo-bleaching was also able to reduce the YFP activity. Four biological independent cells were used in the statistics analysis and the data was represented by mean ± SEM. c, The structure model of JMJ16 [catalytic domain Jumonji with C5HC2 zinc-finger and helical domain (HD)]. The Jumonji domain is colored in green, while the HD plus C5HC2-Zinc finger part is in marine. d–f, Structural comparison of JMJ16 model and peptide-bound Jumonji domains. d, JMJ14-H3K4me3 complex (PDB code: 5YKO): The Jumonji domain and HD -C5HC2 domain are colored in violet and yellow, respectively. e, Human JMJD2A-H3K9me3 complex (PDB code: 2OQ6): The Jumonji domain and the extended helixes region (294-355aa) are colored in wheat and gray, respectively. f, Mouse JMJD3-H3K27me3 complex (PDB code: 4EZH): The Jumonji domain and the C-helix and GATAL zinc-finger are colored in light blue and light pink, respectively. JMJ16 is colored in the same scheme as in Supplementary Fig. 4c. The H3 peptides are shown in space-filling representation.
Extended Data Fig. 5 Analyses of the target genes regulated by MMD1 and JMJ16.
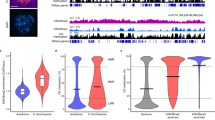
a, The histogram shows up and down-expressed genes regulated by JMJ16 in meiocytes. b, Venn diagram shows genes co-regulated by both MMD1 and JMJ16. c, Validation of JMJ16 expression by anti-HA antibody in JMJ16-HA transgenic plants driven with native promoter. The experiment was repeated for 3 times with similar results. d, Analysis of condensin subunit gene expression in leaves of WT and jmj16. Genes with a reduction in jmj16 meiocytes as shown in Fig. 6b exhibit elevated expression or no significant alternation in leaves. P-values are 0.00686 for CAPE1, 1.91 × 10−4 for CAPE2, 9.36 × 10−4 for CAPG, 1.43 × 10−4 for CAPG2, 0.033 for CAPH and 0.031 for CAPC. e, f, Selected genes such as FLC show significantly increased expression in jmj16 leaves (e), but no significant changes in jmj16 meiocytes (f). The SMC3 gene is a control. P-values are 8.35 × 10−5 for FLC, 0.00144 for COL1, 0.0202 for UNE15, 0.021 for PPME1, 0.011 for PRF5, 0.013 for GRP16), 1.27 × 10−3 for IAA2, 9.37 × 10−4 for IAA7, 2.79 × 10−4 for IAA19, 0.044 for IAA30, 1.45 × 10−3 for AT3G38790, 0.0133 for AT3G28840, 0.0103 for AT3G28980 and 0.016 for AT3G28830 in Extended Data Fig. 5e; P-value (SMC3) = 0.0098 in Extended Data Fig. 5f. g, h, Detection of H3K4me3 (g) and H3K9me3 (h) occupancy in condensin genes in inflorescences of WT and mmd1. P-value (CAPH-P) = 1.94 × 10−4. * P < 0.05, ** P < 0.01, two-tailed Student’s t test with two-sided. Data are represented as mean ± SEM from three independent experiments.
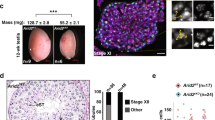
Extended Data Fig. 6 The Arabidopsis jmj16 mutant shows defects in fertility and meiotic chromosome condensation.
a, Illustration of the gene, protein organizations and T-DNA insertional positions of JMJ16. b, The JMJ16 expression level was analyzed by RT-qPCR in WT and jmj16 mutants. P1-4 represents primers’ position on JMJ16 marked in a. Data are represented as mean ± SEM from three independent experiments. c, d, Morphology of fertility was examined and homozygous mutants show partial sterility with abnormally short siliques, scale bar = 5 cm and inviable pollen grains (red pollens are alive and dark/green pollens are dead) by Alexzander staining of anthers, scale bar = 10μm. Independent plants for WT (n = 45), jmj16-1(n = 40), jmj16-2 (n = 25) and JMJ16 complementation transgenic lines (n = 20) were used with similar results. Twenty independent anthers for each genotype were used with similar results. e, Genetic analyses of meiotic chromosome condensation defects in jmj16, mmd1 and mmd1 jmj16 double mutants. Diplotene cells for WT (n = 21), mmd1 (n = 20), jmj16-1 (n = 25) and mmd1 jmj16-1 (n = 30) were used with similar results. Diakinesis cells for WT (n = 25), mmd1 (n = 30), jmj16-1 (n = 31) and mmd1 jmj16-1 (n = 20) were used with similar results; Metaphase I cells for WT (n = 20), mmd1 (n = 24), jmj16-1 (n = 30) and mmd1 jmj16-1 (n = 20) were used with similar results. f, The estimation of metaphase I chromatin areas (circled by yellow lines) in WT (n = 17), mmd1 (n = 11), jmj16-1 (n = 12) and mmd1 jmj16-1 double mutants (n = 16). Data are represented as mean ± SEM. Scale bar = 5μm. P-values are 1.34 × 10−13 (mmd1/ WT), 1.26 × 10−8 (jmj16-1/WT), 5.65 × 10−15 (mmd1 jmj16-1/WT), 1.18 × 10−4 (mmd1/jmj16-1), 1.88 × 10−6 (jmj16-1/mmd1 jmj16-1) and 0.296 (mmd1/mmd1 jmj16-1). * P < 0.05, ** P < 0.01, two-tailed Student’s t test.
Extended Data Fig. 7 The expression of synthetic JMJ16-N with MMD PHD domain partially recues meiotic chromosome condensation defect in mmd1 in vivo.
a, Illustration of synthetic constructs between the PHD domain of MMD1 (PHDMMD1) and the JMJ16-N without (Construct A) or with (Construct B) ZF (zinc finger domain). b, The expression level of JMJ16 in transgenic plant meiocytes was analyzed by RT-qPCR using the primers in the JJMJ16-N. The data are represented by mean ± SEM by three replicates with similar results. P-values are 1.51 × 10−4 (Construct A-1/WT), 1.19 × 10−5 (Construct A-2/WT), 4.54 × 10−4 (Construct B-1/WT) and 4.39 × 10−5 (Construct B-2/WT). c, Meiotic prophase chromosome behaviors of WT, mmd1 and constructs A and B transgenic plants are stained by DAPI in zygotene, pachytene, diakinesis and anaphase II, respectively. Zygotene cells for WT (n = 20), mmd1 (n = 20), Construct A-1 (n = 22), Construct A-2 (n = 21), Construct B-1 (n = 21) and Construct B-2 (n = 20) were used with similar results; Pachytene cells for WT (n = 15), mmd1 (n = 14), Construct A-1 (n = 14), Construct A-2 (n = 10), Construct B-1 (n = 12) and Construct B-2 (n = 20) were used with similar results; Diakinesis cells for WT (n = 20), mmd1 (n = 21), Construct A-1 (n = 20), Construct A-2 (n = 15), Construct B-1 (n = 21) and Construct B-2 (n = 19) were used with similar results; Anaphase II cells for WT (n = 20), mmd1 (n = 24), Construct A-1 (n = 22), Construct A-2 (n = 20), Construct B-1 (n = 12) and Construct B-2 (n = 20) were used with similar results. d, Estimation of chromatin areas of metaphase I chromosome in male meiocytes in Fig. 6i. The areas of chromatins in construct A and construct B were defined by circle drawing tools and area detection by Image J software and the number of cells is labeled in the column. We used cm2 as the area unit that is calculated by the pictures enlarged by the microscope in the centi-meter level, however, the real size of chromatins is micro-meter. P-values are 3.17 ×10−9 (mmd1/WT), 7.7 × 10−11 (Construct B-1/WT) and 5.33 × 10−19 (Construct B-2/WT). The data are represented as mean ± SD. e, RT-qPCR analyses of the CAP-D3 expression in meiocytes of WT, mmd1 and transgenic lines. The data are represented as mean ± SEM with three biological independent replicates. P-values are 3.09 ×10−4 (mmd1/WT), 7.37 × 10−5 (Construct B-1/WT) and 7.74 × 10−7 (Construct B-2/WT). f, The binding affinity of construct A and B. Well 1 and 2 are controls from WT to show H3K4me3 and H3K9me3. FLAG IP was conducted in WT (Well 3), A (Well 4) and B (Well 5) with anti-FLAG. The IP lysates were detected by H3K4me3, H3K9me3 and FLAG antibodies. Construct B only binds H3K4me3 and Construct A binds to both H3K9me3 and H3K4me3. Two replicates for Western blotting were performed. g–i, The complementation of full length MMD1 and JMJ16 in mmd1 mutants. ACTIN::MMD1-FLAG and ACTIN::JMJ16-HA are transformed into mmd1 heterozygotes. g, In the offspring of mmd1 homozygotes, MMD1 and JMJ16 were detected by MMD1 and HA antibodies. h, The siliques used for WT (n = 120), mmd1 (n = 145) and JMJ16 + MMD1(mmd1) (n = 168). i, Meiotic metaphase I chromosome phenotypes used for WT (n = 24), mmd1 (n = 20) and JMJ16 + MMD1(mmd1) (n = 27). Scale bar = 5μm. * P < 0.05, ** P < 0.01, two-tailed Student’s t test.
Supplementary information
Supplementary Information
Supplementary Tables 1–3. Supplementary Table 1: summary of JmjC demethylases towards multiple substrate specificities in plants and animals. JmjC-containing demethylates in plants and animals showed multiple demethylation activities for different histone modifications. Supplementary Table 2: peptides used in this study. Peptides used in the MALDI–TOF-MS and FP are listed in this table with amino-acid sequences, catalogue numbers and providers. Supplementary Table 3: primers used in this study. Primer names, sequences and applications were noted in this table.
Supplementary Data
Genes with downregulated and upregulated expression in jmj16 meiocytes. Sheet 1: 1,633 upregulated genes in jmj16 male meiocytes (log2[fdc] ≥ 1). Sheet 2: 4,626 downregulated genes in jmj16 male meiocytes (log2[fdc] ≤ −1). Sheet 3: 543 coregulated genes with upregulated expression in mmd1 (stage 4–7 anthers) (log2[fdc] ≥ 1) and jmj16 male meiocytes (log2[fdc] ≥ 1). Sheet 4: 802 coregulated genes with downregulated expression in mmd1 (stage 4–7 anthers) (log2[fdc] ≤ −1) and jmj16 male meiocytes (log2[fdc] ≤ −1). Sheet 5: biological processes annotation in upregulated genes or downregulated genes in jmj16 meiocytes. The transcriptome data were obtained from two replicates. For each replicate, about 20,000 male meiocytes were isolated from anthers (see Methods). The statistical significance (P value) of the differential expression of genes was calculated using a two-tailed Student’s t-test; FDR-adjusted P ≤ 0.05 was considered to be significant. The GO enrichment analysis was conducted by AgriGO (http://bioinfo.cau.edu.cn/agriGO/analysis.php), and statistics were compiled by FDR correction; FDR-adjusted P ≤ 0.05 was considered to be significant.
Source data
Source Data Fig. 1
Unprocessed western blots.
Source Data Fig. 2
Unprocessed western blots.
Source Data Fig. 2
Statistical source data.
Source Data Fig. 3
Unprocessed western blots.
Source Data Fig. 3
Statistical source data.
Source Data Fig. 4
Unprocessed western blots.
Source Data Fig. 4
Statistical source data.
Source Data Fig. 5
Statistical source data.
Source Data Fig. 6
Statistical source data.
Source Data Extended Data Fig. 1
Unprocessed gels.
Source Data Extended Data Fig. 2
Unprocessed western blots and gels.
Source Data Extended Data Fig. 2
Statistical source data.
Source Data Extended Data Fig. 3
Unprocessed western blots.
Source Data Extended Data Fig. 3
Statistical source data.
Source Data Extended Data Fig. 5
Statistical source data.
Source Data Extended Data Fig. 6
Statistical source data.
Source Data Extended Data Fig. 7
Unprocessed western blots.
Source Data Extended Data Fig. 7
Statistical source data.
Rights and permissions
About this article
Cite this article
Wang, J., Yu, C., Zhang, S. et al. Cell-type-dependent histone demethylase specificity promotes meiotic chromosome condensation in Arabidopsis. Nat. Plants 6, 823–837 (2020). https://doi.org/10.1038/s41477-020-0697-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41477-020-0697-0
This article is cited by
-
Developmentally dependent reprogramming of the Arabidopsis floral transcriptome under sufficient and limited water availability
BMC Plant Biology (2024)
-
Regulation of micro- and small-exon retention and other splicing processes by GRP20 for flower development
Nature Plants (2024)
-
Resequencing of global Lotus corniculatus accessions reveals population distribution and genetic loci, associated with cyanogenic glycosides accumulation and growth traits
BMC Biology (2023)
-
Plant synthetic epigenomic engineering for crop improvement
Science China Life Sciences (2022)