Abstract
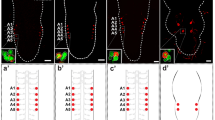
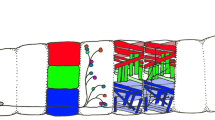
The central nervous system (CNS) of Drosophila is comprised of the brain and the ventral nerve cord (VNC), which are the homologous structures of the vertebrate brain and the spinal cord, respectively. Neurons of the CNS arise from neural stem cells called neuroblasts (NBs). Each neuroblast gives rise to a specific repertory of cell types whose fate is unknown in most lineages. A combination of spatial and temporal genetic cues defines the fate of each neuron. We studied the origin and specification of a group of peptidergic neurons present in several abdominal segments of the larval VNC that are characterized by the expression of the neuropeptide GPB5, the GPB5-expressing neurons (GPB5-ENs). Our data reveal that the progenitor NB that generates the GPB5-ENs also generates the abdominal leucokinergic neurons (ABLKs) in two different temporal windows. We also show that these two set of neurons share the same axonal projections in larvae and in adults and, as previously suggested, may both function in hydrosaline regulation. Our genetic analysis of potential specification determinants reveals that Klumpfuss (klu) and huckebein (hkb) are involved in the specification of the GPB5 cell fate. Additionally, we show that GPB5-ENs have a role in starvation resistance and longevity; however, their role in desiccation and ionic stress resistance is not as clear. We hypothesize that the neurons arising from the same neuroblast lineage are both architecturally similar and functionally related.









Similar content being viewed by others
References
Al-Anzi B, Armand E, Nagamei P, Olszewski M, Sapin V, Waters C, Zinn K, Wyman RJ, Benzer S (2010) The leucokinin pathway and its neurons regulate meal size in Drosophila. Curr Biol 20:969–978
Alvarez-Rivero J, Moris-Sanz M, Estacio-Gomez A, Montoliu-Nerin M, Diaz-Benjumea FJ, Herrero P (2017) Variability in the number of abdominal leucokinergic neurons in adult Drosophila melanogaster. J Comp Neurol 525:639–660
Baonza A, Garcia-Bellido A (2000) Notch signaling directly controls cell proliferation in the Drosophila wing disc. Proc Natl Acad Sci U S A 97:2609–2614
Benito-Sipos J, Estacio-Gomez A, Moris-Sanz M, Baumgardt M, Thor S, Diaz-Benjumea FJ (2010) A genetic cascade involving klumpfuss, nab and castor specifies the abdominal leucokinergic neurons in the Drosophila CNS. Development 137:3327–3336
Birkholz O, Rickert C, Berger C, Urbach R, Technau GM (2013) Neuroblast pattern and identity in the Drosophila tail region and role of doublesex in the survival of sex-specific precursors. Development 140:1830–1842
Birkholz O, Rickert C, Nowak J, Coban IC, Technau GM (2015) Bridging the gap between postembryonic cell lineages and identified embryonic neuroblasts in the ventral nerve cord of Drosophila melanogaster. Biol Open 4:420–434
Bossing T, Udolph G, Doe CQ, Technau GM (1996) The embryonic central nervous system lineages of Drosophila melanogaster. I. Neuroblast lineages derived from the ventral half of the neuroectoderm. Dev Biol 179:41–64
Brody T, Odenwald WF (2000) Programmed transformations in neuroblast gene expression during Drosophila CNS lineage development. Dev Biol 226:34–44
Calleja M, Moreno E, Pelaz S, Morata G (1996) Visualization of gene expression in living adult Drosophila. Science 274:252–255
Cheah PY, Chia W, Yang X (2000) Jumeaux, a novel Drosophila winged-helix family protein, is required for generating asymmetric sibling neuronal cell fates. Development 127:3325–3335
Chu-LaGraff Q, Schmid A, Leidel J, Bronner G, Jackle H, Doe CQ (1995) Huckebein specifies aspects of CNS precursor identity required for motoneuron axon pathfinding. Neuron 15:1041–1051
Doe CQ (1992) Molecular markers for identified neuroblasts and ganglion mother cells in the Drosophila central nervous system. Development 116:855–863
Doe CQ (2017) Temporal patterning in the Drosophila CNS. Annu Rev Cell Dev Biol 33:219–240
Doe CQ, Technau GM (1993) Identification and cell lineage of individual neural precursors in the Drosophila CNS. Trends Neurosci 16:510–514
Estacio-Gomez A, Moris-Sanz M, Schafer AK, Perea D, Herrero P, Diaz-Benjumea FJ (2013) Bithorax-complex genes sculpt the pattern of leucokinergic neurons in the Drosophila central nervous system. Development 140:2139–2148
Froldi F, Cheng LY (2016) Understanding how differentiation is maintained: lessons from the Drosophila brain. Cell Mol Life Sci 73:1641–1644
Grosskortenhaus R, Robinson KJ, Doe CQ (2006) Pdm and Castor specify late-born motor neuron identity in the NB7-1 lineage. Genes Dev 20:2618–2627
Harris, R. M., B. D. Pfeiffer, G. M. Rubin, and J. W. Truman. 2015. Neuron hemilineages provide the functional ground plan for the Drosophila ventral nervous system, Elife, 4
Henrique, D., and F. Schweisguth. 2019. Mechanisms of notch signaling: a simple logic deployed in time and space, Development, 146
Herrera SC, Martin R, Morata G (2013) Tissue homeostasis in the wing disc of Drosophila melanogaster: immediate response to massive damage during development. PLoS Genet 9:e1003446
Herrero P, Estacio-Gomez A, Moris-Sanz M, Alvarez-Rivero J, Diaz-Benjumea FJ (2014) Origin and specification of the brain leucokinergic neurons of Drosophila: similarities to and differences from abdominal leucokinergic neurons. Dev Dyn 243:402–414
Isshiki T, Pearson B, Holbrook S, Doe CQ (2001) Drosophila neuroblasts sequentially express transcription factors which specify the temporal identity of their neuronal progeny. Cell 106:511–521
Jan YN, L. Y. (Jan. 1994) Neuronal cell fate specification in Drosophila. Curr Opin Neurobiol 4:8–13
Kambadur R, Koizumi K, Stivers C, Nagle J, Poole SJ, Odenwald WF (1998) Regulation of POU genes by castor and hunchback establishes layered compartments in the Drosophila CNS. Genes Dev 12:246–260
Kumar A, Bello B, Reichert H (2009) Lineage-specific cell death in postembryonic brain development of Drosophila. Development 136:3433–3442
Lacin H, Truman JW (2016) Lineage mapping identifies molecular and architectural similarities between the larval and adult Drosophila central nervous system. Elife 5:e13399
Lee G, Kim J, Kim Y, Yoo S, Park JH (2018) Identifying and monitoring neurons that undergo metamorphosis-regulated cell death (metamorphoptosis) by a neuron-specific caspase sensor (Casor) in Drosophila melanogaster. Apoptosis 23:41–53
Lopez-Arias B, Dorado B, Herrero P (2011) Blockade of the release of the neuropeptide leucokinin to determine its possible functions in fly behavior: chemoreception assays. Peptides 32:545–552
Mellerick DM, Kassis JA, Zhang SD, Odenwald WF (1992) Castor encodes a novel zinc finger protein required for the development of a subset of CNS neurons in Drosophila. Neuron 9:789–803
Namiki S, Dickinson MH, Wong AM, Korff W, Card GM (2018) The functional organization of descending sensory-motor pathways in Drosophila. Elife 7:e34272
Nassel DR (2002) Neuropeptides in the nervous system of Drosophila and other insects: multiple roles as neuromodulators and neurohormones. Prog Neurobiol 68:1–84
Nassel DR, Cantera R, Karlsson A (1992) Neurons in the cockroach nervous system reacting with antisera to the neuropeptide leucokinin I. J Comp Neurol 322:45–67
Rocco, DA, Paluzzi JV (2016) Functional role of the heterodimeric glycoprotein hormone, GPA2/GPB5, and its receptor, LGR1: an invertebrate perspective. Gen Comp Endocrinol 234:20–27
Rocco DA, Kim DH, Paluzzi JV (2017) Immunohistochemical mapping and transcript expression of the GPA2/GPB5 receptor in tissues of the adult mosquito, Aedes aegypti. Cell Tissue Res 369:313–330
Schmidt H, Rickert C, Bossing T, Vef O, Urban J, Technau GM (1997) The embryonic central nervous system lineages of Drosophila melanogaster. II. Neuroblast lineages derived from the dorsal part of the neuroectoderm. Dev Biol 189:186–204
Sellami A, Agricola HJ, Veenstra JA (2011) Neuroendocrine cells in Drosophila melanogaster producing GPA2/GPB5, a hormone with homology to LH, FSH and TSH. Gen Comp Endocrinol 170:582–588
Spana EP, Doe CQ (1996) Numb antagonizes Notch signaling to specify sibling neuron cell fates. Neuron 17:21–26
Terhzaz S, O'Connell FC, Pollock VP, Kean L, Davies SA, Veenstra JA, Dow JA (1999) Isolation and characterization of a leucokinin-like peptide of drosophila melanogaster. J Exp Biol 202:3667–3676
Truman JW, Bate M (1988) Spatial and temporal patterns of neurogenesis in the central nervous system of Drosophila melanogaster. Dev Biol 125:145–157
Truman JW, Moats W, Altman J, Marin EC, Williams DW (2010) Role of Notch signaling in establishing the hemilineages of secondary neurons in Drosophila melanogaster. Development 137:53–61
Tuthill JC, Wilson RI (2016) Mechanosensation and adaptive motor control in insects. Curr Biol 26:R1022–R1R38
Udolph G, Rath P, Chia W (2001) A requirement for Notch in the genesis of a subset of glial cells in the Drosophila embryonic central nervous system which arise through asymmetric divisions. Development 128:1457–1466
Ulvklo C, MacDonald R, Bivik C, Baumgardt M, Karlsson D, Thor S (2012) Control of neuronal cell fate and number by integration of distinct daughter cell proliferation modes with temporal progression. Development 139:678–689
Urbach R, Technau GM (2003) Molecular markers for identified neuroblasts in the developing brain of Drosophila. Development 130:3621–3637
Veverytsa L, Allan DW (2013) Subtype-specific neuronal remodeling during Drosophila metamorphosis. Fly (Austin) 7:78–86
Vierbuchen T, Ostermeier A, Pang ZP, Kokubu Y, Sudhof TC, Wernig M (2010) Direct conversion of fibroblasts to functional neurons by defined factors. Nature 463:1035–1041
Yang X, Bahri S, Klein T, Chia W (1997) Klumpfuss, a putative Drosophila zinc finger transcription factor, acts to differentiate between the identities of two secondary precursor cells within one neuroblast lineage. Genes Dev 11:1396–1408
Zandawala M, Marley R, Davies SA, Nassel DR (2018) Characterization of a set of abdominal neuroendocrine cells that regulate stress physiology using colocalized diuretic peptides in Drosophila. Cell Mol Life Sci 75:1099–1115
Acknowledgements
We are grateful to JA Veenstra for providing the GPB5-Gal4 and UAS-GPB5RNAi flies and Beatriz Fraile for her technical help in the laboratory. We thank the Bloomington Stock Center for providing fly stocks and the Confocal Microscopy Service of the CBM-SO for technical imaging assistance. We appreciate Melinda Modrell’s assistance with the English language.
Funding
This work was supported by a Spanish Ministerio de Ciencia e Innovación (grant number BFU2014-53761 (to F.J.D-B.)) and by institutional grants from the Fundación Ramón Areces and Banco de Santander to the CBM-SO.
Author information
Authors and Affiliations
Contributions
All authors had full access to all of the data in the study and take responsibility for the accuracy of the data analysis. Study concept and design: FJDB and PH. Acquisition of data: LDP, LMP and PH. Analysis and interpretation of data: LDP, LMP, FJDB and PH. Drafting of manuscript: PH. Obtained funding: FJDB. Study supervision: FJDB and PH.
Corresponding author
Ethics declarations
Ethical approval
This article does not contain any studies with human participants performed by any of the authors.
All procedures performed in this study involving animals were in accordance with the ethical standards of the institution at which the studies were conducted (Consejo Superior de Investigaciones Cientificas/Spanish National Research Council). The only animal used by the authors is insects (Drosophila melanogaster). The use of insects as an experimental system does not require ethical approval.
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Díaz-de-la-Peña, L., Maestro-Paramio, L., Díaz-Benjumea, F.J. et al. Temporal groups of lineage-related neurons have different neuropeptidergic fates and related functions in the Drosophila melanogaster CNS. Cell Tissue Res 381, 381–396 (2020). https://doi.org/10.1007/s00441-020-03231-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00441-020-03231-8