Abstract
Introduction
The metabolic alterations reflecting the influence of prostate cancer cells can be captured through metabolomic profiling.
Objective
To characterize the plasma metabolomic profile in prostatic intraepithelial neoplasia (PIN) and prostate cancer (PCa).
Methods
Metabolomics analyses were performed in plasma samples from individuals classified as non-cancerous control (n = 36), with PIN (n = 16), or PCa (n = 27). Untargeted [26 moieties identified after pre-processing by gas chromatography/mass spectrometry (GC/MS)] and targeted [46 amino acids, carbohydrates, organic acids and fatty acids by GC/MS, and 16 nucleosides and amino acids by ultra performance liquid chromatography-triple quadrupole/mass spectrometry (UPLC-TQ/MS)] analyses were performed. Prostate specific antigen (PSA) concentrations were measured in all samples. In PCa patients, the Gleason scores were determined.
Results
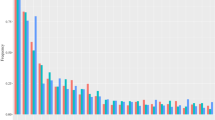
The metabolites that were best discriminated (p < 0.05, FDR < 0.2) for the Kruskal–Wallis test with Dunn’s post-hoc comparing the control versus the PIN and PCa groups included isoleucine, serine, threonine, cysteine, sarcosine, glyceric acid, among several others. PIN was mainly characterized by alterations on steroidogenesis, glycine and serine metabolism, methionine metabolism and arachidonic acid metabolism, among others. In the case of PCa, the most predominant metabolic alterations were ubiquinone biosynthesis, catecholamine biosynthesis, thyroid hormone synthesis, porphyrin and purine metabolism. In addition, we identified metabolites that were correlated to the PSA [i.e. hypoxanthine (r = − 0.60, p < 0.05; r = − 0.54, p < 0.01) and uridine (r = − 0.58, p < 0.05; r = − 0.50, p < 0.01) in PIN and PCa groups, respectively] and metabolites that were significantly different in PCa patients with Gleason score < 7 and ≥ 7 [i.e. arachidonic acid, median (P25–P75) = 883.0 (619.8–956.4) versus 570.8 (505.6–651.8), respectively (p < 0.01)].
Conclusions
This human plasma metabolomic assessment contributes to the understanding of the unique metabolic features exhibited in PIN and PCa and provides a list of metabolites that can have the potential to be used as biomarkers for early detection of disease progression and management.



Similar content being viewed by others
Abbreviations
- PIN:
-
Prostatic intraepithelial neoplasia
- PCa:
-
Prostate cancer
- GC–MS:
-
Gas chromatography/mass spectrometry
- UPLC-TQ/MS:
-
Ultra performance liquid chromatography-triple quadrupole/mass spectrometry
- PSA:
-
Prostate-specific antigen
- MSTFA:
-
N-Methyl-N-trimethylsilyltrifluoroacetamide
- BSA:
-
Bovine serum albumin
- IQR:
-
Interquartile range
- FDR:
-
False discovery rate
References
Baenke, F., Peck, B., Miess, H., & Schulze, A. (2013). Hooked on fat: the role of lipid synthesis in cancer metabolism and tumour development. Disease Models & Mechanisms,6(6), 1353–1363.
Bray, F., Ferlay, J., Soerjomataram, I., Siegel, R. L., Torre, L. A., & Jemal, A. (2018). Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA a Cancer Journal for Clinicians,68(6), 394–424.
Bok, R., Lee, J., Sriram, R., Keshari, K., Sukumar, S., Daneshmandi, S., et al. (2019). The role of lactate metabolism in prostate cancer progression and metastases revealed by dual-agent hyperpolarized 13C MRSI. Cancers,11(2), 257.
Brierley, J. D., Gospodarowicz, M. K., & Wittekind, C. (Eds.). (2016). TNM classification of malignant tumours. Hoboken: Wiley.
Chakroborty, D., Sarkar, C., Basu, B., Dasgupta, P. S., & Basu, S. (2009). Catecholamines regulate tumor angiogenesis. Cancer Research,69(9), 3727–3730.
Choi, W. I., Jeon, B. N., Park, H., Yoo, J. Y., Kim, Y. S., Koh, D. I., et al. (2008). Proto-oncogene FBI-1 (Pokemon) and SREBP-1 synergistically activate transcription of fatty-acid synthase gene (FASN). Journal of Biological Chemistry,283(43), 29341–29354.
De Vogel, S., Ulvik, A., Meyer, K., Ueland, P. M., Nygård, O., Vollset, S. E., et al. (2014). Sarcosine and other metabolites along the choline oxidation pathway in relation to prostate cancer: A large nested case–control study within the JANUS cohort in Norway. International Journal of Cancer,134(1), 197–206.
Eidelman, E., Twum-Ampofo, J., Ansari, J., & Siddiqui, M. M. (2017). The metabolic phenotype of prostate cancer. Frontiers in Oncology,7, 131.
Epstein, J. I., Allsbrook, W. C., Jr., Amin, M. B., Egevad, L. L., & ISUP Grading Committee. (2005). The 2005 International Society of Urological Pathology (ISUP) consensus conference on Gleason grading of prostatic carcinoma. The American Journal of Surgical Pathology,29(9), 1228–1242.
Freeman, M. R., & Solomon, K. R. (2004). Cholesterol and prostate cancer. Journal of Cellular Biochemistry,91(1), 54–69.
Gómez-Cebrián, N., Rojas-Benedicto, A., Albors-Vaquer, A., López-Guerrero, J. A., Pineda-Lucena, A., & Puchades-Carrasco, L. (2019). Metabolomics contributions to the discovery of prostate cancer biomarkers. Metabolites,9(3), 48.
Kelly, R. S., Sinnott, J. A., Rider, J. R., Ebot, E. M., Gerke, T., Bowden, M., et al. (2016b). The role of tumor metabolism as a driver of prostate cancer progression and lethal disease: Results from a nested case-control study. Cancer & metabolism,4(1), 22.
Kelly, R. S., Vander Heiden, M. G., Giovannucci, E., & Mucci, L. A. (2016a). Metabolomic biomarkers of prostate cancer: Prediction, diagnosis, progression, prognosis, and recurrence. Cancer Epidemiology and Prevention Biomarkers,25(6), 887–906.
Krashin, E., Piekiełko-Witkowska, A., Ellis, M., & Ashur-Fabian, O. (2019). Thyroid hormones and cancer: A comprehensive review of preclinical and clinical studies. Frontiers in Endocrinology,10, 59.
Krupp, D., Doberstein, N., Shi, L., & Remer, T. (2012). Hippuric acid in 24-hour urine collections is a potential biomarker for fruit and vegetable consumption in healthy children and adolescents. The Journal of Nutrition,142(7), 1314–1320.
Kumar, D., Gupta, A., Mandhani, A., & Sankhwar, S. N. (2015). Metabolomics-derived prostate cancer biomarkers: Fact or fiction? Journal of Proteome Research,14(3), 1455–1464.
Kumar, D., Gupta, A., Mandhani, A., & Sankhwar, S. N. (2016). NMR spectroscopy of filtered serum of prostate cancer: A new frontier in metabolomics. The Prostate,76(12), 1106–1119.
Leitzmann, M. F., & Rohrmann, S. (2012). Risk factors for the onset of prostatic cancer: Age, location, and behavioral correlates. Clinical epidemiology,4, 1–11.
Locasale, J. W. (2013). Serine, glycine and one-carbon units: Cancer metabolism in full circle. Nature Reviews Cancer,13(8), 572.
Lokhov, P. G., Dashtiev, M. I., Moshkovskii, S. A., & Archakov, A. I. (2010). Metabolite profiling of blood plasma of patients with prostate cancer. Metabolomics,6(1), 156–163.
Lucarelli, G., Ditonno, P., Bettocchi, C., Spilotros, M., Rutigliano, M., Vavallo, A., et al. (2013b). Serum sarcosine is a risk factor for progression and survival in patients with metastatic castration-resistant prostate cancer. Future Oncology,9(6), 899–907.
Lucarelli, G., Fanelli, M., Larocca, A. M. V., Germinario, C. A., Rutigliano, M., Vavallo, A., et al. (2012). Serum sarcosine increases the accuracy of prostate cancer detection in patients with total serum PSA less than 4.0 ng/ml. The Prostate,72(15), 1611–1621.
Lucarelli, G., Loizzo, D., Ferro, M., Rutigliano, M., Vartolomei, M. D., Cantiello, F., et al. (2019). Metabolomic profiling for the identification of novel diagnostic markers and therapeutic targets in prostate cancer: An update. Expert Review of Molecular Diagnostics,19(5), 377–387.
Lucarelli, G., Rutigliano, M., Bettocchi, C., Palazzo, S., Vavallo, A., Galleggiante, V., et al. (2013a). Spondin-2, a secreted extracellular matrix protein, is a novel diagnostic biomarker for prostate cancer. The Journal of Urology,190(6), 2271–2277.
Ludwig, J. A., & Weinstein, J. N. (2005). Biomarkers in cancer staging, prognosis and treatment selection. Nature Reviews Cancer,5(11), 845–856.
Markin, P. A., Brito, A., Moskaleva, N., et al. (2020). Plasma sarcosine measured by gas chromatography-mass spectrometry distinguishes prostatic intraepithelial neoplasia and prostate cancer from benign prostate hyperplasia. Laboratory Medicine. https://doi.org/10.1093/labmed/lmaa008.
Martínez, A., Knappskog, P. M., & Haavik, J. A. (2001). Structural approach into human tryptophan hydroxylase and its implications for the regulation of serotonin biosynthesis. Current Medicinal Chemistry,8(9), 1077–1091.
Midttun, Ø., McCann, A., Aarseth, O., et al. (2016). Combined measurement of 6 fat-soluble vitamins and 26 water-soluble functional vitamin markers and amino acids in 50 μl of serum or plasma by high-throughput mass spectrometry. Analytical Chemistry,88, 10427–10436.
Mondul, A. M., Moore, S. C., Weinstein, S. J., Karoly, E. D., Sampson, J. N., & Albanes, D. (2015). Metabolomic analysis of prostate cancer risk in a prospective cohort: The alpha-tocopherol, beta-carotene cancer prevention (ATBC) study. International Journal of Cancer,137(9), 2124–2132.
Mostaghel, E. A., Solomon, K. R., Pelton, K., Freeman, M. R., & Montgomery, R. B. (2012). Impact of circulating cholesterol levels on growth and intratumoral androgen concentration of prostate tumors. PLoS ONE,7(1), e30062.
Patel, M. I., Kurek, C., & Dong, Q. (2008). The arachidonic acid pathway and its role in prostate cancer development and progression. The Journal of Urology,179(5), 1668–1675.
Pérez-Rambla, C., Puchades-Carrasco, L., García-Flores, M., Rubio-Briones, J., López-Guerrero, J. A., & Pineda-Lucena, A. (2017). Non-invasive urinary metabolomic profiling discriminates prostate cancer from benign prostatic hyperplasia. Metabolomics,13(5), 52.
Shestakova, K., Brito, A., Mesonzhnik, N. V., Moskaleva, N. E., Kurynina, K. O., Grestskaya, N. M., et al. (2018). Rabbit plasma metabolomic analysis of Nitroproston®: A multi target natural prostaglandin based-drug. Metabolomics,14(9), 112.
Šimundić, A. M. (2009). Measures of diagnostic accuracy: Basic definitions. EJIFCC,19(4), 203–211.
Slatkoff, S., Gamboa, S., Zolotor, A. J., Mounsey, A. L., & Jones, K. (2011). PSA testing: When it’s useful, when it’s not. The Journal of Family Practice,60(6), 357–360.
Soto-Guzman, A., Navarro-Tito, N., Castro-Sanchez, L., Martinez-Orozco, R., & Salazar, E. P. (2010). Oleic acid promotes MMP-9 secretion and invasion in breast cancer cells. Clinical & Experimental Metastasis,27(7), 505–515.
Spratlin, J. L., Serkova, N. J., & Eckhardt, S. G. (2009). Clinical applications of metabolomics in oncology: A review. Clinical Cancer Research,15(2), 431–440.
Sreekumar, A., Poisson, L. M., Rajendiran, T. M., Khan, A. P., Cao, Q., Yu, J., et al. (2009). Metabolomic profiles delineate potential role for sarcosine in prostate cancer progression. Nature,457(7231), 910.
Stabler, S., Koyama, T., Zhao, Z., Martinez-Ferrer, M., Allen, R. H., Luka, Z., et al. (2011). Serum methionine metabolites are risk factors for metastatic prostate cancer progression. PLoS ONE,6(8), e22486.
Struck-Lewicka, W., Kaliszan, R., & Markuszewski, M. J. (2014). Analysis of urinary nucleosides as potential cancer markers determined using LC–MS technique. Journal of Pharmaceutical and Biomedical Analysis,101, 50–57.
Thysell, E., Surowiec, I., Hörnberg, E., Crnalic, S., Widmark, A., Johansson, A. I., et al. (2010). Metabolomic characterization of human prostate cancer bone metastases reveals increased levels of cholesterol. PLoS ONE,5, 12.
US Department of Health and Human Services. (2001). Bioanalytical method validation, guidance for industry. https://www.fda.gov/cder/guidance/4252fnl.htm.
van der Mijn, J. C., Kuiper, M. J., Siegert, C. E., Wassenaar, A. E., van Noesel, C. J., & Ogilvie, A. C. (2017). Lactic acidosis in prostate cancer: Consider the Warburg effect. Case Reports in Oncology,10(3), 1085–1091.
Weiner, A. B., Matulewicz, R. S., Eggener, S. E., & Schaeffer, E. M. (2017). Increasing incidence of metastatic prostate cancer in the United States (2004–2013). Prostate Cancer and Prostatic Diseases,20, 283–288.
Windelberg, A., Årseth, O., Kvalheim, G., & Uealand, P. (2005). Automated assay for the determination of methylmalonic acid, total homocysteine, and related amino acids in human serum or plasma by means of methylchloroformate derivatization and gas chromatography-mass spectrometry. Clinical Chemistry,51, 2103–2109.
Wu, H., Liu, T., Ma, C., Xue, R., Deng, C., Zeng, H., et al. (2011). GC/MS-based metabolomic approach to validate the role of urinary sarcosine and target biomarkers for human prostate cancer by microwave-assisted derivatization. Analytical and Bioanalytical Chemistry,401(2), 635–646.
Yang, P., Cartwright, C. A., Li, J. I., Wen, S., Prokhorova, I. N., Shureiqi, I., et al. (2012). Arachidonic acid metabolism in human prostate cancer. International Journal of Oncology,41(4), 1495–1503.
Funding
This study was funded by Project 5-100 Sechenov University Grant.
Author information
Authors and Affiliations
Contributions
The author responsibilities—PAM participated in sample collection, conducted biochemical analyses, performed statistical analyses, interpreted biological and clinical information, and provided input to the manuscript. AB conceptualized the study, performed statistical analyses, interpreted biological and clinical information, and wrote the manuscript. NM conducted biochemical analyses, interpreted biological information and provided input to the manuscript. MRL interpreted metabolic information and provided input to the manuscript. EVL, YVS, YVL, VYM, NVP, DVE and AVL were part of patient recruitment, sample collection, medical history and clinical procedures. SAA conceived the main study and supervised the study. SAA and AB have final responsibility for all parts of this research.
Corresponding authors
Ethics declarations
Conflict of interest
None of the authors have a conflict of interest to declare.
Ethical approval
This research was approved by the Ethics Committee at the I.M Sechenov First Moscow State Medical University, Moscow, Russia. Written signed informed consent was obtained from each volunteer before entry into the study. The study was performed in conformity with the ethical principles for medical research involving humans stated in the Declaration of Helsinki.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
11306_2020_1694_MOESM1_ESM.docx
Supplementary Material: Supplementary table 1: Flow rates for separation of purines and pyrimidines. Supplementary table 2: Non-significant comparison across groups untargeted metabolomics. Supplementary table 3: Non-significant comparison across groups targeted metabolomics. Supplementary table 4: Enrichment analysis control group versus PIN. Supplementary table 5: Enrichment analysis control group versus PCa. Supplementary table 6: Spearman correlations untargeted and targeted metabolomics versus PSA across all groups. Supplementary table 7: Non-significant comparisons untargeted metabolomics according to Gleason score in the PCa group. Supplementary table 8: Non-significant comparisons targeted metabolomics according to Gleason score in the PCa group. Supplementary file1 (DOCX 69 kb)
Rights and permissions
About this article
Cite this article
Markin, P.A., Brito, A., Moskaleva, N. et al. Plasma metabolomic profile in prostatic intraepithelial neoplasia and prostate cancer and associations with the prostate-specific antigen and the Gleason score. Metabolomics 16, 74 (2020). https://doi.org/10.1007/s11306-020-01694-y
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11306-020-01694-y