Abstract
Inflammatory bowel disease (IBD), a chronic relapsing gastrointestinal inflammatory disease, mainly comprises ulcerative colitis (UC) and Crohn’s disease (CD). Although the mechanisms and pathways of IBD have been widely examined in recent decades, its exact pathogenesis remains unclear. Studies have focused on the discovery of new therapeutic targets and application of precision medicine. Recently, a strong connection between IBD and noncoding RNAs (ncRNAs) has been reported. ncRNAs include microRNAs (miRNAs), long noncoding RNAs (lncRNAs), and circular RNAs (circRNAs). The contributions of lncRNAs and circRNAs in IBD are less well-studied compared with those of miRNAs. However, lncRNAs and circRNAs are likely to drive personalized therapy for IBD. They will enable accurate diagnosis, prognosis, and prediction of therapeutic responses and promote IBD therapy. Herein, we briefly describe the molecular functions of lncRNAs and circRNAs and provide an overview of the current knowledge of the altered expression profiles of lncRNAs and circRNAs in patients with IBD. Further, we discuss how these RNAs are involved in the nosogenesis of IBD and are emerging as biomarkers.
Similar content being viewed by others
Facts
-
long noncoding RNAs (lncRNAs) and circular RNAs (circRNAs) are involved in the pathogenesis of inflammatory bowel disease (IBD).
-
Moreover, certain lncRNAs and circRNAs are potential biomarkers of IBD.
-
The contributions of lncRNAs and circRNAs in IBD will become hot spots in future studies.
Open questions
-
lncRNAs and circRNAs show altered expression profile in patients with IBD compared with those in healthy controls.
-
Which and how are lncRNAs and circRNAs involved in the internal mechanism of IBD?
-
Will lncRNAs and circRNAs serve as clinical biomarkers of IBD?
Introduction
Inflammatory bowel disease (IBD) is a chronic inflammatory disease within the gastrointestinal tract. The two major subtypes of IBD are ulcerative colitis (UC) and Crohn’s disease (CD)1. As the occurrence of IBD is continuously increasing, particularly in developing countries, this disease has become a concern worldwide2. There are two main types of treatments for IBD: induction therapy and maintenance therapy. Currently, mucosal healing is regarded as a new therapeutic goal for reducing the rate of re-hospitalizations, operation, and disability3,4. Commonly prescribed drugs for IBD include 5-aminosalicylic acid (5-ASA), corticosteroids, immunosuppressants, thalidomide, and biologicals5,6. However, these therapies cause adverse reactions, lead to a poor quality of life, and cannot achieve the treatment goals for IBD7,8. The etiology of IBD is not completely understood. Understanding the pathogenesis of IBD will help to explore better therapies and reduce the burden on the healthcare system. Increasing evidence has shown that IBD is likely the result of the reciprocal action between genetic factors, environmental factors, and the gut microbiome9.
Recently, scientists have determined the functions of many noncoding RNAs (ncRNAs) in IBD pathogenesis. Approximately 240 risk loci related to IBD have been identified in Genome Wide Association Study10. Most IBD-associated genetic loci are located outside of protein-coding regions and appear to affect ncRNAs11. ncRNAs are indispensable gene regulators at the transcriptional and translational levels and are related to IBD pathobiology12. ncRNAs are chiefly composed of microRNA (miRNA), long noncoding RNA (lncRNA), and circular RNA (circRNA). Among them, miRNAs have been the most thoroughly studied and numerous altered expression profiles of miRNA have been found in IBD13. Moreover, miRNAs are associated with inflammatory pathways in IBD, such as cytokine and chemokine regulation, deregulated autophagy, intestinal epithelial permeability, and necrosis factor-ĸB (NF-ĸB) activation14,15,16,17. We also determined that miRNA can be used in clinical assessment and to regulate intestinal barrier function in patients with IBD18,19,20.
Although lncRNAs and circRNAs have not been thoroughly explored in IBD, they play a crucial role in tumor angiogenesis and carcinogenesis21,22,23,24. Additionally, various lncRNAs and circRNAs have been identified as biomarkers for tumor diagnosis and prognosis25,26,27. Furthermore, some studies suggested that lncRNAs and circRNAs could be promising therapeutic targets of multiple diseases28,29,30,31. Several strategies have been proposed based on the roles of lncRNAs and circRNAs32,33,34,35. Of these methods, antisense oligos36, RNA interference37, small molecules38, and the CRISPR-Cas9 system39 are the leading candidates. ABX464, a small-molecule drug targeting lncRNA 0599–205, has strong anti-inflammatory effects in the dextran sulfate sodium (DSS)-induced colitis model40. lncRNA and circRNA will clearly become the focus of research on IBD.
In this review, we summarize the current knowledge of the molecular functions and roles of lncRNAs and circRNAs in IBD.
Molecular functions of lncRNAs and circRNAs
lncRNAs are long-chain (more than 200 nucleotides) non-coding RNAs that lack open reading frames41. lncRNAs can regulate gene expression transcriptionally, post-transcriptionally, or by guiding chromatin-modifying complexes into specific genomic loci42. There are four major mechanisms of action encompassing most discovered mechanisms. First, lncRNAs can serve as signals to determine the time and location of gene regulation41. They can react with diverse stimuli and reliably reflect the actions of signaling pathways or transcription factors41. Moreover, lncRNAs can serve as decoys, integrating with DNA-binding proteins to avoid combining with DNA recognition elements or interacting with miRNA, thus reducing the degradation of peculiar RNA43. Additionally, lncRNAs can serve as scaffolds, joining several proteins to generate ribonucleoprotein complexes (lncRNA-RNPs)43. In many diverse biological processes, lncRNA acts as a central platform, maintaining precise control over intermolecular interactions and signaling events43. Finally, lncRNAs can serve as guides, directing the specific protein complexes to position at specific targets43. lncRNAs can control target gene expression and generate both neighboring and distant genetic changes43. However, an individual lncRNA may possess several mechanisms of action, and each mode is not necessarily exclusive41.
circRNAs are stable, evolutionarily conserved, and single-stranded RNA molecules. Unlike linear RNAs, circRNAs are closed-loop type RNAs with joined 3′ and 5′ ends44. There are four types of circRNAs, namely exonic circRNAs (ecircRNAs), circular intronic RNAs (ciRNAs), exon–intron circRNAs (EIciRNAs), and intergenic circRNAs45. circRNAs carry out parental gene transcription in addition to a small amount of protein coding46,47. As miRNA sponges, they can regulate RNA expression by adsorbing miRNAs48. Furthermore, circRNAs can interact with RNA-binding proteins to influence certain physiological processes49. circRNAs also act as gene transcription regulators50. ecircRNAs are biosynthesized by back-splicing and affect the transcription of linear RNAs which is carried out by canonical splicing51. Back-splicing competes with canonical splicing when these processes require the same exon and site51. In addition, ecircRNA likely acts as an “mRNA trap” because of isolation of the translation start site and inhibition of RNA translation52. ciRNAs and EIciRNAs interact with polymerase II complex and regulate the expression of parental genes, which is a prerequisite for the other functions of these RNAs50. ecircRNAs are found in the cytoplasm, and some can be loaded into ribosomes, which subsequently translate RNA into peptides or proteins53.
Roles of lncRNAs in IBD
Although numerous studies on IBD have focused on coding-protein genes, lncRNAs are also highly expressed in IBD patients54,55,56 compared with healthy controls. Cumulative data from Genome Wide Association Study revealed a connection between IBD and lncRNA polymorphisms57. Although the functions of lncRNAs in pathological processes and diseases development are less well-studied compared with those in miRNAs, some of lncRNAs are related to IBD pathogenesis (Table 1).
lncRNAs and intestinal epithelial barrier dysregulation
Intestinal epithelial cells (IECs) array and make up intestinal barrier to block a variety of noxious substances such as the microbiota, microbial products, and antigens in the lumen. The specialized structures in the intestinal barrier comprise tight junctions (TJs) and adherent junctions (AJs), ensuring the function of the epithelial barrier58. Studies in patients with IBD showed that intestinal barrier function is disrupted in both active and quiescent disease states59,60. Furthermore, disrupted intestinal barrier, reduction of junctional proteins, and increased intestinal permeability were observed in patients with CD61. Increased epithelial permeability has also been observed in the inactive phase and is strongly predictive of clinical relapse. Destruction of the epithelial barrier is an initial characteristic of disease relapse, suggesting that it plays an initiating role of mucosal inflammation. Many studies have revealed the connection between lncRNAs and the intestinal epithelial barrier.
NEAT1
lncRNA nuclear paraspeckle assembly transcript 1 (NEAT1) is an inflammatory cytokine regulator related to the innate immune response62. NEAT1 is also a key component of the ribonucleoprotein complexes regulating DNA-mediated activation of the innate immune response63. Liu et al.64 reported that compared with control groups, NEAT1 was over-expressed in the intestinal tissues, serum, and exosomes of DSS-induced mice, and in tumor necrosis factor (TNF)-α-induced inflammatory cell models. Similarly, epithelial cell permeability was increased in the above mice and cell models compared with in control groups64. NEAT1 suppression reversed the effects in TNF-α- and DSS-induced IBD models, decreased epithelial cells permeability, and enhanced intestinal epithelial integrity64. However, Birkl et al.65 found that TNF-α may be essential for mucosa repair in the early stage of inflammation. NEAT1 suppression also promoted macrophage polarization towards alternatively activated macrophages (“M2”) rather than classically activated macrophages (“M1”) and inhibited inflammation64. These results revealed that NEAT1 is involved in IBD pathogenesis by regulating intestinal epithelial barrier function and the key mediators involved in disease should be precisely targeted during treatment. Photodynamic therapy (PDT) is a promising therapy for IBD66, particularly low-dose PDT. Farve et al.67 demonstrated that delta-aminolevulinic acid (δ-ALA)-induced low-dose PDT alleviated T-cell-mediated mice colitis and adverse events were negligible. Wang et al.68 verified that 5-ALA-induced PDT relieved DSS-induced colitis in mice through the NEAT1-miRNA204–5p axis.
H19
lncRNA H19 is transcribed from the H19 gene on chromosome 11 (Brannan et al.69). H19 can be found in multiple tissues during the embryonic stage but is silenced after birth70. Under extensive pathological conditions, H19 over-expression is universally detected. Intestinal H19 was dramatically upregulated in mice colitis models, as well as in inflamed colonic tissues from patients with IBD71. Inflammation-induced H19 was observed in IECs. H19 induced by the inflammatory cytokine IL-22 promoted IEC proliferation, epithelial regeneration, and mucosal healing71. Mechanistically, H19 antagonized negative regulators of IECs proliferation, such as p53 protein, miRNA-34a, and let-7, and increased the expression of multiple cell growth-promoting genes in the epithelium71. Other researchers also observed the negative effects of H19. Highly expressed H19 repressed the function of mRNAs encoding TJ protein ZO-1 and AJ protein E-cadherin by releasing miR-675, leading to epithelial barrier damage72. HuR over-expression prevented miR-675 from releasing from H19, promoted ZO-1 and E-cadherin generation, and eliminated H19-induced barrier malfunction72. In contrast, targeted HuR deletion increased the abundance of miR-675 in the intestinal barrier and postponed the recovery of intestinal barrier in mice suffering from pathological stimulus72.
Vitamin D receptor (VDR) is a receptor of 1,25 (OH)2D3 in humans73. 1,25(OH)2D3 is the Vitamin D active form73 and prevents gut damaged by certain destructive reagents74. In multiple tissues, VDR plays an important role in regulating inflammation and carcinogenesis75,76. lncRNA H19 over-expression in UC tissues may decrease VDR and disrupt intestinal epithelial barrier function which is involved in the development of UC77. H19 over-expression significantly decreased ZO-1, occludin, and VDR levels and impaired the function of the Caco-2 monolayer barrier77. The disruptive effect of H19 was partly due to miR-675–5p, which targeted the 3′-untranslated region of VDR mRNA77. miR-675–5p inhibitors can increase ZO-1 and VDR levels77. Therefore, the interaction between lncRNA H19 and VDR signaling may contribute to studies on therapeutic targets for UC.
SPRY4-IT1
Transcribed from the SPRY4 gene, lncRNA SPRY4-IT1 does not code for proteins78. The RNA-binding proteins HuR acts as critical regulator of TJ proteins in the intestine, and HuR dysregulation results in disruption of the epithelial barrier in vitro and in vivo79,80. The 3′-untranslated regions of mRNAs encoding the TJ proteins such as claudin-1, -3, occludin, and JAM-1 include several SPRY4-IT1-binding sites, SPRY4-IT1 silencing caused TJ mRNAs to shift from high-translating sections to low-translating sections in polyribosomes81. SPRY4-IT1 pull-down led to intestinal epithelial barrier dysfunction by reducing the stability of TJ mRNAs81. The process of SPRY4-IT1 regulated TJ mRNAs was enhanced when SPRY4-IT1 associated with HuR81. However, SPRY4-IT1 can interact with TJ mRNAs directly without interacting with HuR81. HuR silencing decreased rather than obstructed the association of SPRY4-IT1 with these TJ mRNAs81. Increasing SPRY4-IT1 levels in the gut showed protective effects by increasing TJ protein expression81. As patients with IBD have increased intestinal permeability, novel molecular therapies aiming to over-express lncRNA SPRY4-IT1 may control gut permeability in specific clinical settings.
CRNDE
The lncRNA colorectal neoplasia differentially expressed (CRNDE) was highly expressed in colorectal adenomas and carcinomas82. CRNDE may be involved in tumorigenesis by regulating miRNAs83. Yang et al.84 suggested that CRNDE is also associated with IBD progression. CRNDE was highly expressed in tissues from DSS-induced mice colitis and human colonic epithelial cells models84. CRNDE inhibition reduced DSS-induced cell apoptosis and cleaved caspase-3, and the apoptotic rate was significantly decreased84. In DSS-induced cell models, CRNDE suppressed miRNA-495 and increased suppressor of cytokine signaling (SOCS1)84. miRNA-495 has been found to be decreased in UC and prevented IEC apoptosis through the JAK signaling pathway85. SOCS1 restricted cytokine receptor signaling86 and promoted IFN-γ-induced IEC apoptosis87. The CRNDE/miR-495/SOCS1 axis was also validated in DSS-induced mice colitis models84. The clinical features of these mice were alleviated after interfering with CRNDE expression, showing improvement in body weight loss and a reduction in bloody stools84. Therefore, lncRNA CRNDE is a potential target for regulating IECs apoptosis through the CRNDE/miR-495/SOCS1 axis.
Other lncRNAs
The lncRNA CDKN2B-AS1 has more than 20 spliced variants containing canonical spliced linear RNA and back-spliced circular RNA molecules88. The longest linear and major circular RNA shape of CDKN2B-AS1 were decreased in UC colon tissues88. Reducing the levels of both linear and circular CDKN2B-AS1 enhanced the barrier formation ability of colonic epithelium by disrupting Claudin-2 expression88. A reduction in CDKN2B-AS1 improved barrier function, showing that the absence of CDKN2B-AS1 in patients may play a protective role after damage88. Xiao et al.89 discovered that elevation of lncRNA uc.173 promoted intestinal epithelium growth. Reduced uc.173 levels slowed IEC renewal by interacting with the pri-miR-195 transcript, resulting in miRNA-195 degradation89. lncRNA BC012900 over-expression resulted in inhibition of IEC proliferation and increased the susceptibility of these cells to apoptosis56. This likely occurred by increasing the abundance of PPM1A (protein phosphatase, Mg2+/Mn2+-dependent, 1A)56. Chen et al.90 reported that elevated PlncRNA1 levels prevent intestinal epithelial barrier injury. Furthermore, PlncRNA1 regulates the level of miRNA-34c90. These two ncRNAs supported the regular effect of the intestinal barrier by mediating the production of TJ proteins ZO-1 and occludin, as well as MAZ90. Another lncRNA, colon cancer–associated transcript–1 (CCAT1), over-expressed in IBD tissues compared with in normal tissues and may be associated with the development of IBD91. CCAT1 can serve as miRNA-185–3p sponge and maintain the stability of myosin light chain kinase (MLCK) mRNA by decreasing miRNA-185–3p binding to MLCK mRNA in Caco-2 cells91. MLCK and its phosphorylation product regulated TJs assembly and increased intestinal permeability91. The positive correlation between CCAT1 and MLCK accelerates IBD development91.
lncRNAs and immune homeostasis dysregulation
IBD is an inflammation disease of the intestinal mucosa, as well as a sustained and aberrant immune disorder, caused by defects in the regulation of intestinal mucosal immunity92. NF-ĸB is a representative immune response factor that can translocate into the nucleus when NF-ĸB inhibitory protein is phosphorylated and then degraded93. This results the transcription of target genes such as interleukin-1β (IL-1β), interleukin-8 (IL-8), and interferon-γ (IFN-γ)94. Some studies reported that excessive inflammatory incidents, such as NF-ĸB activation and high pro-inflammatory cytokines expression, contribute to colitis95,96. Excessive accumulation of immune cells in the gut and induction of complex inflammatory networks make it difficult to explain the roles of individual cytokines and immune pathways as well as the precise etiology and the pathogenesis of IBD97. Interleukin-1b, IL-6, IL-8, and TNF stimulate NF-ĸB, which triggers the transcription of pro-inflammatory cytokines. Regulatory T lymphocytes (Tregs) are an important subset of T lymphocytes. They can limit the functions of immune cells and maintain immunity. Treg dysfunction is attributed to CD and disease severity98.
IFNG-AS1
The lncRNA IFNG-AS1 is located at chromosome 12 in human and is close to IFNG. IFNG-AS1 was increased in patients with active UC compared with in both healthy controls and non-inflamed tissues of patients with UC56. Using human UC samples, mice colitis models, and Jurkat T cell models, Padua et al. found that IFNG-AS1 was related to the IBD single nucleotide polymorphism (SNP) rs7134599 (Padua et al.99). There is a positive link between IFNG-AS1 over-expression and the crucial inflammatory cytokine IFNG expression in immune cells99. Moreover, Rankin et al.100 illustrated that the IFNG-AS1 gene is located beside the inflammatory cytokine IL-22 gene and extensively regulates the pro-inflammatory cascade. IFNG-AS1 may promote the effects of Th1 cytokines (IFNG, IL-2) and reduce the effects of Th2 cytokines (IL-10, IL-13) through an MLL/SET1 mechanism100. Overall, lncRNA IFNG-AS1 is a potential target for treating patients with colitis.
DQ786243
Tregs dysfunction is involved in CD and its severity98. Forkhead box P3 (Foxp3) and cAMP response element-binding protein (CREB) are transcription factors required for the generation, function, and development of Tregs101,102. Zhang et al.103 discovered that the expression of the lncRNA DQ786243 and CREB were increased in the blood of patients with active CD compared with those in the inactive CD and healthy controls. Interestingly, Foxp3 expression was decreased in the blood of patients with inactive CD compared with that in active CD or healthy controls103. DQ786243 may have a significant effect on regulating CREB and Foxp3 genes103. DQ786243 transfection in Jurkat cells promoted CREB and Foxp3 expression as well as CREB phosphorylation in vitro103. As the expression of CREB and Foxp3 in the blood of patients with CD is not significantly correlated, CREB phosphorylation rather than CREB itself may affect Foxp3 expression103. Moreover, the DQ786243, CREB, and Foxp3 mRNAs are related to C-reactive protein (CRP), which is a vital serum biomarker of inflammation103. These finding suggest that lncRNA DQ786243 is involved in CD pathogenesis and may regulate Tregs function by affecting CREB and Foxp3 expression.
LINC01882
There is variation in the genetic locus of protein tyrosine phosphatase 2 (PTPN2) in IBD104. PTPN2 regulates cytokines signaling by acting on multiple phosphorylated proteins105. A study of patients with CD demonstrated a link between the SNP rs2542151 and lower levels of PTPN2 protein in colonic fibroblasts, as well as the formation of aberrant autophagosomes in IECs106. PTPN2 locus SNPs are related to changes in the lncRNA LINC01882, which is primary expressed in T cells and involved in autoimmune diseases, including IBD107. LINC01882 may participate in IL-2 expression, which affects differentiation, immune responses, and homeostasis of various lymphocytes, including Tregs107. Changes in the number of Tregs can contribute to the progress of autoimmune diseases107. However, this study mainly focused on rheumatoid arthritis, and the relationship between LINC01882 and IBD requires further analysis.
Other lncRNAs
The lncRNA ROCKI negatively regulated its cognate encoding gene, myristoylated alanine-rich protein kinase C (MARCKS), by constituting a compound at the MARCKS promoter, which then promoted inflammatory cytokine and chemokine production108. The expression of MARCKS, mediated by ROCKI, may contribute to IBD108. Quan et al.109 examined Roseburia intestinalis flagellin-induced lncRNA expression profiles and found that lncRNA HIF1A-AS2 inactivated the NF-ĸB/JNK pathway and decreased the expression of cytokines IL-1β, IL-6, IL-12, and TNF-α. HIF1A-AS2 was effective for alleviating inflammatory responses in vitro and in vivo; therefore, HIF1A-AS2 may be a negative modulator of intestinal inflammation. The lncRNA ANRIL located at chromosome 9p21 is significantly downregulated in patients with UC110. ANRIL inhibition remarkably reversed the effects of injury by improving cell viability, suppressing cell apoptosis, and reducing inflammatory cytokine production111. The suppressive effects of lncRNA ANRIL were achieved through the TLR4/MyD88/NF-ĸB pathway, which further inhibited UC development111. Li et al.112 identified numerous lncRNAs differentially expressed in the mucosa of CD and predicted a lncRNA-miRNA/TF mRNA network. Most of these lncRNAs are related to cell signaling pathways and immune reactions112. This network is conductive to improve the efficiency of CD gene searches and provides a foundation for follow-up studies112.
lncRNAs as IBD biomarkers
IBD has a large influence on the quality of life and health care system. Clinical manifestations, endoscopic evaluation, imaging methods, and histopathological examinations are commonly used in IBD treatment. However, the clinical features of IBD differ between individuals, and ~25% patients have extraintestinal features before diagnosis113. Endoscopy and histopathological examinations are known as the “gold standard” of IBD diagnosis114,115. But both approaches heavily rely on skilled clinicians, and many adopt alternative methods as a result116. These factors all contribute to the difficulty of diagnosis. Consequently, researchers prefer the use of biomarkers, such as C-reactive protein (CRP), calprotectin, lactoferrin, and others. However, sensitive and specific biomarkers for IBD are lacking. Many lncRNAs have been shown to be involved in IBD. Changes in lncRNA levels can be applied for monitoring of IBD. lncRNAs should be exploited for IBD diagnosis and prognosis, as well as for predicting therapeutic responses. Many lncRNAs can serve as biomarkers for the clinical evaluation of patients with IBD (Table 2).
lncRNA as prognostic and diagnostic biomarkers in IBD
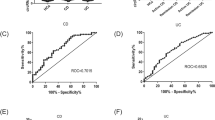
Wang et al.117 demonstrated that in tissues and plasma samples from patients with IBD, lncRNA DIO3OS was significantly downregulated whereas lncRNA KIF9-AS1 and LINC01272 were significantly upregulated compared with in healthy controls117. KIF9-AS1, LINC01272, and DIO3OS have latent diagnostic value for IBD117. The areas under the ROC curve (AUCs) between these three lncRNAs in patients with IBD and healthy controls are mostly higher than 0.76 (Wang et al.117). In summary, the expression of lncRNA KIF9-AS1, LINC01272, and DIO3OS in tissues and plasma samples from IBD patients differed from that in healthy controls and has potential diagnostic value for IBD detection117. Ge et al.118 illustrated that the level of lncRNA ANRIL distinguished patients with CD from healthy controls. The AUC value of ANRIL was 0.803 (Ge et al.118). Interestingly, lncRNA ANRIL can also distinguish the active stage of CD from the remission stage, with an AUC value of 0.839 (Ge et al.118). ANRIL showed negative correlations with disease risk, disease activity, and pro-inflammatory cytokines levels but positive correlations with anti-inflammatory cytokines levels118. lncRNAs acted as biomarkers in both early and late disease stages, even when complications were present. Over half of patients with CD develop complications over time, such as fistulae and stenosis119. The lncRNA RP11–679B19.1 was shown to be associated with recurrent fibrostenotic CD, but its detailed mechanism remains unknown120.
lncRNAs as predictors of therapeutic response in IBD
The lncRNA ANRIL can serve as a biomarker under multiple conditions. Changes in ANRIL expression are associated with the infliximab treatment response in patients with CD. ANRIL from responders of infliximab treatment was increased, whereas that from unresponsive individuals remained stable118. ANRIL upregulation in the intestinal mucosa could act as a marker for assessing the response to infliximab treatment in patients with CD118. Haberman et al.121 reported that based on the intestinal biopsies of the pediatric patients with IBD under treatment, who underwent diagnostic endoscopies, lncRNA HNF4A-AS1 and LINC01272 expression was significantly correlated with severe mucosal ulcers. In addition, LINC01272 showed a significantly positive correlation with calprotectin S100A8, which is currently used as a clinical biomarker of tissue inflammation. However, HNF4A-AS1 was negatively correlated with calprotectin S100A8 (Haberman et al.121). LINC01272 was specifically expressed in myeloid dendritic cells (DC), monocytes, and neutrophils, whereas HNF4A-AS1 was specifically expressed in epithelial cells121. Tissue-specific lncRNA HNF4A-AS1 and LINC01272 expression resulted in the development of a novel lncRNA-directed therapy with fewer off-target effects121. Corticosteroids are commonly prescribed drugs for IBD. Glucocorticoids (GCs) with anti-inflammatory and immunosuppressive effects are used to induce remission in IBD patients122. However, ~20% of patients applying GCs developed resistance to GCs, and 40% of patients maintained clinical remission, relying on GCs. In poor responders to GCs, the levels of lncRNA growth arrest-specific 5 (GAS5) were higher than those in good responders; therefore, GAS5 may be associated with GCs resistance123,124. A later study demonstrated that the expression of GAS5 differed between GCs-sensitive and GCs-resistant cells, and GAS5 is positively correlated with GCs resistance in children with IBD125. Additionally, endogenous GAS5 affects GCs effectiveness, likely because it accumulates in the cytoplasm and plays a role at the post-transcriptional level125. Overall, lncRNA GAS5 can be considered as a novel candidate marker and shows potential for use in the personalization of GCs therapy125.
Roles of circRNAs in IBD
circRNAs are related to a large number of biological processes and diseases. For example, circQTL SNPs are significantly enriched for the Genome Wide Association Study variants associated with various diseases, particularly IBD, schizophrenia, and type II diabetes mellitus126. As a regulator of gene expression, circRNA acts on genetic variation and phenotypic changes. Some circRNAs have been demonstrated to participate in the nosogenesis of IBD and colitis-associated cancer (CAC; Table 3). Although many circRNAs have been identified in humans, functional studies of circRNAs in IBD have not been widely conducted. Thus, analyzing alterations in circRNA profiles and their roles are likely to reveal fundamental molecular mechanisms in IBD.
circRNAs and intestinal epithelial barrier dysregulation
The intestinal epithelium is a cell monolayer constituting an important gut barrier. Intestinal stem cells (ISCs) are rapidly self-renewed and can differentiate into the intestinal epithelium127. Lgr5+ ISCs are an ISC subgroup. circRNA circPan3 (transcribed from Pan3) was over-expressed in human Lgr5+ ISCs and mouse Lgr5-GFP + ISCs128. circPan3 increased the level of IL-13 receptor subunit (IL-13Rα1) by binding to the mRNA of IL-13Rα1 in ISCs and improved the self-renewal capacity of ISCs128. circPan3 deletion in human Lgr5+ ISCs impaired ISC self-renewal and epithelium regeneration capacity128. Similarly, circPan3 bound to IL-13Rα1 mRNA in mice ISCs to preserve its stability and allowed IL-13Rα1 expression in these cells128. In summary, the self-renewal of both human and mice ISCs required the presence of circRNA circPan3 through the IL-13Rα1-mediated signaling pathway.
Although the intestinal epithelial barrier can block most pathogens, several pathogenic bacteria can escape from the barrier and invade IECs129. Autophagy can target and degrade cytoplasmic pathogens in lysosomes130. ATG16L1 is produced from the Atg16l1 gene and has a vital role in autophagy and intestinal epithelium homeostasis. HuR is an RNA-binding protein that is a vital post-transcriptional regulator in the intestinal epithelium131,132. HuR and circRNA circPABPN1 regulated the expression of ATG16L1 in the intestinal epithelium133. Highly expressed circPABPN1 repressed HuR binding to Atg16l1 mRNA in IECs, and then prevented HuR-induced ATG16L1 translation, as well as reduced ATG16L1 production133. The HuR interaction with circPABPN1 partly regulated autophagy by modulating ATG16L1 translation, suggesting that the HuR/circPABPN1/ATG16L1 axis is related to the nosogenesis of IBD and other mucosal disorders133.
circRNAs and immune homeostasis dysregulation
circRNA_103516 was remarkably increased in the active period compared with in emission period of both CD and UC and was positively correlated with disease activity (CD activity index, Mayo, CRP, and erythrocyte sedimentation rate)134. In patients with circRNA_103516 was positively correlated with pro-inflammatory cytokines and negatively correlated with anti-inflammatory cytokine134. This suggests that circRNA_103516 carried out a pro-inflammatory function through inflammation and immune events involved in IBD. Furthermore, the AUC values of circRNA_103516 for UC and CD were 0.687 and 0.790, respectively134. circRNA_103516 presented substantial clinical value for CD and UC, and may therefore also be considered as a novel biomarker for IBD134. In patients with stricture and penetrating CD, the positive prevalence of circRNA_103516 was higher. Thus, circRNA_103516 may contribute to stricture and penetrating behavior of CD134. Moreover, miRNA-19b may inhibit SOCS3 to participate in IEC chemokine production135. circRNA_103516 was negatively correlated with hsa-miRNA-19b-1–5p in patients with CD but not with UC134. circRNA_103516 may be implicated in the molecular mechanisms of CD through hsa-miRNA-19b-1–5p sponging134.
Qiao et al.136 found that circRNA-102685 is highly expressed in the colon tissues of patients with CD and potentially regulates miRNA-146. miRNA-146b relieved gut inflammation by activating NF-ĸB in animal experiments137 and affected the functions of multiple immune cells, such as, Tregs cells and dendritic cells138. In addition, circRNA-102685 was involved in other pathways, such as the chemokine signaling pathway and apoptosis. Those pathways have been identified as being involved in IBD139,140,141. However, the results of Qiao et al.136 are not comprehensive because limited tissue specimens were examined. Generally, circRNA-102685 expression may be related to CD pathogenesis136.
circRNAs as IBD biomarkers
Identifying diagnostic and prognostic biomarkers of IBD will help predict disease behavior and monitor treatment responses. circRNAs with ring structures are more stable than linear RNAs in tissues and body fluids142. This property suggests that circRNAs can act as promising biomarkers of IBD. Some circRNAs were found to be latent biomarkers of IBD (Table 2). Yin et al.143 demonstrated that four circRNAs (004662, 092520, 102610, and 103124) were significantly upregulated in peripheral blood mononuclear cells of patients with CD compared with that those in healthy controls. Furthermore, compared with patients with UC, circRNA_004662 showed higher expression in patients with CD143. The AUC values of these four circRNAs (092520, 102610, 004662, and 103124) were 0.66, 0.78, 0.85, and 0.74, respectively, making these circRNAs potential diagnostic biomarkers of CD143. Considering the observed diagnosis values (P-value, sensitivity, specificity, and AUC), circRNA_004662 may emerge as a promising biomarker to differentiate CD from UC143. circRNA_004662 was an ecircRNA back-spliced from the exon of superoxide dismutase 2 gene and protected cells by mitochondrial reactive oxygen species detoxification144. circRNA_004662 may be related to the mammalian target of rapamycin pathway which limits pro-inflammatory cytokines and enhances anti-inflammatory responses145.
circRNAs in CAC
The CAC risk is increased in patients with IBD, which is in directly linked to the extent and duration of inflammation146. circRNAs with diverse functions are new hotspots for researchers studying the noncoding cancer genome147. The altered expression profiles of circRNAs may be correlated with CAC at the transcription level.
Yuan et al.148 reported that altered expression profiles of circRNAs are involved in CAC development. Compared with in normal colonic tissues, mmu_circRNA_001801, mmu_circRNA_002987, and mmu_circRNA_001155 were the most increased circRNAs, and mmu_circRNA_00287, mmu_circRNA_003037, and mmu_circRNA_001226 were the most decreased circRNAs148. Moreover, this study explored the possible connections between circRNAs and miRNAs and attempted to establish a network. It was found that mmu_ circRNA_001226 and mmu_circRNA_000287 were the first two key points in this network148. The mmu-circ-001226/mmu-circ-000287-miRNA-mRNA network may be the potential mechanism for CAC148.
Conclusion
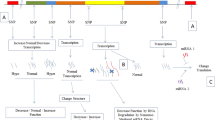
For the past few decades, IBD has become a global health concern149. Its precise pathogenesis remains incompletely understood. Although research on lncRNAs and circRNAs in IBD is still in the early stages, many lncRNAs and circRNAs have been implicated in IBD pathogenesis and have shown promising prospects for clinical applications (Fig. 1). Multiple aspects of lncRNAs, circRNAs, and their functions in IBD require further investigation. Technical tools used to identify the biological mechanism of ncRNAs in human IBD should be improved. Moreover, the complexity of IBD nosogenesis and limited available information indicated that a single lncRNA or circRNA may not entirely explain IBD. Based on the close interactions between lncRNAs, circRNAs, and IBD, it is crucial to further elucidate the molecular mechanisms of these RNAs in IBD, in addition exploring promising therapeutic approaches.
Roles of lncRNA/circRNA in IBD are roughly classified into internal mechanisms and clinical applications. Internal mechanisms mainly focus on lncRNAs/circRNAs regulating immune homeostasis and the intestinal barrier. Clinical applications mainly include biomarkers and therapeutic predictors. lncRNAs/circRNAs can also function as therapeutic targets of IBD with continued research and technique development. lncRNA long noncoding RNA, circRNA circular RNA, IBD inflammatory bowel disease, siRNA small interfering RNA.
Inhibition or enhancement of lncRNAs and circRNAs may be useful for IBD treatment. For efficiency, the development of antagonists or mimics of lncRNAs and circRNAs must be based on tissue-specific or cell type-specific characteristic. Rigorous clinical trials are required to assess the effect and security of these promising treatments. In summary, lncRNAs and circRNAs are promising areas of research for investigating IBD pathogenesis and potential clinical applications.
References
Zhao, M. & Burisch, J. Impact of genes and the environment on the pathogenesis and disease course of inflammatory bowel disease. Dig. Dis. Sci. 64, 1759–1769 (2019).
Ng, S. C. et al. Worldwide incidence and prevalence of inflammatory bowel disease in the 21st century: a systematic review of population-based studies. Lancet 390, 2769–2778 (2017).
Huang, S. et al. Mucosal healing is associated with the reduced disabling disease in Crohn′s disease. Clin. Transl. Gastroenterol. 10, e00015 (2019).
Shah, S. C., Colombel, J. F., Sands, B. E. & Narula, N. Systematic review with meta-analysis: mucosal healing is associated with improved long-term outcomes in Crohn′s disease. Aliment. Pharmacol. Ther. 43, 317–333 (2016).
Harbord, M. et al. Third european evidence-based consensus on diagnosis and management of ulcerative colitis. Part 2: current management. J. Crohns Colitis 11, 769–784 (2017).
Matsuoka, K. et al. Evidence-based clinical practice guidelines for inflammatory bowel disease. J. Gastroenterol. 53, 305–353 (2018).
Hazel, K. & O’Connor, A. Emerging treatments for inflammatory bowel disease. Ther. Adv. Chronic Dis. 11, 2040622319899297 (2020).
Targownik, L. E. & Bernstein, C. N. Infectious and malignant complications of TNF inhibitor therapy in IBD. Am. J. Gastroenterol. 108, 1835–1842 (2013).
Sartor, R. B. Genetics and environmental interactions shape the intestinal microbiome to promote inflammatory bowel disease versus mucosal homeostasis. Gastroenterology 139, 1816–1819 (2010).
Wawrzyniak, M. & Scharl, M. Genetics and epigenetics of inflammatory bowel disease. Swiss Med. Wkly 148, e00015 (2018).
Hrdlickova, B., de Almeida, R. C., Borek, Z. & Withoff, S. Genetic variation in the non-coding genome: involvement of micro-RNAs and long non-coding RNAs in disease. Biochim. Biophys. Acta Mol. Basis Dis. 1842, 1910–1922 (2014).
Yarani, R., Mirza, A. H., Kaur, S. & Pociot, F. The emerging role of lncRNAs in inflammatory bowel disease. Exp. Mol. Med. 50, 1–14 (2018).
Paraskevi, A. et al. Circulating microRNA in inflammatory bowel disease. J. Crohns Colitis 6, 900–904 (2012).
Chen, W.-X., Ren, L.-H. & Shi, R.-H. Implication of miRNAs for inflammatory bowel disease treatment: systematic review. World J. Gastrointest. Pathophysiol. 5, 63–70 (2014).
Feng, X. et al. Up-regulation of microRNA-126 may contribute to athogenesis of ulcerative colitis via regulating NF-kappaB inhibitor I kappa B alpha. PLoS ONE 7, e52782 (2012).
Lu, C. et al. Mir106b and mir93 prevent removal of bacteria from epithelial cells by disrupting ATG16L1-mediated autophagy. Gastroenterology 146, 188–199 (2014).
Ye, D., Guo, S., Al-Sadi, R. & Ma, T. Y. MicroRNA regulation of intestinal epithelial tight junction permeability. Gastroenterology 141, 1323–1333 (2011).
Chen, P. et al. Circulating microRNA146b-5p is superior to C-reactive protein as a novel biomarker for monitoring inflammatory bowel disease. Aliment. Pharmacol. Ther. 49, 733–743 (2019).
Li, L. et al. Cytokine IL9 triggers the pathogenesis of inflammatory bowel disease through the miR21-CLDN8 pathway. Inflamm. Bowel Dis. 24, 2211–2223 (2018).
Wang, H. et al. Pro-inflammatory miR-223 mediates the cross-talk between the IL23 pathway and the intestinal barrier in inflammatory bowel disease. Genome Biol. 17, 58 (2016).
Yan, B., Zhang, W., Mao, X. W. & Jiang, L. Y. Circular RNA ciRS-7 correlates with advance disease and poor prognosis, and its down - regulation inhibits cells proliferation while induces cells apoptosis in non-small cell lung cancer. Eur. Rev. Med. Pharmacol. Sci. 22, 8712–8721 (2018).
Yu, L. et al. The circular RNA Cdr1as act as an oncogene in hepatocellular carcinoma through targeting miR-7 expression. PLoS ONE 11, e0158347 (2016).
Yu, Y. et al. Long non-coding RNA PVT1 promotes cell proliferation and migration by silencing ANGPTL4 expression in cholangiocarcinoma. Mol. Ther. Nucleic Acids 13, 503–513 (2018).
Zhao, J. et al. LncRNA PVT1 promotes angiogenesis via activating the STAT3/VEGFA axis in gastric cancer. Oncogene 37, 4094–4109 (2018).
Chen, H., Wang, K., Pei, D. & Xu, H. Appraising circular RNAs as novel biomarkers for the diagnosis and prognosis of gastric cancer: a pair-wise meta-analysis. J. Clin. Lab. Anal. e23303 (2020).
Dang, H. X. et al. Long non-coding RNA LCAL62 / LINC00261 is associated with lung adenocarcinoma prognosis. Heliyon 6, e03521 (2020).
Chen, Q. et al. Plasma Long non-coding RNA RP11-438N5.3 as a novel biomarker for non-small cell lung cancer. Cancer Manag. Res. 12, 1513–1521 (2020).
Yang, H. et al. CircPTK2 (hsa_circ_0005273) as a novel therapeutic target for metastatic colorectal cancer. Mol. Cancer 19, 13 (2020).
Ni, H. et al. Inhibition of circHIPK3 prevents angiotensin II-induced cardiac fibrosis by sponging miR-29b-3p. Int. J. Cardiol. 292, 188–196 (2019).
Jin, L. et al. LncRNAs as therapeutic targets for autophagy-involved cardiovascular diseases: a review of molecular mechanism and therapy strategy. Curr. Med. Chem. (2020).
Pu, Y., Zhao, H., Wu, X., Mei, M. & Shen, B. The long noncoding RNA Ptprd-IR is a novel molecular target for TGF-beta1-mediated nephritis. Int. J. Biochem. Cell Biol. 122, 105742 (2020).
Liu, T., Ye, P., Ye, Y., Lu, S. & Han, B. Circular RNA hsa_circRNA_002178 silencing retards breast cancer progression via microRNA-328-3p-mediated inhibition of COL1A1. J. Cell. Mol. Med. 24, 2189–2201 (2020).
Xu, L. et al. CRISPR-edited stem cells in a patient with HIV and acute lymphocytic leukemia. N. Engl. J. Med. 381, 1240–1247 (2019).
Zhang, Y.-L., Hu, H.-Y., You, Z.-P., Li, B.-Y. & Shi, K. Targeting long non-coding RNA MALAT1 alleviates retina neurodegeneration in diabetic mice. Int. J. Ophthalmol. 13, 213–219 (2020).
Meng, L. et al. Towards a therapy for Angelman syndrome by targeting a long non-coding RNA. Nature 518, 409–412 (2015).
Bajan, S. & Hutvagner, G. RNA-based therapeutics: from antisense oligonucleotides to miRNAs. Cells 9, 137 (2020).
Wilson, R. C. & Doudna, J. A. Molecular mechanisms of RNA interference. Annu. Rev. Biophys. 42, 217–239 (2013).
Donlic, A. & Hargrove, A. E. Targeting RNA in mammalian systems with small molecules. Wiley Interdiscip. Rev. RNA 9, e1477 (2018).
Wang, H., La Russa, M. & Qi, L. S. CRISPR/Cas9 in genome editing and beyond. Annu. Rev. Biochem. 85, 227–264 (2016).
Vautrin, A. et al. Both anti-inflammatory and antiviral properties of novel drug candidate ABX464 are mediated by modulation of RNA splicing. Sci. Rep. 9, 792 (2019).
Wang, K. C. & Chang, H. Y. Molecular mechanisms of long noncoding RNAs. Mol. Cell 43, 904–914 (2011).
Ghosal, S., Das, S. & Chakrabarti, J. Long noncoding RNAs: new players in the molecular mechanism for maintenance and differentiation of pluripotent stem cells. Stem Cells Dev. 22, 2240–2253 (2013).
Rinn, J. L. & Chang, H. Y. Genome regulation by long noncoding RNAs. Annu. Rev. Biochem. 81, 145–166 (2012).
Hansen, T. B. et al. Natural RNA circles function as efficient microRNA sponges. Nature 495, 384–388 (2013).
Wang, F., Nazarali, A. J. & Ji, S. Circular RNAs as potential biomarkers for cancer diagnosis and therapy. Am. J. Cancer Res. 6, 1167–1176 (2016).
Memczak, S. et al. Circular RNAs are a large class of animal RNAs with regulatory potency. Nature 495, 333–338 (2013).
Jeck, W. R. et al. Circular RNAs are abundant, conserved, and associated with ALU repeats. RNA 19, 141–157 (2013).
Kulcheski, F. R., Christoff, A. P. & Margis, R. Circular RNAs are miRNA sponges and can be used as a new class of biomarker. J. Biotechnol. 238, 42–51 (2016).
Turner, M., Galloway, A. & Vigorito, E. Noncoding RNA and its associated proteins as regulatory elements of the immune system. Nat. Immunol. 15, 484–491 (2014).
Lei, K. et al. The mechanism and function of circular RNAs in human diseases. Exp. Cell Res. 368, 147–158 (2018).
Hou, L.-D. & Zhang, J. Circular RNAs: an emerging type of RNA in cancer. Int. J. Immunopathol. Pharmacol. 30, 1–6 (2017).
Meng, X. et al. Circular RNA: an emerging key player in RNA world. Brief. Bioinform. 18, 547–557 (2017).
Pamudurti, N. R. et al. Translation of CircRNAs. Mol. Cell 66, 9–21 (2017).
Chen, D. et al. Plasma long noncoding RNA expression profile identified by microarray in patients with Crohn′s disease. World J. Gastroenterol. 22, 4716–4731 (2016).
Mirza, A. H. et al. Transcriptomic landscape of lncRNAs in inflammatory bowel disease. Genome Med. 7, 39 (2015).
Wu, F., Huang, Y., Dong, F. & Kwon, J. H. Ulcerative colitis-associated long noncoding RNA, BC012900, regulates intestinal epithelial cell apoptosis. Inflamm. Bowel Dis. 22, 782–795 (2016).
Mirza, A. H., Kaur, S., Brorsson, C. A. & Pociot, F. Effects of GWAS-associated genetic variants on lncRNAs within IBD and T1D candidate loci. PLoS ONE 9, e105723 (2014).
Turner, J. R. Intestinal mucosal barrier function in health and disease. Nat. Rev. Immunol. 9, 799–809 (2009).
Mehta, S., Nijhuis, A., Kumagai, T., Lindsay, J. & Silver, A. Defects in the adherens junction complex (E-cadherin/beta-catenin) in inflammatory bowel disease. Cell Tissue Res. 360, 749–760 (2015).
Foerster, C. Tight junctions and the modulation of barrier function in disease. Histochem. Cell Biol. 130, 55–70 (2008).
Zeissig, S. et al. Changes in expression and distribution of claudin 2, 5 and 8 lead to discontinuous tight junctions and barrier dysfunction in active Crohn′s disease. Gut 56, 61–72 (2007).
Imamura, K. et al. Long noncoding RNA NEAT1-dependent SFPQ relocation from promoter region to paraspeckle mediates IL8 expression upon immune stimuli. Mol. Cell 53, 393–406 (2014).
Morchikh, M. et al. HEXIM1 and NEAT1 long non-coding RNA form a multi-subunit complex that regulates DNA-mediated innate immune response. Mol. Cell 67, 387–399 (2017).
Liu, R. et al. Inhibition of lncRNA NEAT1 suppresses the inflammatory response in IBD by modulating the intestinal epithelial barrier and by exosome-mediated polarization of macrophages. Int. J. Mol. Med. 42, 2903–2913 (2018).
Birkl, D. et al. TNF alpha promotes mucosal wound repair through enhanced platelet activating factor receptor signaling in the epithelium. Mucosal Immunol. 12, 909–918 (2019).
Ortner, M. E. J. et al. Successful photodynamic therapy for nonresectable cholangiocarcinoma: A randomized prospective study. Gastroenterology 125, 1355–1363 (2003).
Favre, L. et al. Low dose endoluminal photodynamic therapy improves murine T cell-mediated colitis. Endoscopy 43, 604–616 (2011).
Wang, K. et al. Neat1-miRNA204-5p-PI3K-AKT axis as a potential mechanism for photodynamic therapy treated colitis in mice. Photodiagnosis Photodyn. Ther. 24, 349–357 (2018).
Brannan, C. I., Dees, E. C., Ingram, R. S. & Tilghman, S. M. The product of the H19 gene may function as an RNA. Mol. Cell. Biol. 10, 28–36 (1990).
Bartolomei, M. S., Zemel, S. & Tilghman, S. M. Parental imprinting of the mouse H19 gene. Nature 351, 153–155 (1991).
Geng, H. et al. In inflamed intestinal tissues and epithelial cells, interleukin 22 signaling increases expression of H19 long noncoding RNA, which promotes mucosal regeneration. Gastroenterology 155, 144–155 (2018).
Zou, T. et al. H19 long noncoding RNA regulates intestinal epithelial barrier function via microRNA 675 by interacting with RNA-binding protein HuR. Mol. Cell. Biol. 36, 1332–1341 (2016).
Haussler, M. R. et al. The nuclear vitamin D receptor: biological and molecular regulatory properties revealed. J. Bone Miner. Res. 13, 325–349 (1998).
Chen, S.-W. et al. Protective effect of 1,25-dihydroxyvitamin D3 on lipopolysaccharide-induced intestinal epithelial tight junction injury in caco-2 cell monolayers. Inflammation 38, 375–383 (2015).
Mousa, A., Misso, M., Teede, H., Scragg, R. & de Courten, B. Effect of vitamin D supplementation on inflammation: protocol for a systematic review. BMJ Open 6, e010804 (2016).
Feldman, D., Krishnan, A. V., Swami, S., Giovannucci, E. & Feldman, B. J. The role of vitamin D in reducing cancer risk and progression. Nat. Rev. Cancer 14, 342–357 (2014).
Chen, S.-W. et al. Effect of long noncoding RNA H19 overexpression on intestinal barrier function and its potential role in the pathogenesis of ulcerative colitis. Inflamm. Bowel Dis. 22, 2582–2592 (2016).
Mazar, J. et al. The functional characterization of long noncoding RNA SPRY4-IT1 in human melanoma cells. Oncotarget 5, 8959–8969 (2014).
Yu, T.-X. et al. Competitive binding of CUGBP1 and HuR to occludin mRNA controls its translation and modulates epithelial barrier function. Mol. Biol. Cell 24, 85–99 (2013).
Yu, T.-X. et al. Chk2-dependent HuR phosphorylation regulates occludin mRNA translation and epithelial barrier function. Nucleic Acids Res. 39, 8472–8487 (2011).
Xiao, L. et al. Long noncoding RNA SPRY4-IT1 regulates intestinal epithelial barrier function by modulating the expression levels of tight junction proteins. Mol. Biol. Cell 27, 617–626 (2016).
Graham, L. D. et al. Colorectal neoplasia differentially expressed (CRNDE), a novel gene with elevated expression in colorectal adenomas and adenocarcinomas. Genes Cancer 2, 829–840 (2011).
Chen, Z. et al. LncRNA CRNDE promotes hepatic carcinoma cell proliferation, migration and invasion by suppressing miR-384. Am. J. Cancer Res. 6, 2299–2309 (2016).
Yang, F., Li, X.-F., Cheng, L.-N. & Li, X.-L. Long non-coding RNA CRNDE promotes cell apoptosis by suppressing miR-495 in inflammatory bowel disease. Exp. Cell Res. 382, 111484 (2019).
Chu, X.-Q. et al. Overexpression of microRNA-495 improves the intestinal mucosal barrier function by targeting STAT3 via inhibition of the JAK/STAT3 signaling pathway in a mouse model of ulcerative colitis. Pathol. Res. Pract. 214, 151–162 (2018).
Calabrese, V. et al. SOCS1 links cytokine signaling to p53 and senescence. Mol. Cell 36, 754–767 (2009).
Cui, X. et al. The suppressor of cytokine signaling SOCS1 promotes apoptosis of intestinal epithelial cells via p53 signaling in Crohn′s disease. Exp. Mol. Pathol. 101, 1–11 (2016).
Rankin, C. R. et al. Linear and circular CDKN2B-AS1 expression is associated with inflammatory bowel disease and participates in intestinal barrier formation. Life Sci. 231, 116571 (2019).
Xiao, L. et al. Long noncoding RNA uc.173 promotes renewal of the intestinal mucosa by inducing degradation of microRNA 195. Gastroenterology 154, 599–611 (2018).
Chen, T., Xue, H., Lin, R. & Huang, Z. MiR-34c and PIncRNA1 mediated the function of intestinal epithelial barrier by regulating tight junction proteins in inflammatory bowel disease. Biochem. Biophys. Res. Commun. 486, 6–13 (2017).
Ma, D. et al. CCAT1 lncRNA promotes inflammatory bowel disease malignancy by destroying intestinal barrier via downregulating miR-185-3p. Inflamm. Bowel Dis. 25, 862–874 (2019).
Geremia, A. & Arancibia-Carcamo, C. V. Innate lymphoid cells in intestinal inflammation. Front. Immunol. 8, 1296 (2017).
Baker, R. G., Hayden, M. S., Ghosh, S. & NF-kappa, B. inflammation, and metabolic disease. Cell Metab. 13, 11–22 (2011).
Shih, V. F.-S., Tsui, R., Caldwell, A. & Hoffmann, A. A single NF kappa B system for both canonical and non-canonical signaling. Cell Res. 21, 86–102 (2011).
Wang, X. et al. Oroxyloside prevents dextran sulfate sodium-induced experimental colitis in mice by inhibiting NF-kappa B pathway through PPAR gamma activation. Biochem. Pharmacol. 106, 70–81 (2016).
Perse, M. & Cerar, A. Dextran sodium sulphate colitis mouse model: traps and tricks. J. Biomed. Biotechnol. 2012, 718617 (2012).
Abraham, C. & Cho, J. H. Inflammatory bowel disease. N. Engl. J. Med. 361, 2066–2078 (2009).
Boden, E. K. & Snapper, S. B. Regulatory T cells in inflammatory bowel disease. Curr. Opin. Gastroenterol. 24, 733–741 (2008).
Padua, D. et al. A long noncoding RNA signature for ulcerative colitis identifies IFNG-AS1 as an enhancer of inflammation. Am. J. Physiol. Gastrointest. Liver Physiol. 311, G446–G457 (2016).
Rankin, C. R. et al. The IBD-associated long noncoding RNA IFNG-AS1 regulates the balance between inflammatory and anti-inflammatory cytokine production after T-cell stimulation. Am. J. Physiol. Gastrointest. Liver Physiol. 318, G34–G40 (2020).
Zhang, L. & Zhao, Y. The regulation of Foxp3 expression in regulatory CD4(+)CD25(+)T cells: multiple pathways on the road. J. Cell. Physiol. 211, 590–597 (2007).
Kim, H.-P. & Leonard, W. J. CREB/ATF-dependent T cell receptor-induced FoxP3 gene expression: a role for DNA methylation. J. Exp. Med. 204, 1543–1551 (2007).
Qiao, Y. Q. et al. LncRNA DQ786243 affects Treg related CREB and Foxp3 expression in Crohn′s disease. J. Biomed. Sci. 20, 87 (2013).
Franke, A. et al. Replication of signals from recent studies of Crohn′s disease identifies previously unknown disease loci for ulcerative colitis. Nat. Genet. 40, 713–715 (2008).
Doody, K. M., Bourdeau, A. & Tremblay, M. L. T-cell protein tyrosine phosphatase is a key regulator in immune cell signaling: lessons from the knockout mouse model and implications in human disease. Immunol. Rev. 228, 325–341 (2009).
Scharl, M. et al. Protein tyrosine phosphatase nonreceptor type 2 regulates autophagosome formation in human intestinal cells. Inflamm. Bowel Dis. 18, 1287–1302 (2012).
Houtman, M. et al. T cells are influenced by a long non-coding RNA in the autoimmune associated PTPN2 locus. J. Autoimmun. 90, 28–38 (2018).
Zhang, Q. et al. The long noncoding RNA ROCKI regulates inflammatory gene expression. EMBO J. 38, e100041 (2019).
Quan, Y. et al. Roseburia intestinalis-derived flagellin is a negative regulator of intestinal inflammation. Biochem. Biophys. Res. Commun. 501, 791–799 (2018).
Liao, K. et al. The research progress of lncRNA involved in the regulation of inflammatory diseases. Mol. Immunol. 101, 182–188 (2018).
Qiao, C. et al. Long noncoding RNA ANRIL contributes to the development of ulcerative colitis by miR-323b-5p/TLR4/MyD88/NF-kappa B pathway. Biochem. Biophys. Res. Commun. 508, 217–224 (2019).
Li, N. & Shi, R. Expression alteration of long non-coding RNAs and their target genes in the intestinal mucosa of patients with Crohn′s disease. Clin. Chim. Acta 494, 14–21 (2019).
Vavricka, S. R. et al. Chronological order of appearance of extraintestinal manifestations relative to the time of IBD diagnosis in the Swiss inflammatory bowel disease cohort. Inflamm. Bowel Dis. 21, 1794–1800 (2015).
Sinh, P. & Shen, B. Endoscopic evaluation of surgically altered bowel in patients with inflammatory bowel diseases. Inflamm. Bowel Dis. 21, 1459–1471 (2015).
Vucelic, B. Inflammatory bowel diseases: controversies in the use of diagnostic procedures. Dig. Dis. 27, 269–277 (2009).
Canavese, G. et al. The diagnosis of inflammatory bowel disease is often unsupported in clinical practice. Dig. Liver Dis. 47, 20–23 (2015).
Wang, S. et al. KIF9-AS1, LINC01272 and DIO3OS lncRNAs as novel biomarkers for inflammatory bowel disease. Mol. Med. Rep. 17, 2195–2202 (2018).
Ge, Q., Dong, Y., Lin, G. & Cao, Y. Long noncoding RNA antisense noncoding RNA in the INK4 locus correlates with risk, severity, inflammation and infliximab efficacy in Crohn′s disease. Am. J. Med. Sci. 357, 134–142 (2019).
Cosnes, J. et al. Long-term evolution of disease behavior of Crohn′s disease. Inflamm. Bowel Dis. 8, 244–250 (2002).
Visschedijk, M. C. et al. Genomic and expression analyses identify a disease-modifying variant for fibrostenotic Crohn′s disease. J. Crohns Colitis 12, 582–588 (2018).
Haberman, Y. et al. Long ncRNA landscape in the ileum of treatment-naive early-onset Crohn disease. Inflamm. Bowel Dis. 24, 346–360 (2018).
Ford, A. C. et al. Glucocorticosteroid therapy in inflammatory bowel disease: systematic review and meta-analysis. Am. J. Gastroenterol. 106, 590–599 (2011).
Lucafo, M. et al. Differential expression of GAS5 in rapamycin-induced reversion of glucocorticoid resistance. Clin. Exp. Pharmacol. Physiol. 43, 602–605 (2016).
Lucafo, M. et al. Long noncoding RNA GAS5: a novel marker involved in glucocorticoid response. Curr. Mol. Med. 15, 94–99 (2015).
Lucafo, M. et al. Role of the long non-coding RNA growth arrest-specific 5 in glucocorticoid response in children with inflammatory bowel disease. Basic Clin. Pharmacol. Toxicol. 122, 87–93 (2018).
Liu, Z. et al. Detection of circular RNA expression and related quantitative trait loci in the human dorsolateral prefrontal cortex. Genome Biol. 20, 99 (2019).
Clevers, H. The intestinal crypt, a prototype stem cell compartment. Cell 154, 274–284 (2013).
Zhu, P. et al. IL-13 secreted by ILC2s promotes the self-renewal of intestinal stem cells through circular RNA circPan3. Nat. Immunol. 20, 183–194 (2019).
Backhed, F., Ley, R. E., Sonnenburg, J. L., Peterson, D. A. & Gordon, J. I. Host-bacterial mutualism in the human intestine. Science 307, 1915–1920 (2005).
Boya, P., Reggiori, F. & Codogno, P. Emerging regulation and functions of autophagy. Nat. Cell Biol. 15, 713–720 (2013).
Xiao, L. & Wang, J.-Y. RNA-binding proteins and microRNAs in gastrointestinal epithelial homeostasis and diseases. Curr. Opin. Pharmacol. 19, 46–53 (2014).
Giammanco, A. et al. Intestinal epithelial HuR modulates distinct pathways of proliferation and apoptosis and attenuates small intestinal and colonic tumor development. Cancer Res. 74, 5322–5335 (2014).
Li, X.-X. et al. Interaction between HuR and circPABPN1 modulates autophagy in the intestinal epithelium by altering ATG16L1 translation. Mol. Cell. Biol. 40, e00492 (2020).
Ye, Y.-L. et al. Increased circulating circular RNA_103516 is a novel biomarker for inflammatory bowel disease in adult patients. World J. Gastroenterol. 25, 6273–6288 (2019).
Cheng, X. et al. MiR-19b downregulates intestinal SOCS3 to reduce intestinal inflammation in Crohn′s disease. Sci. Rep. 5, 10397 (2015).
Qiao, Y. Q., Cai, C. W., Shen, J., Zheng, Q. & Ran, Z. H. Circular RNA expression alterations in colon tissues of Crohn′s disease patients. Mol. Med. Rep. 19, 4500–4506 (2019).
Nata, T. et al. MicroRNA-146b improves intestinal injury in mouse colitis by activating nuclear factor-kappa B and improving epithelial barrier function. J. Gene Med. 15, 249–260 (2013).
Xu, X.-M. & Zhang, H.-J. MiRNAs as new molecular insights into inflammatory bowel disease: crucial regulators in autoimmunity and inflammation. World J. Gastroenterol. 22, 2206–2218 (2016).
Mudter, J. & Neurath, M. F. Apoptosis of T cells and the control of inflammatory bowel disease: therapeutic implications. Gut 56, 293–303 (2007).
Nunes, T., Bernardazzi, C. & de Souza, H. S. Cell death and inflammatory bowel diseases: apoptosis, necrosis, and autophagy in the intestinal epithelium. Biomed. Res. Int. 2014, 218493 (2014).
Singh, U. P. et al. Chemokine and cytokine levels in inflammatory bowel disease patients. Cytokine 77, 44–49 (2016).
Bolha, L., Ravnik-Glavac, M. & Glavac, D. Circular RNAs: biogenesis, function, and a role as possible cancer biomarkers. Int. J. Genomics 2017, 6218353 (2017).
Yin, J. et al. Circular RNA expression profile in peripheral blood mononuclear cells from Crohn disease patients. Medicine 98, e16072 (2019).
Pias, E. K. et al. Differential effects of superoxide dismutase isoform expression on hydroperoxide-induced apoptosis in PC-12 cells. J. Biol. Chem. 278, 13294–13301 (2003).
Weichhart, T. & Saeemann, M. D. The multiple facets of mTOR in immunity. Trends Immunol. 30, 218–226 (2009).
Beaugerie, L. & Itzkowitz, S. H. Cancers complicating inflammatory bowel disease. N. Engl. J. Med. 372, 1441–1452 (2015).
Kristensen, L. S., Hansen, T. B., Veno, M. T. & Kjems, J. Circular RNAs in cancer: opportunities and challenges in the field. Oncogene 37, 555–565 (2018).
Yuan, G. et al. Comprehensive analysis of differential circular RNA expression in a mouse model of colitis-induced colon carcinoma. Mol. Carcinog. 57, 1825–1834 (2018).
Molodecky, N. A. et al. Increasing incidence and prevalence of the inflammatory bowel diseases with time, based on systematic review. Gastroenterology 142, 46–54 (2012).
Acknowledgements
This study was funded by a grant from the National Natural Science Foundation of China (#81870374, #81670498), Guangzhou Science and Technology Department (#202002030041), Guangdong Science and Technology (#2017A030306021), and the Fundamental Research Funds for the Central Universities (#19ykzd11). We would like to thank Editage (editage.cn) for English language editing.
Author information
Authors and Affiliations
Contributions
Guarantor of the article: S.Z., S.Z. and Y.H. designed the study. S.Z., L.L., and G.Z. wrote and revised the manuscript. P.C., Y.W., J.H., and M.C. revised the contents of the manuscript. All authors approved the final version of the manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Edited by H.-U. Simon
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Lin, L., Zhou, G., Chen, P. et al. Which long noncoding RNAs and circular RNAs contribute to inflammatory bowel disease?. Cell Death Dis 11, 456 (2020). https://doi.org/10.1038/s41419-020-2657-z
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41419-020-2657-z
This article is cited by
-
Cross talk between bacterial and human gene networks enriched using ncRNAs in IBD disease
Scientific Reports (2023)
-
lncRNA H-19 and miR-200a implication and frequency of lncRNA H-19 rs2170425 SNP in ulcerative colitis and Crohn’s disease
Comparative Clinical Pathology (2023)
-
Serum long noncoding RNA H19/micro RNA-675-5p axis as a probable diagnostic biomarker in inflammatory bowel disease
Molecular Biology Reports (2023)
-
Role of circular RNAs in retinoblastoma
Functional & Integrative Genomics (2023)
-
Updated immunomodulatory roles of gut flora and microRNAs in inflammatory bowel diseases
Clinical and Experimental Medicine (2022)