Abstract
Tuberculosis is one of the leading causes of death across the world. The treatment regimens for tuberculosis are well established, but still the control of the disease faces many challenges such as lengthy treatment protocols, drug resistance and toxicity. In the present work, mycolic acid methyl transferase (MmaA1), a protein involved in the maturation of mycolic acids in the biochemical pathway of the Mycobacterium, was studied for novel drug discovery. The homology model of the MmaA1 protein was built and validated by using computational techniques. The MmaA1 protein has 286 amino acid residues consisting of 10 α-helices and 7 β-sheets. The active site of the MmaA1 protein was identified using CASTp, SiteMap and PatchDock. Virtual screening studies were performed with two small molecule ligand databases: Asinex synergy and Diverse_Elite_Gold_Platinum databases having a total of 43,446 molecules and generated 1,30,814 conformers against the predicted and validated active site of the MmaA1 protein. Binding analysis showed that the residues ASP 19, PHE 22, TRP 30, TYR 32, TRP 74 and ALA 77 of MmaA1 protein have consistent interactions with the ligands. The hit ligands were further filtered by in silico ADME properties to eliminate potentially toxic molecules. Of the top 10 molecules, 3-(2-morpholinoacetamido)-N-(1,4-dihydro-4-oxoquinazolin-6-yl) benzamide was synthesised and screened for in vitro anti-TB activity against Mtb H37Rv using MABA assay. The compound and its intermediates exhibited good in vitro anti-TB activity which can be taken up for future lead optimisation studies.
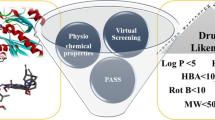
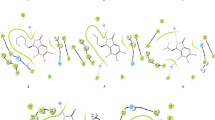
Graphical abstract
Structure based virtual screening study was performed using a validated homology model against small molecules from two virtual compound libraries. Synthesised the lead compound 3-(2-morpholinoacetamido)-N-(1,4-dihydro-4-oxoquinazolin-6-yl)benzamide obtained from virtual screening. In vitro activity against Mtb H37Rv has given a promising result.













Similar content being viewed by others
References
World Health Organisation (2019) Global tuberculosis. https://apps.who.int/iris/handle/10665/274453 Accessed on 10 Feb 2020
WHO Global Tuberculosis Programme (2002) An expanded DOTS framework for effective Tuberculosis control. Stop TB communicable diseases. Geneva: World Health Organisation. WHO/CDS/TB/2002.297
Osborne R (2013) First novel anti-tuberculosis drug in 40 years. Nat Biotech 31(2):89–90. https://doi.org/10.1038/nbt0213-89
Li Y, Sun F, Zhang W (2019) Bedaquiline and delamanid in the treatment of multi drug resistant tuberculosis: promising but challenging. Drug Dev Res 80:98–105. https://doi.org/10.1002/ddr.21498
Villemagne B, Crauste C, Flipo M, Baulard AR, Willand N (2012) Tuberculosis: the drug development pipeline at a glance. Eur J Med Chem 51:1–16. https://doi.org/10.1016/j.ejmech.2012.02.033
Marais BJ (2013) History of tuberculosis and drug resistance. New Engl J Med 386(1):88–90. https://doi.org/10.1056/NEJMc1212308
Nagasree KP, Murali Krishna Kumar M (2016) Chapter 1 In: Antitubercular drug therapy—past, present and future. Science Publishing Group, New York, pp 1–14. ISBN: 978-1-940366-14-2
Companico Andre, Moreira R, Lopes RF (2018) Drug discovery in tuberculosis: new drug targets and antimycobacterial agents. Eur J Med Chem 150:525–545. https://doi.org/10.1016/j.ejmech.2018.03.020
Brennan PJ (2003) Structure, function and biogenesis of the cell wall of mycobacterium tuberculosis. Tuberculosis 83:91–97. https://doi.org/10.1016/S1472-9792(02)00089-6
Marrakchi H, Laneelle MA, Daffe M (2014) Mycolic acids: structures, biosynthesis and beyond. Chem Biol 21(1):67–85. https://doi.org/10.1016/j.chembiol.2013.11.011
Barry CE III, Lee RE, Mdluli K, Sampson AE, Schroeder BG, Slayden RA, Yuan Y (1998) Mycolic acids: structure, biosynthesis and physiological functions. Prog Lipid Res 37:143–179. https://doi.org/10.1016/s0163-7827(98)00008-3
Asselineau C, Asselineau J, Laneellee G, Laneelle MA (2002) The biosynthesis of mycolic acids by Mycobacteria: current and alternative hypotheses. Prog Lipid Res 41(6):501–523. https://doi.org/10.1016/s0163-7827(02)00008-5
Arora P, Goyal A, Natarajan VT, Rajakumara E, Verma P, Gupta R, Yousuf M, Trivedi OA, Mohanty D, Tyagi A, Sankaranarayan R, Gokhale RS (2009) Mechanistic and functional insights into fatty acid activation in mycobacterium tuberculosis. Nat Chem Biol 5:166–173. https://doi.org/10.1038/nchembio.143
Yuan Y, Mead D, Schroeder BG, Zhu Y, Barry CE III (1998) The biosynthesis of mycolic acids in mycobacterium tuberculosis. Enzymatic methy(lene) transfer to acyl carrier protein bound meromycolic acid in vitro. J Biol Chem 273(33):21282–21290. https://doi.org/10.1074/jbc.273.33.21282
Takayama K, Wang C, Besra GS (2005) Pathway to synthesis and processing of mycolic acids in mycobacterium tuberculosis. Clin Microbiol Rev 18(1):81–101. https://doi.org/10.1128/cmr.18.1.81-101.2005
Yuan Y, Crane DC, Musser JM, Sreevatsan S, Barry CE III (1997) MMAS-1, the branch point between cis and trans cyclopropane ring containing oxygenated mycolates in mycobacterium tuberculosis. J Biol Chem 272(15):10041–10049. https://doi.org/10.1074/jbc.272.15.10041
Brossier F, Veziris N, Truffot-Pernot C, Jarlier V, Sougakoff W (2011) Molecular investigation of resistance to the anti tuberculosis drug Ethionamide in multidrug-resistant clinical isolates of mycobacterium tuberculosis. Anti Microb Agent Chemother 55(1):355–360. https://doi.org/10.1128/AAC.01030-10
Cade CE, Dlouhy AC, Medzihradszky KF, Sala-Castillo SP, Ghiladi RA (2010) Isoniazid-resistance conferring mutations in mycobacterium tuberculosis KatG: catalase, peroxidise and INH-NADH adduct formation activities. Prot Sci 9(3):458–474. https://doi.org/10.1002/pro.324
Gasteiger E, Gattiker A, Hoogland C, Ivanyi I, Appel RD, Bairoch A (2003) Expasy: the proteomics server for in-depth protein knowledge and analysis. Nucleic Acids Res 31:3784–3788. https://doi.org/10.1093/nar/gkg563
Berman HM, Westbrook J, Feng Z, Gilliland G, Bhat TN, Weissig H, Shindyalov IN, Bourne PE (2000) The protein data bank. Nucleic Acids Res 28(1):235–242. https://doi.org/10.1093/nar/28.1.235
Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ (1990) Basic local alignment search tool. J Mol Biol 215:403–410. https://doi.org/10.1016/S0022-2836(05)80360-2
Christian C, Jonathan DB, Geoffrey JB (2008) The Jpred3 secondary structure prediction server. Nucleic Acids Res 36:197–201. https://doi.org/10.1093/nar/gkn238
Thompson JD, Higgins DG, Gibson TJ (1994) ClustalW: improving the sensitivity of sequence alignment through sequence weighting, position—specific gap penalties and weight matrix choice. Nucleic Acids Res 22:4673–4680. https://doi.org/10.1093/nar/22.22.4673
Webb B, Sali A (2014) Comparative protein structure modeling using Modeller. Curr Protoc Bioinfo 5(6):1–32. https://doi.org/10.1002/0471250953.bi0506s47
Laskowski RA, MacArthur MW, Moss DS, Thornton JM (1993) PROCHECK: a program to check the stereochemical quality of protein structures. J Appl Crystallogr 26:283–291. https://doi.org/10.1107/S0021889892009944
Kamy Z, Eisenberg D (1994) The three dimensional profile method using residue preference as a continuous function of residue environment. Prot Sci 3:687–695. https://doi.org/10.1002/pro.5560030416
Wiedersten M, Sippl MJ (2007) ProSA-web: interactive web service for the recognition of errors in three-dimensional structures of proteins. Nucleic Acids Res 35:407–441. https://doi.org/10.1093/nar/gkm290
Sheik SS, Sundararajan P, Hussain ASZ, Sekar KK (2002) Ramachandran plot on the web. Bioinformatics 18(11):1548–1549. https://doi.org/10.1093/bioinformatics/18.11.1548
Carrascoza F, Zaric S, Silaghi-Dumitrescu R (2014) Computational study of protein secondary structure elements: Ramachandran plots revisited. J Mol Graph Model 50:125–133. https://doi.org/10.1016/j.jmgm.2014.04.001
Dundas J, Ouyang Z, Tseng J (2006) CASTp: Computed atlas of surface topography of proteins with structural and topographical mapping of functionally annotated residues. Nucleic Acids Res 34:116–118. https://doi.org/10.1093/nar/gkl282
Halgren T (2007) New method for fast and accurate binding site identification and analysis. Chem Biol Drug Des 69:146–148. https://doi.org/10.1111/j.1747-0285.2007.00483.x
Duhovny DS, Inbar Y, Nussinov R, Wolfson HJ (2005) Patchdock and SymmDock: servers for rigid and symmetric Docking. Nucleic Acids Res 33:363–367. https://doi.org/10.1093/nar/gki481
Julien V, Fabienne B, Fanny B (2009) S-adenosyl-N-decyl-aminoethyl, a potent bisubstrate inhibitor of mycobacterium tuberculosis mycolic acid methyl transferases. J Biol Chem 284(29):19321–19330. https://doi.org/10.1074/jbc.M809599200
Shoichet BK (2004) Virtual screening of chemical libraries. Nature 432:862–865. https://doi.org/10.1038/nature03197
Kitchen DB (2004) Docking and scoring in virtual screening for drug discovery: methods and applications. Nat Rev Drug Discov 3:935–949. https://doi.org/10.1038/nrd1549
Friesner RA, Banks JL, Murphy RB, Halgren TA, Klicic JJ, Mainz DT, Repasky MP, Knoll EH, Shelley M, Perry JK, Shaw DE, Francis P, Shenkin PS (2004) Glide: a new approach for rapid, accurate docking and scoring. Method and assessment of docking accuracy. J Med Chem 47(7):1739–1749. https://doi.org/10.1021/jm0306430
Jorgensen WL, Maxwell DS, Tirado-Rives J (1996) Development and testing of the OPLS All-Atom force field on conformational energetics and properties of organic liquids. J Am Chem Soc 118:11225–11236. https://doi.org/10.1021/ja9621760
Kaminski GA, Friesner RA, Tirado-Rives J, Jorgensen WL (2001) Evaluation and reparametrization of the OPLS-AA force field for proteins via comparison with accurate quantum chemical calculations on peptides. J Phys Chem B 105(28):6474–6487. https://doi.org/10.1021/jp003919d
LigPrep, version 2.4 (2010) Schrodinger, LLC, New York, NY
Friesner RA, Murphy RB, Repasky MP, Frye LL, Greenwood JR, Halgren TA, Sanschagrin PC, Mainz DT (2006) Extra precision glide: docking and scoring incorporating a model of hydrophobic enclosure for protein–ligand complexes. J Med Chem 49:6177–6196. https://doi.org/10.1021/jm051256o
Sean E, Waller CL, Swaan PW, Cruciani G, Wrighton SA, Wikel JH (2000) Progress in predicting human ADME parameters in silico. J Pharmacol Tox Methods 44(1):251–272. https://doi.org/10.1016/s1056-8719(00)00109-x
Valerio LG Jr (2009) In silico toxicology for the pharmaceutical sciences. Toxicol Appl Pharmacol 241:356–370. https://doi.org/10.1016/j.taap.2009.08.022
QikProp (2010) Schrodinger, LLC, New York, NY
Tahlan S, Ramasamy K, Lim SM, Shah SAA, Mani V, Narasimhan B (2018) Design, synthesis and therapeutic potential of 3-(2-(1H-benzo[d]imidazol-2-ylthio)acetamido)-N-(substituted phenyl) benzamide analogues. Chem Cent J 12(1):139. https://doi.org/10.1186/s13065-018-0513-3
Wayland EN, Rieke RD (1962) New synthetic route to 6-nitroisatin via nitration of 3-indole aldehyde. J Org Chem 27(6):2250–2252. https://doi.org/10.1021/jo01053a529
Gabriel FR, Silva BV, Martinez ST, Pinto AC (2015) Anthranilic acids from isatin: an efficient, versatile and environmentally friendly method. Ann Acad Braz Sci 87(3):1525–1529. https://doi.org/10.1590/0001-3765201520140289
Sharma GVR, Robert AR (2012) Synthetic strategies to Quinazolinones. Int J Adv Pharm Biol Chem 1(3):337–341 ISSN: 2277-4688
Valeur E, Bradley M (2009) Amide bond formation: beyond the myth of coupling reagents. Chem Soc Rev 38(2):38606–38631. https://doi.org/10.1039/B701677H
Nakayama GR, Caton MC, Nova MP, Parandoosh Z (1997) Assessment of the Alamar Blue assay for cellular growth and viability in vitro. J Immunol Methods 204(2):205–208. https://doi.org/10.1016/s0022-1759(97)00043-4
Daisy Vanitha J, Paramasivan CN (2004) Evaluation of Microplate Alamar Blue assay for drug susceptibility testing of Mycobacterium avium complex isolates. Diagn Microbiol Infect Dis 49:179–182. https://doi.org/10.1016/j.diagmicrobio.2004.04.003
Laskowsky RA, Watson DJ, Thornton MJ (2005) ProFunc: a server for predicting protein function from 3D structure. Nucleic Acids Res 33:89–93. https://doi.org/10.1093/nar/gki414
The PyMOL Molecular Graphics System, Version 2.0, Schrodinger LLC
Ramakrishna D, Ramasree D, Bhargavi K, Vishwanath R, Shantiprada V, Rajender V, Uma V (2016) Suppressor of cytokine signalling-3 as a drug target for type2 diabetes mellitus: a structure guided approach. Chem Select 1:2502–2514. https://doi.org/10.1002/jmr.2706
Ramesh M, Rajender V, Hymavathi V, Kiran Kumar M, Vasavi M, Uma V (2017) Identification of small molecular ligands as potent inhibitors of fatty acid metabolism in Mycobacterium tuberculosis. J Mol Struct 1150:227–241. https://doi.org/10.1016/j.molstruc.2017.08.090
Rajender V, Kiran Kumar M, Vasavi M, Ramasree D, Hymavathi V, Ramesh M, Uma V (2018) Identification of small molecular inhibitors for efflux protein Rv2688c of Mycobacterium tuberculosis. Biotechnol Appl Biochem 65(4):608–621. https://doi.org/10.1002/bab.1647
Accelrys Discovery Studio Visualiser, version 3.5
Acknowledgements
The author HV acknowledges Department of Science and Technology (DST), Government of India, for financial support under the Women Scientist Scheme (WOS-A) vide reference no. SR/WOS-A/CS-1031/2014. The authors HV, VM, KKM, RV, RM and UV acknowledge the Principal and Head, Department of Chemistry, University College of Sciences, Osmania University, Hyderabad, India, for providing the facilities to carry out this work. The authors HV and MMK acknowledge the Principal, AU College of Pharmaceutical Sciences, Andhra University, Visakhapatnam, India, for providing the facilities to carry out this work.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Veeravarapu, H., Malkhed, V., Mustyala, K.K. et al. Structure-based drug design, synthesis and screening of MmaA1 inhibitors as novel anti-TB agents. Mol Divers 25, 351–366 (2021). https://doi.org/10.1007/s11030-020-10107-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11030-020-10107-0