Abstract
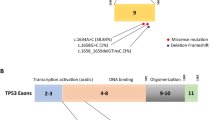
The ERCC4/FANCQ gene is a potential candidate gene for susceptibility to hereditary breast cancer, being a participant of the Fanconi anemia (FA)/BRCA pathway required for DNA repair. ERCC4 encodes XPF endonuclease which mainly participates in nucleotide excision repair (NER) and interstrand crosslink (ICL) repair. Heterozygous mutations in ERCC4 have been identified in various cancers. In this study the heterozygous mutation c.2395C>T (p.Arg799Trp) in ERCC4 was found in a hereditary breast cancer patient using next-generation sequencing. Further screening for the ERCC4*p.Arg799Trp mutation in 966 breast cancer patients and 686 control individuals revealed heterozygous mutation carriers in both groups, but no statistically significant differences in the frequency of the mutant allele between the two samples were found. The results of our study suggest that the ERCC4*p.Arg799Trp mutation is not associated with high risk of breast cancer, although further studies are needed to evaluate the clinical significance of this mutation.




Similar content being viewed by others
REFERENCES
Sijbers, A.M., de Laat, W.L., Ariza, R.R., et al., Xeroderma pigmentosum group F caused by a defect in a structure-specific DNA repair endonuclease, Cell, 1996, vol. 86, pp. 811—822. https://doi.org/10.1016/S0092-8674(00)80155-5
Kashiyama, K., Nakazawa, Y., Pilz, D.T., et al., Malfunction of nuclease ERCC1-XPF results in diverse clinical manifestations and causes Cockayne syndrome, xeroderma pigmentosum, and Fanconi anemia, Am. J. Hum. Genet., 2013, vol. 92, pp. 807—819. https://doi.org/10.1016/j.ajhg.2013.04.007
Bogliolo, M., Schuster, B., Stoepker, C., et al., Mutations in ERCC4, encoding the DNA-repair endonuclease XPF, cause Fanconi anemia, Am. J. Hum. Genet., 2013, vol. 92, pp. 800—806. https://doi.org/10.1016/j.ajhg.2013.04.002
Ferri, D., Orioli, D., and Botta, E., Heterogeneity and overlaps in nucleotide excision repair disorders,Clin. Genet., 2020, vol. 97, pp. 12—24. https://doi.org/10.1111/cge.13545
Marín, M., Ramírez, M.J., Carmona, M.A., et al., Functional comparison of XPF missense mutations associated to multiple DNA repair disorders,Genes, 2019, vol. 10. pii. E60. https://doi.org/10.3390/genes10010060
Wang, Y., Yu, M., Yang, J.-X., et al.,Genomic comparison of endometrioid endometrial carcinoma and its precancerous lesions in Chinese patients by high-depth next generation sequencing, Front. Oncol., 2019, vol. 9, p. 123.https://doi.org/10.3389/fonc.2019.00123
Donner, I., Katainen, R., Sipilä, L.J., et al., Germline mutations in young non-smoking women with lung adenocarcinoma,Lung Cancer, 2018, vol. 122, pp. 76—82. https://doi.org/10.1016/j.lungcan.2018.05.027
Chan, S.H., Lim, W.K., Ishak, N., et al., Germline mutations in cancer predisposition genes are frequent in sporadic sarcomas, Sci. Rep., 2017, vol. 7, p. 10660. https://doi.org/10.1038/s41598-017-10333-x
Berwick, M., Satagopan, J.M., Ben-Porat, L., et al., Genetic heterogeneity among Fanconi anemia heterozygotes and risk of cancer, Cancer Res., 2007, vol. 67, no. 19, pp. 9591—9596. https://doi.org/10.1158/0008-5472.CAN-07-1501
Cantor, S.B., Bell, D.W., Ganesan, S., et al., BACH1, a novel helicase-like protein, interacts directly with BRCA1 and contributes to its DNA repair function, Cell, 2001, vol. 105, no. 1, pp. 149—160. https://doi.org/10.1016/S0092-8674(01)00304-X
Sawyer, S.L., Tian, L., Kähkönen, M., et al., Biallelic mutations in BRCA1 cause a new Fanconi anemia subtype, Cancer Discov., 2015, vol. 5, no. 2, pp. 135—142. https://doi.org/10.1158/2159-8290.CD-14-1156
Domchek, S.M., Tang, J., Stopfer, J., et al., Biallelic deleterious BRCA1 mutations in a woman with early-onset ovarian cancer, Cancer Discov., 2013, vol. 3, pp. 399—405.https://doi.org/10.1158/2159-8290.CD-12-0421
Motycka, T.A., Bessho, T., Post, S.M., et al., Physical and functional interaction between the XPF/ERCC1 endonuclease and hRad52, J. Biol. Chem., 2004, vol. 279, pp. 13634—13639. https://doi.org/10.1074/jbc.M313779200
McDaniel, L.D. and Schultz, R.A., XPF/ERCC4 and ERCC1: their products and biological roles, Adv. Exp. Med. Biol., 2008, vol. 637, pp. 65—82.
Fekairi, S., Scaglione, S., Chahwan, C., et al., Human SLX4 is a Holliday junction resolvase subunit that binds multiple DNA repair/recombination endonucleases, Cell, 2009, vol. 138, pp. 78—89.https://doi.org/10.1016/j.cell.2009.06.029
Fadda, E., Conformational determinants for the recruitment of ERCC1 by XPA in the nucleotide excision repair (NER) pathway: structure and dynamics of the XPA binding motif, Biophys. J., 2013, vol. 104, pp. 2503—2511. https://doi.org/10.1016/j.bpj.2013.04.023
Manandhar, M., Boulware, K.S., and Wood, R.D., The ERCC1 and ERCC4 (XPF) genes and gene products, Gene, 2015, vol. 569, no. 2, pp. 153—161. https://doi.org/10.1016/j.gene.2015.06.026
Bermisheva, M.A., Bogdanova, N.V., Gilyazova, I.R., et al., Ethnic features of genetic susceptibility to breast cancer, Russ. J. Genet., 2018, vol. 45, no. 2, pp. 226—234. https://doi.org/10.1134/S1022795418020047
Doi, H., Koyano, Sh., Miyatake, S., et al., Cerebellar ataxia-dominant phenotype in patients with ERCC4 mutations, J. Hum. Genet., 2018, vol. 63, pp. 417—423. https://doi.org/10.1038/s10038-017-0408-5
Niraj, J., Färkkilä, A., and D’Andrea, A.D., The Fanconi anemia pathway in cancer,Annu. Rev. Cancer Biol., 2019, vol. 3,pp. 457—478. https://doi.org/10.1146/annurev-cancerbio-030617-050422
Wang, L.C. and Gautier, J., The Fanconi anemia pathway and ICL repair: implications for cancer therapy, Crit. Rev. Biochem. Mol. Biol., 2010, vol. 45, no. 5, pp. 424—439.https://doi.org/10.3109/10409238.2010.502166
Dong, H., Nebert, D.W., Bruford, E.A., et al., Update of the human and mouse Fanconi anemia genes, Hum. Genomics, 2015, vol. 9, p. 32. https://doi.org/10.1186/s40246-015-0054-y
Sijbers, A.M., van Voorst Vader, P.C., Snoek, J.W., et al., Homozygous R788W point mutation in the XPF gene of a patient with xerodermapigmentosum and late-onset neurologic disease, J. Invest. Dermatol., 1998, vol. 110, pp. 832—836.https://doi.org/10.1046/j.1523-1747.1998.00171.x
Mori, T., Yousefzadeh, M.J., Faridounnia, M., et al., ERCC4 variants identified in a cohort of patients with segmental progeroid syndromes, Hum. Mut., 2018, vol. 39, pp. 255—265. https://doi.org/10.1002/humu.23367
Osorio, A., Bogliolo, M., Fernandez, V., et al., Evaluation of rare variants in the new Fanconi anemia gene ERCC4 (FANCQ) as familial breast/ovarian cancer susceptibility alleles, Hum. Mut., 2013, vol. 34, no. 12, pp. 1615—1618. https://doi.org/10.1002/humu.22438
Kohlhase, S., Bogdanova, N.V., Schurmann, P., et al., Mutation analysis of the ERCC4/FANCQ gene in hereditary breast cancer, PLoS One, 2014, vol. 9, no. 1. e85334. https://doi.org/10.1371/journal.pone.0085334
Funding
This work was carried out as part of the state task of the Ministry of Education and Science of the Russian Federation (no. AAAA-A16-116020350032-1) with the support of the Russian Foundation of Basic Research no. 17-44-020498 r_а, 17-29-06014 ofi_m, and the Program for the Development of Bioresource Collections no. 007-030164/2, as well as with use of the equipment of the Biomika Shared Access Center and the Unique Scientific Complex KODINK.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest. The authors declare that they have no conflict of interest.
Statement of compliance with standards of research involving humans as subjects. All procedures performed in a study involving people comply with the ethical standards of the institutional and/or national committee for research ethics and the 1964 Helsinki Declaration and its subsequent changes or comparable ethical standards. Informed consent from each of the participants in the studywas obtained.
Additional information
Translated by K. Lazarev
Rights and permissions
About this article
Cite this article
Bermisheva, M.A., Gilyazova, I.R., Zinnatullina, G.F. et al. Analysis of Rare Variant c.2395C>T (p.Arg799Trp) in Gene ERCC4 in Breast Cancer Patients from Bashkortostan. Russ J Genet 56, 627–632 (2020). https://doi.org/10.1134/S1022795420050026
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1134/S1022795420050026