Abstract
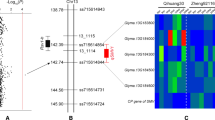
Pythium sylvaticum is one of the most prevalent and aggressive Pythium species causing seedling and root rot of soybean. In this study, two recombinant inbred line populations (POP1 and POP2) and a genome-wide association study (GWAS) panel were used for mapping loci conferring partial resistance to P. sylvaticum in soybean using a greenhouse assay. POP1 (“E09014” × “E05226-T”) and POP2 (“E05226-T” × ‘E09088”) each contains 113 and 79 lines, respectively, and all lines were genotyped using the SoySNP6K BeadChip. QTL mapping using composite interval mapping (CIM) identified 5 QTL on soybean chromosomes of 10 (q10.1 andq10.2), 18 (q18.1 and q18.2), and 20 (q20.1), and each QTL explained 9.7–16.6% of phenotypic variation. The GWAS panel consisted of 214 soybean lines and was genotyped using the SoySNP50K BeadChip. A total of 7 significant SNP markers were identified on chromosomes 10, 18, and 20. Markers Gm10_42965189_G_T, Gm10_42975806_T_C, and Gm10_43004105_A_C were closely linked with q10.1 (<50 kb). Marker Gm18_7898429_A_C was co-localized with q18.2 and was located within the coding region of Glyma.18 g081700. Further investigation revealed that Gm20_2245263_G_A co-localized with qRRW20, a QTL previously identified for partial resistance to P. irregulare. All of the QTL identified in this study can be incorporated into soybean lines to improve enduring partial resistance to Pythium diseases using marker assisted breeding.




Similar content being viewed by others
References
Allen TW, Bradley CA, Sisson AJ, Byamukama E, Chilvers MI, Coker CM, Collins AA, Damicone JP, Dorrance AE, Dufault NS, Esker PD, Faske TR, Giesler LJ, Grybauskas AP, Hershman DE, Hollier CA, Isakeit T, Jardine DJ, Kelly HM, Kemerait RC, Kleczewski NM, Koenning SR, Kurle JE, Malvick DK, Markell SG, Mehl HL, Mueller DS, Mueller JD, Mulrooney RP, Nelson BD, Newman MA, Osborne L, Overstreet C, Padgett GB, Phipps PM, Price PP, Sikora EJ, Smith DL, Spurlock TN, Tande CA, Tenuta AU, Wise KA, Wrather JA (2017) Soybean yield loss estimates due to diseases in the United States and Ontario, Canada, from 2010 to 2014. Plant Health Progress 18:19–27
Arsenault-Labrecque G, Sonah H, Lebreton A, Labbe C, Marchand G, Xue A, Belzile F, Knaus BJ, Grunwald NJ, Belanger RR (2018) Stable predictive markers for Phytophthora sojae avirulence genes that impair infection of soybean uncovered by whole genome sequencing of 31 isolates. BMC Biol 16:80
Bates D, Maechler M, Bolker B, Walker S (2015) lme4: linear mixed-effects models using Eigen and S4. R package version 1.1-7. 2014
Bradbury PJ, Zhang Z, Kroon DE, Casstevens TM, Ramdoss Y, Buckler ES (2007) TASSEL: software for association mapping of complex traits in diverse samples. Bioinformatics 23:2633–2635
Broders KD, Lipps PE, Paul PA, Dorrance AE (2007) Characterization of Pythium spp. associated with corn and soybean seed and seedling disease in Ohio. Plant Dis 91:727–735
Broders KD, Wallhead MW, Austin GD, Lipps PE, Paul PA, Mullen RW, Dorrance AE (2009) Association of soil chemical and physical properties withPythium species diversity, community composition, and disease incidence. Phytopathology 99:957–967
Churchill GA, Doerge RW (1994) Empirical threshold values for quantitative trait mapping. Genetics 138:963–971
Coser SM, Chowda Reddy RV, Zhang J, Mueller DS, Mengistu A, Wise KA, Allen TW, Singh A, Singh AK (2017) Genetic architecture of charcoal rot (Macrophomina phaseolina) resistance in soybean revealed using a diverse panel. Front Plant Sci 8:1626
Dorrance AE, Berry SA, Lipps PE (2004) Characterization ofPythium spp. from three Ohio fields for pathogenicity on corn and soybean and metalaxyl sensitivity. Plant Health Progress 5:10
Ellis ML, McHale LK, Paul PA, St. Martin SK, Dorrance AE (2013) Soybean germplasm resistant to and molecular mapping of resistance quantitative trait loci derived from the soybean accession PI 424354. Crop Science 53:1008–1021
Hartman GL, Rupe JC, Sikora EJ, Domier LL, Davis JA, Steffey KL (2016) Compendium of soybean diseases and pests, 5th edn. https://doi.org/10.1094/9780890544754
Huang X, Han B (2014) Natural variations and genome-wide association studies in crop plants. Annu Rev Plant Biol 65:531–551
Hwang EY, Song Q, Jia G, Specht JE, Hyten DL, Costa J, Cregan PB (2014) A genome-wide association study of seed protein and oil content in soybean. BMC Genomics 15:1
Klepadlo M, Balk CS, Vuong TD, Dorrance AE, Nguyen HT (2019) Molecular characterization of genomic regions for resistance to Pythium ultimum var. ultimum in the soybean cultivar Magellan. Theor Appl Genet 132:405–417
Levesque CA, Brouwer H, Cano L, Hamilton JP, Holt C, Huitema E, Raffaele S, Robideau GP, Thines M, Win J, Zerillo MM, Beakes GW, Boore JL, Busam D, Dumas B, Ferriera S, Fuerstenberg SI, Gachon CM, Gaulin E, Govers F, Grenville-Briggs L, Horner N, Hostetler J, Jiang RH, Johnson J, Krajaejun T, Lin H, Meijer HJ, Moore B, Morris P, Phuntmart V, Puiu D, Shetty J, Stajich JE, Tripathy S, Wawra S, van West P, Whitty BR, Coutinho PM, Henrissat B, Martin F, Thomas PD, Tyler BM, De Vries RP, Kamoun S, Yandell M, Tisserat N, Buell CR (2010) Genome sequence of the necrotrophic plant pathogen Pythium ultimum reveals original pathogenicity mechanisms and effector repertoire. Genome Biol 11:R73
Lin F, Zhao M, Ping J, Johnson A, Zhang B, Abney TS, Hughes TJ, Ma J (2013) Molecular mapping of two genes conferring resistance to Phytophthora sojae in a soybean landrace PI 567139B. Theor Appl Genet 126:2177–2185
Lin F, Wani SH, Collins PJ, Wen Z, Gu C, Chilvers MI, Wang D (2018) Mapping quantitative trait loci for tolerance to Pythium irregulare in soybean (Glycine max L.). G3 (Bethesda) 8:3155–3161
Liu K, Muse SV (2005) PowerMarker: an integrated analysis environment for genetic marker analysis. Bioinformatics 21:2128–2129
Matthiesen RL, Ahmad AA, Robertson AE (2016) Temperature affects aggressiveness and fungicide sensitivity of four Pythium spp. that cause soybean and corn damping off in Iowa. Plant Dis 100:583–591
May KJ, Whisson SC, Zwart RS, Searle IR, Irwin JA, Maclean DJ, Carroll BJ, Drenth A (2002) Inheritance and mapping of 11 avirulence genes inPhytophthora sojae. Fungal Genet Biol 37:1–12
Nei M, Takezaki N (1983) Estimation of genetic distances and phylogenetic trees from DNA analysis
Nyquist WE, Baker RJ (1991) Estimation of heritability and prediction of selection response in plant populations. Crit Rev Plant Sci 10:235–322
Papa KE, Campbell WA, Hendrix FF Jr (1967) Sexuality in Pythium sylvaticum: heterothallism. Mycologia 59:589–595
Pilet-Nayel ML, Moury B, Caffier V, Montarry J, Kerlan MC, Fournet S, Durel CE, Delourme R (2017) Quantitative resistance to plant pathogens in pyramiding strategies for durable crop protection. Front Plant Sci 8:1838
Price AL, Patterson NJ, Plenge RM, Weinblatt ME, Shadick NA, Reich D (2006) Principal components analysis corrects for stratification in genome-wide association studies. Nat Genet 38:904–909
Robinson GK (1991) That BLUP is a good thing: the estimation of random effects. Stat Sci 6:15–32
Rod KS, Walker DR, Bradley CA (2018) Evaluation of major ancestors of North American soybean cultivars for resistance to three Pythium species that cause seedling blight. Plant Dis 102:2241–2252
Rojas JA, Jacobs JL, Napieralski S, Karaj B, Bradley CA, Chase T, Esker PD, Giesler LJ, Jardine DJ, Malvick DK, Markell SG, Nelson BD, Robertson AE, Rupe JC, Smith DL, Sweets LE, Tenuta AU, Wise KA, Chilvers MI (2017) Oomycete species associated with soybean seedlings in North America-part II: diversity and ecology in relation to environmental and edaphic factors. Phytopathology 107:293–304
Rosso ML, Rupe JC, Chen P, Mozzoni LA (2008) Inheritance and genetic mapping of resistance to damping-off caused by in ‘archer’ soybean. Crop Sci 48:2215–2222
Serrano M, Robertson AE (2018) The effect of cold stress on damping-off of soybean caused by Pythium sylvaticum. Plant Dis 102:2194–2200
Serrano M, McDuffee D, Robertson AE (2018) Damping-off caused byPythium sylvaticum on soybeans subjected to periods of cold stress is reduced by seed treatments. Can J Plant Pathol 40:571–579
Silva Lda C, Wang S, Zeng ZB (2012) Composite interval mapping and multiple interval mapping: procedures and guidelines for using windows QTL cartographer. Methods Mol Biol 871:75–119
Stasko AK, Wickramasinghe D, Nauth BJ, Acharya B, Ellis ML, Taylor CG, McHale LK, Dorrance AE (2016) High-density mapping of resistance QTL towardPhytophthora sojae, Pythium irregulare, and Fusarium graminearum in the same soybean population. Crop Sci 56:2476–2492
Urrea K, Rupe J, Chen P, Rothrock CS (2017) Characterization of Seed Rot Resistance to in Soybean. Crop Science 57:1394–1403
Wen Z, Tan R, Yuan J, Bales C, Du W, Zhang S, Chilvers MI, Schmidt C, Song Q, Cregan PB, Wang D (2014) Genome-wide association mapping of quantitative resistance to sudden death syndrome in soybean. BMC Genomics 15:809
Wen Z, Tan R, Zhang S, Collins PJ, Yuan J, Du W, Gu C, Ou S, Song Q, An YC, Boyse JF, Chilvers MI, Wang D (2018) Integrating GWAS and gene expression data for functional characterization of resistance to white mould in soya bean. Plant Biotechnol J 16:1825–1835
Zitnick-Anderson KK, Nelson BD Jr (2015) Identification and pathogenicity of Pythium on soybean in North Dakota. Plant Dis 99:31–38
Zitnick-Anderson KK, Norland JE, Del Rio Mendoza LE, Fortuna AM, Nelson BD Jr (2017) Probability models based on soil properties for predicting presence-absence of Pythium in soybean roots. Microb Ecol 74:550–560
Acknowledgment
We thank Janette Jacobs, Randy Laurenz, John Boyse, Robert Stoutenburg, and Dr. Wenyan Du for the help in conducting this research. We are also thankful to University Grants Commission (UGC), India, for providing Raman Postdoctoral Fellowship (5–20/2016(IC)) to SHW.
Funding
We thank the funding support from Michigan Soybean Promotion Committee, USDA National Institute of Food and Agriculture (Hatch project 1011788) and AgBioResearch at Michigan State University (Project No. MICL02013).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Supplementary Figure 1
Population structures of soybean lines in GWAS panel. A: Neighbor-joining tree of 214 soybean accessions. B: PCA plots of the first two components of 214 soybean accessions. Subgroup 1 (Coser et al. 2017) contained mainly landraces from China, Japan, and Korea. Subgroup 2 (green) contained improved lines from Michigan State University and other places in the United States. Subgroup 3 (blue) mainly consisted of improved lines from Michigan State University (Supplementary Table 1). (JPG 138 kb)
Supplementary Figure 2
Genome-wide average LD decay estimated in GWAS panel. The position of euchromatin and heterochromatin of each chromosome were based on Wm82.a2 (Gmax2.0). The LD decay rate indicated by r2 was dropped to 0.2 at 354 kb and 7500 kb in euchromatic and heterochromatic regions, respectively. This result was similar with the mean LD estimated by Hwang et al. (2014) using the same method (360 kb and 9600 kb in euchromatic and heterochromatic regions, respectively). On whole chromosome level, r2 was dropped to half its maximum value at 395 kb, which was similar with the LD decay of improved lines (370 kb) reported by Wen et al. (2018) (JPG 38 kb)
Supplementary Figure 3
Genome-wide association study of partial resistance to P. sylvaticum in soybean. A: Manhattan plots for RRW-BLUP in GWAS panel. The -log10(P) values from a genome-wide scan were plotted against the position on each of the 20 chromosomes. The horizontal red line indicated the genome-wide significance threshold (FDR < 0.05).B: Quantile-quantile plot in the GWAS panel. (JPG 145 kb)
Supplementary Table 1
(XLSX 18 kb)
Supplementary Material
(XLSX 32 kb)
Rights and permissions
About this article
Cite this article
Lin, F., Wani, S.H., Collins, P.J. et al. QTL mapping and GWAS for identification of loci conferring partial resistance to Pythium sylvaticum in soybean (Glycine max (L.) Merr). Mol Breeding 40, 54 (2020). https://doi.org/10.1007/s11032-020-01133-9
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11032-020-01133-9