Abstract
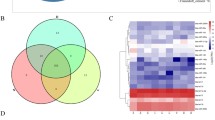
Breast milk plays an essential role for offspring development; however, there lacks evidence of how specific milk components like nucleic acids mechanistically function to regulate neonate development. Previously, we found that maternal high-fat diet (HFD) not only significantly affected mRNA and miRNA content of the secreted milk transcriptome in mice but also affected the duodenal proteome of suckling pups. Here, we hypothesized that nucleic acids differentially expressed in milk of HFD fed dams are related to differentially abundant proteins in offspring duodenum nursed by HFD dams. We tested this hypothesis by analyzing one-to-one relationships in RNA-seq data of milk transcriptomes from control (10% kcal fat) and HFD (60% kcal fat) fed mice and liquid chromatography–tandem mass spectrometry (LC-MS/MS) duodenal proteome data from pups exposed to milk. Ten percent of differentially abundant duodenal proteins between controls and HFD-exposed pups had predicted upregulation or downregulation based on differential milk RNA content. Of these, 76% were targets of upregulated miRNA, and linear regression analysis indicated relationships (p < 0.05) between multiple milk miRNA counts and duodenal protein abundance. Duodenal proteins that were potential targets of milk miRNA enriched Gene Ontology (GO) terms and KEGG pathways related to cytoskeletal structure and neural development, suggesting potential regulation of pup enteric nervous system. One-to-one relationships between milk miRNA content and protein abundance in neonate duodenum support the potential for milk miRNAs regulating neonate development. Identification of milk miRNAs that changed in response to maternal diet will enable design of mechanistic studies that test effects on neonate.




Similar content being viewed by others
References
Aghazadeh Y, Papadopoulos V (2016) The role of the 14-3-3 protein family in health, disease, and drug development. Drug Discov Today 21(2):278–287
Alsaweed M, Lai CT, Hartmann PE, Geddes DT, Kakulas F (2016) Human milk cells contain numerous miRNAs that may change with milk removal and regulate multiple physiological processes. Int J Mol Sci 17(6)
Arntz OJ, Pieters BC, Oliveira MC, Broeren MG, Bennink MB, de Vries M, van Lent PL, Koenders MI, van den Berg WB, van der Kraan PM, van de Loo FA (2015) Oral administration of bovine milk derived extracellular vesicles attenuates arthritis in two mouse models. Mol Nutr Food Res 59(9):1701–1712
Ballard O, Morrow AL (2013) Human milk composition: nutrients and bioactive factors. Pediatr Clin N Am 60(1):49–74
Breastfeeding S o (2012) Breastfeeding and the use of human milk. Pediatrics 129(3):e827–e841
Chen Y, Wang J, Yang S, Utturkar S, Crodian J, Cummings S, Thimmapuram J, San Miguel P, Kuang S, Gribskov M, Plaut K, Casey T (2017) Effect of high-fat diet on secreted milk transcriptome in midlactation mice. Physiol Genomics 49(12):747–762
Cornell B, Toyo-Oka K (2017) 14-3-3 Proteins in brain development: neurogenesis, neuronal migration and neuromorphogenesis. Front Mol Neurosci 10:318
Del Giudice M, Bo S, Grigolon S, Bosia C (2018) On the role of extrinsic noise in microRNA-mediated bimodal gene expression. PLoS Comput Biol 14(4):e1006063
Djuranovic S, Nahvi A, Green R (2011) A parsimonious model for gene regulation by miRNAs. Science 331(6017):550–553
Dobransky T, Brewer D, Lajoie G, Rylett RJ (2003) Phosphorylation of 69-kDa choline acetyltransferase at threonine 456 in response to amyloid-beta peptide 1-42. J Biol Chem 278(8):5883–5893
Franks A, Airoldi E, Slavov N (2017) Post-transcriptional regulation across human tissues. PLoS Comput Biol 13(5):e1005535
Gao N, Luo J, Uray K, Qian A, Yin S, Wang G, Wang X, Xia Y, Wood JD, Hu H (2012) CaMKII is essential for the function of the enteric nervous system. PLoS One 7(8):e44426
García-Segura L, Pérez-Andrade M, Miranda-Ríos J (2013) The emerging role of microRNAs in the regulation of gene expression by nutrients. Journal of Nutrigenetics and Nutrigenomics 6(1):16–31
Ginsberg SD, Mufson EJ, Alldred MJ, Counts SE, Wuu J, Nixon RA, Che S (2011) Upregulation of select rab GTPases in cholinergic basal forebrain neurons in mild cognitive impairment and Alzheimer’s disease. J Chem Neuroanat 42(2):102–110
Giriko C, Andreoli CA, Mennitti LV, Hosoume LF, Souto TOS, Silva AV, Mendes-da-Silva C (2013) Delayed physical and neurobehavioral development and increased aggressive and depression-like behaviors in the rat offspring of dams fed a high-fat diet. Int J Dev Neurosci 31(8):731–739
Gu Y, Li M, Wang T, Liang Y, Zhong Z, Wang X, Zhou Q, Chen L, Lang Q, He Z, Chen X, Gong J, Gao X, Li X, Lv X (2012) Lactation-related microRNA expression profiles of porcine breast milk exosomes. PLoS One 7(8):e43691
Gu Q, Cuevas E, Raymick J, Kanungo J, Sarkar S (2020) Downregulation of 14-3-3 proteins in Alzheimer’s disease. Mol Neurobiol 57(1):32–40
Hanna S, El-Sibai M (2013) Signaling networks of Rho GTPases in cell motility. Cell Signal 25(10):1955–1961
Hnia K, Ramspacher C, Vermot J, Laporte J (2015) Desmin in muscle and associated diseases: beyond the structural function. Cell Tissue Res 360(3):591–608
Hol EM, Capetanaki Y (2017) Type III intermediate filaments desmin, glial fibrillary acidic protein (GFAP), vimentin, and peripherin. Cold Spring Harb Perspect Biol 9(12)
Huangda W, Sherman BT, Lempicki RA (2009) Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat Protoc 4(1):44–57
Johnson CD, Barlow-Anacker AJ, Pierre JF, Touw K, Erickson CS, Furness JB, Epstein ML, Gosain A (2018) Deletion of choline acetyltransferase in enteric neurons results in postnatal intestinal dysmotility and dysbiosis. FASEB J 32(9):4744–4752
Kelly J, Moyeed R, Carroll C, Albani D, Li X (2019) Gene expression meta-analysis of Parkinson’s disease and its relationship with Alzheimer’s disease. Mol Brain 12(1):16
Krämer A, Green J, Pollard J, Tugendreich S (2014) Causal analysis approaches in ingenuity pathway analysis. Bioinformatics 30(4):523–530
Laubier J, Castille J, Le Guillou S, Le Provost F (2015) No effect of an elevated miR-30b level in mouse milk on its level in pup tissues. RNA Biol 12(1):26–29
Maragkakis M, Reczko M, Simossis VA, Alexiou P, Papadopoulos GL, Dalamagas T, Giannopoulos G, Goumas G, Koukis E, Kourtis K, Vergoulis T, Koziris N, Sellis T, Tsanakas P, Hatzigeorgiou AG (2009) DIANA-microT web server: elucidating microRNA functions through target prediction. Nucleic Acids Res 37(Web Server issue):W273–W276
Melnik BC, Kakulas F, Geddes DT, Hartmann PE, John SM, Carrera-Bastos P, Cordain L, Schmitz G (2016) Milk miRNAs: simple nutrients or systemic functional regulators? Nutr Metab (Lond) 13:42
Mendes-da-Silva C, Giriko C, Mennitti LV, Hosoume LF, Souto TOS, Silva AV (2014) Maternal high-fat diet during pregnancy or lactation changes the somatic and neurological development of the offspring. Arq Neuropsiquiatr 72(2):136–144
Munch EM, Harris RA, Mohammad M, Benham AL, Pejerrey SM, Showalter L, Hu M, Shope CD, Maningat PD, Gunaratne PH, Haymond M, Aagaard K (2013) Transcriptome profiling of microRNA by Next-Gen deep sequencing reveals known and novel miRNA species in the lipid fraction of human breast milk. PLoS One 8(2):e50564
Oliveros, J. C. 2007-2015. Venny. An interactive tool for comparing lists with Venn’s diagrams.
Povero D, Eguchi A, Li H, Johnson CD, Papouchado BG, Wree A, Messer K, Feldstein AE (2014) Circulating extracellular vesicles with specific proteome and liver microRNAs are potential biomarkers for liver injury in experimental fatty liver disease. PLoS One 9(12):e113651
Suarez-Trujillo A, Chen Y, Aduwari C, Cummings S, Kuang S, Buhman KK, Hedrick V, Sobreira TJP, Aryal UK, Plaut K, Casey T (2019) Maternal high-fat diet exposure during gestation, lactation, or gestation and lactation differentially affects intestinal morphology and proteome of neonatal mice. Nutr Res 66:48–60
Tang W, Cai P, Huo W, Li H, Tang J, Zhu D, Xie H, Chen P, Hang B, Wang S, Xia Y (2016) Suppressive action of miRNAs to ARP2/3 complex reduces cell migration and proliferation via RAC isoforms in Hirschsprung disease. J Cell Mol Med 20(7):1266–1275
Tian T, Zhu YL, Hu FH, Wang YY, Huang NP, Xiao ZD (2013) Dynamics of exosome internalization and trafficking. J Cell Physiol 228(7):1487–1495
Tian T, Zhu Y-L, Zhou Y-Y, Liang G-F, Wang Y-Y, Hu F-H, Xiao Z-D (2014) Exosome uptake through clathrin-mediated endocytosis and macropinocytosis and mediating miR-21 delivery. J Biol Chem 289(32):22258–22267
Title AC, Denzler R, Stoffel M (2015) Uptake and function studies of maternal milk-derived microRNAs. J Biol Chem 290(39):23680–23691
Viswanatha R, Bretscher A, Garbett D (2014) Dynamics of ezrin and EBP50 in regulating microvilli on the apical aspect of epithelial cells. Biochem Soc Trans 42(1):189–194
Vogel C, Marcotte EM (2012) Insights into the regulation of protein abundance from proteomic and transcriptomic analyses. Nat Rev Genet 13(4):227–232
Warner H, Wilson BJ, Caswell PT (2019) Control of adhesion and protrusion in cell migration by Rho GTPases. Curr Opin Cell Biol 56:64–70
Waters PS, McDermott AM, Wall D, Heneghan HM, Miller N, Newell J, Kerin MJ, Dwyer RM (2012) Relationship between circulating and tissue microRNAs in a murine model of breast cancer. PLoS One 7(11):e50459
Williams MD, Mitchell GM (2012) MicroRNAs in insulin resistance and obesity. Exp Diabetes Res 2012:8
Wolf T, Baier SR, Zempleni J (2015) The intestinal transport of bovine milk exosomes is mediated by endocytosis in human colon carcinoma Caco-2 cells and rat small intestinal IEC-6 cells. J Nutr 145(10):2201–2206
Zhou Q, Li M, Wang X, Li Q, Wang T, Zhu Q, Zhou X, Gao X, Li X (2012) Immune-related microRNAs are abundant in breast milk exosomes. Int J Biol Sci 8(1):118–123
Disclaimer
The content is solely the responsibility of the authors and does not necessarily represent the official views of the National Institutes of Health.
Funding
This project was supported, in part, by the Indiana Clinical and Translational Sciences Institute funded by Award Number UL1TR001108 from the National Institutes of Health, National Center for Advancing Translational Sciences, Clinical and Translational Sciences Award. This activity was also funded by Purdue as part of AgSEED Crossroads funding to support Indiana’s Agriculture and Rural Development.
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
ESM 1
(XLSX 57 kb)
Rights and permissions
About this article
Cite this article
Huff, K., Suárez-Trujillo, A., Kuang, S. et al. One-to-one relationships between milk miRNA content and protein abundance in neonate duodenum support the potential for milk miRNAs regulating neonate development. Funct Integr Genomics 20, 645–656 (2020). https://doi.org/10.1007/s10142-020-00743-y
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10142-020-00743-y