Abstract
Few resistance genes providing defence against the major fungal diseases septoria tritici blotch (STB), septoria nodorum blotch, leaf rust (LR), and an emerging wheat blast disease have been identified in durum wheat. We identified sixteen fungal disease-associated QTL through genome-wide association mapping of 180 inbred lines sampled from a durum wheat Composite Cross-population. Two STB resistance-associated QTL mapped to chromosome 3A, one of which colocalizes with Stb6, a known resistance gene previously identified in bread wheat. This partial resistance could be conferred by a new allele of Stb6 or another paralogous gene. The second locus is associated with a reduction in pycnidia density, a recently identified and poorly understood form of resistance. A resistance QTL strongly associated with LR, and colocalizing with Lr61, was observed in a 3.24 Mbp region on chromosome 6B. QTL mapping of LR resistance following treatment by chitin used in the context of inducer treatment was also investigated. Using a combination of resistance alleles at these loci could confer durable resistance to multiple fungal diseases and aid durum wheat breeders in their fight against these fungal pathogens.







Similar content being viewed by others
References
Adhikari TB, Jackson EW, Gurung S, Hansen JM, Bonman JM (2011) Association mapping of quantitative resistance to Phaeosphaeria nodorum in spring wheat landraces from the USDA national small grains collection. Phytopathology 101(11):1301–1310. https://doi.org/10.1094/phyto-03-11-0076
Aktar-Uz-Zaman M, Tuhina-Khatun M, Hanafi MM, Sahebi M (2017) Genetic analysis of rust resistance genes in global wheat cultivars: an overview. Biotechnol Biotechnol Equip 31(3):431–445. https://doi.org/10.1080/13102818.2017.1304180
Anh VL, Inoue Y, Asuke S, Vy TTP, Anh NT, Wang S, Chuma I, Tosa Y (2018) Rmg8 and Rmg7, wheat genes for resistance to the wheat blast fungus, recognize the same avirulence gene AVR-Rmg8. Mol Plant Pathol 19(5):1252–1256. https://doi.org/10.1111/mpp.12609
Aoun M, Breiland M, Kathryn Turner M, Loladze A, Chao S, Xu SS, Ammar K, Anderson JA, Kolmer JA, Acevedo M (2016) Genome-wide association mapping of leaf rust response in a durum wheat worldwide germplasm collection. Plant Genome. https://doi.org/10.3835/plantgenome2016.01.0008
Appels R, Eversole K, Stein N, Feuillet C, Keller B, Rogers J, Pozniak CJ, Choulet F, Distelfeld A, Poland J, Ronen G, Sharpe AG, Barad O, Baruch K, Keeble-Gagnère G, Mascher M, Ben-Zvi G, Josselin A-A, Himmelbach A, Balfourier F, Gutierrez-Gonzalez J, Hayden M, Koh C, Muehlbauer G, Pasam RK, Paux E, Rigault P, Tibbits J, Tiwari V, Spannagl M, Lang D, Gundlach H, Haberer G, Mayer KFX, Ormanbekova D, Prade V, Šimková H, Wicker T, Swarbreck D, Rimbert H, Felder M, Guilhot N, Kaithakottil G, Keilwagen J, Leroy P, Lux T, Twardziok S, Venturini L, Juhász A, Abrouk M, Fischer I, Uauy C, Borrill P, Ramirez-Gonzalez RH, Arnaud D, Chalabi S, Chalhoub B, Cory A, Datla R, Davey MW, Jacobs J, Robinson SJ, Steuernagel B, van Ex F, Wulff BBH, Benhamed M, Bendahmane A, Concia L, Latrasse D, Bartoš J, Bellec A, Berges H, Doležel J, Frenkel Z, Gill B, Korol A, Letellier T, Olsen O-A, Singh K, Valárik M, van der Vossen E, Vautrin S, Weining S, Fahima T, Glikson V, Raats D, Číhalíková J, Toegelová H, Vrána J, Sourdille P, Darrier B, Barabaschi D, Cattivelli L, Hernandez P, Galvez S, Budak H, Jones JDG, Witek K, Yu G, Small I, Melonek J, Zhou R, Belova T, Kanyuka K, King R, Nilsen K, Walkowiak S, Cuthbert R, Knox R, Wiebe K, Xiang D, Rohde A, Golds T, Čížková J, Akpinar BA, Biyiklioglu S, Gao L, N’Daiye A, Kubaláková M, Šafář J, Alfama F, Adam-Blondon A-F, Flores R, Guerche C, Loaec M, Quesneville H, Condie J, Ens J, Maclachlan R, Tan Y, Alberti A, Aury J-M, Barbe V, Couloux A, Cruaud C, Labadie K, Mangenot S, Wincker P, Kaur G, Luo M, Sehgal S, Chhuneja P, Gupta OP, Jindal S, Kaur P, Malik P, Sharma P, Yadav B, Singh NK, Khurana JP, Chaudhary C, Khurana P, Kumar V, Mahato A, Mathur S, Sevanthi A, Sharma N, Tomar RS, Holušová K, Plíhal O, Clark MD, Heavens D, Kettleborough G, Wright J, Balcárková B, Hu Y, Salina E, Ravin N, Skryabin K, Beletsky A, Kadnikov V, Mardanov A, Nesterov M, Rakitin A, Sergeeva E, Handa H, Kanamori H, Katagiri S, Kobayashi F, Nasuda S, Tanaka T, Wu J, Cattonaro F, Jiumeng M, Kugler K, Pfeifer M, Sandve S, Xun X, Zhan B, Batley J, Bayer PE, Edwards D, Hayashi S, Tulpová Z, Visendi P, Cui L, Du X, Feng K, Nie X, Tong W, Wang L (2018) Shifting the limits in wheat research and breeding using a fully annotated reference genome. Science 361(6403):eaar7191. https://doi.org/10.1126/science.aar7191
Astle W, Balding DJ (2009) Population structure and cryptic relatedness in genetic association studies. Stat Sci 24(4):451–471
Avni R, Nave M, Barad O, Baruch K, Twardziok SO, Gundlach H, Hale I, Mascher M, Spannagl M, Wiebe K, Jordan KW, Golan G, Deek J, Ben-Zvi B, Ben-Zvi G, Himmelbach A, MacLachlan RP, Sharpe AG, Fritz A, Ben-David R, Budak H, Fahima T, Korol A, Faris JD, Hernandez A, Mikel MA, Levy AA, Steffenson B, Maccaferri M, Tuberosa R, Cattivelli L, Faccioli P, Ceriotti A, Kashkush K, Pourkheirandish M, Komatsuda T, Eilam T, Sela H, Sharon A, Ohad N, Chamovitz DA, Mayer KFX, Stein N, Ronen G, Peleg Z, Pozniak CJ, Akhunov ED, Distelfeld A (2017) Wild emmer genome architecture and diversity elucidate wheat evolution and domestication. Science 357(6346):93–97. https://doi.org/10.1126/science.aan0032
Ballini E, Nguyen TT, Morel J-B (2013) Diversity and genetics of nitrogen-induced susceptibility to the blast fungus in rice and wheat. Rice 6(1):32
Bancal M-O, Ben Slimane R, Bancal P (2016) Zymoseptoria tritici development induces local senescence in wheat leaves, without affecting their monocarpic senescence under two contrasted nitrogen nutrition. Environ Exp Bot 132:154–162. https://doi.org/10.1016/j.envexpbot.2016.09.002
Bearchell SJ, Fraaije BA, Shaw MW, Fitt BD (2005) Wheat archive links long-term fungal pathogen population dynamics to air pollution. Proc Natl Acad Sci 102(15):5438–5442
Berruyer R, Adreit H, Milazzo J, Gaillard S, Berger A, Dioh W, Lebrun MH, Tharreau D (2003) Identification and fine mapping of Pi33, the rice resistance gene corresponding to the Magnaporthe grisea avirulence gene ACE1. TAG Theor Appl Genet 1072:1139–1147
Bohland C, Balkenhohl T, Loers G, Feussner I, Grambow HJ (1997) Differential induction of lipoxygenase isoforms in wheat upon treatment with rust fungus elicitor, chitin oligosaccharides, chitosan, and methyl jasmonate. Plant Physiol 114(2):679–685
Brown JKM, Chartrain L, Lasserre-Zuber P, Saintenac C (2015) Genetics of resistance to Zymoseptoria tritici and applications to wheat breeding. Fungal Genet Biol 79:33–41. https://doi.org/10.1016/j.fgb.2015.04.017
Brown-Guedira GL, Singh S, Fritz AK (2003) Performance and mapping of leaf rust resistance transferred to wheat from Triticum timopheevii subsp. armeniacum. Phytopathology 93(7):784–789. https://doi.org/10.1094/phyto.2003.93.7.784
Brunner PC, McDonald BA (2018) Evolutionary analyses of the avirulence effector AvrStb6 in global populations of Zymoseptoria tritici identify candidate amino acids involved in recognition. Mol Plant Pathol 19(8):1836–1846. https://doi.org/10.1111/mpp.12662
Ceresini PC, Castroagudín VL, Rodrigues FÁ, Rios JA, Aucique-Pérez CE, Moreira SI, Alves E, Croll D, Maciel JLN (2018) Wheat blast: past, present, and future. Annu Rev Phytopathol 56(1):427–456. https://doi.org/10.1146/annurev-phyto-080417-050036
Ceresini PC, Castroagudín VL, Rodrigues FÁ, Rios JA, Aucique-Pérez CE, Moreira SI, Croll D, Alves E, de Carvalho G, Maciel JLN, McDonald BA (2019) Wheat blast: from its origins in South America to its emergence as a global threat. Mol Plant Pathol 20(2):155–172. https://doi.org/10.1111/mpp.12747
Chern M, Canlas PE, Fitzgerald HA, Ronald PC (2005) Rice NRR, a negative regulator of disease resistance, interacts with Arabidopsis NPR1 and rice NH1. Plant J 43(5):623–635
Crossa J, Burgueno J, Dreisigacker S, Vargas M, Herrera-Foessel SA, Lillemo M, Singh RP, Trethowan R, Warburton M, Franco J, Reynolds M, Crouch JH, Ortiz R (2007) Association analysis of historical bread wheat germplasm using additive genetic covariance of relatives and population structure. Genetics 177(3):1889–1913
Cruz M, Diniz A, Rodrigues F, Barros E (2011) Foliar application of products on the reduction of blast severity on wheat. Trop Plant Pathol 36:424–428. https://doi.org/10.1590/S1982-56762011000600014
Cullis BR, Smith AB, Coombes N (2006) On the design of early generation variety trials with corrected data. J Agric Biol Environ Stat 11:381–393. https://doi.org/10.1198/108571106X154443
Dadkhodaie NA, Karaoglou H, Wellings CR, Park RF (2011) Mapping genes Lr53 and Yr35 on the short arm of chromosome 6B of common wheat with microsatellite markers and studies of their association with Lr36. Theor Appl Genet 122(479):1432–2242
David J, Holtz Y, Ranwez V, Santoni S, Sarah G, Ardisson M, Poux G, Choulet F, Genthon C, Roumet P, Tavaud-Pirra M (2014) Genotyping by sequencing transcriptomes in an evolutionary pre-breeding durum wheat population. Mol Breed 34(4):1531–1548
Du Fall LA, Solomon PS (2013) The necrotrophic effector SnToxA induces the synthesis of a novel phytoalexin in wheat. New Phytol 200(1):185–200. https://doi.org/10.1111/nph.12356
Dvořák J, Knott D (1990) Location of a Triticum speltoides chromosome segment conferring resistance to leaf rust in Triticum aestivum. Genome 33(6):892–897
Enjalbert J, Dawson JC, Paillard S, Rhoné B, Rousselle Y, Thomas M, Goldringer I (2011) Dynamic management of crop diversity: from an experimental approach to on-farm conservation. CR Biol 334(5–6):458–468
Erland LAE, Turi CE, Saxena PK (2016) Serotonin: an ancient molecule and an important regulator of plant processes. Biotechnol Adv 34(8):1347–1361. https://doi.org/10.1016/j.biotechadv.2016.10.002
Figueroa M, Hammond-Kosack KE, Solomon PS (2018) A review of wheat diseases—a field perspective. Mol Plant Pathol 19(6):1523–1536. https://doi.org/10.1111/mpp.12618
Fones H, Gurr S (2015) The impact of Septoria tritici blotch disease on wheat: an EU perspective. Fungal Genet Biol 79:3–7
Fones HN, Steinberg G, Gurr SJ (2015) Measurement of virulence in Zymoseptoria tritici through low inoculum-density assays. Fungal Genet Biol 79:89–93. https://doi.org/10.1016/j.fgb.2015.03.020
Francki MG (2013) Improving Stagonospora nodorum resistance in wheat: a review. Crop Sci 53(2):355–365. https://doi.org/10.2135/cropsci2012.06.0347
Gao L, Turner MK, Chao S, Kolmer J, Anderson JA (2016) Genome wide association study of seedling and adult plant leaf rust resistance in elite spring wheat breeding lines. PLoS ONE 11(2):e0148671. https://doi.org/10.1371/journal.pone.0148671
Ghaffary SMT, Chawade A, Singh PK (2018) Practical breeding strategies to improve resistance to Septoria tritici blotch of wheat. Euphytica 214(7):122. https://doi.org/10.1007/s10681-018-2205-4
Goyeau H, Park R, Schaeffer B, Lannou C (2006) Distribution of pathotypes with regard to host cultivars in french wheat leaf rust populations. Phytopathology 96(3):264–273. https://doi.org/10.1094/PHYTO-96-0264
Goyeau H, Berder J, Czerepak C, Gautier A, Lanen C, Lannou C (2012) Low diversity and fast evolution in the population of Puccinia triticina causing durum wheat leaf rust in France from 1999 to 2009, as revealed by an adapted differential set. Plant Pathol 61(4):761–772. https://doi.org/10.1111/j.1365-3059.2011.02554.x
Gurung S, Mamidi S, Bonman JM, Xiong M, Brown-Guedira G, Adhikari TB (2014) Genome-wide association study reveals novel quantitative trait loci associated with resistance to multiple leaf spot diseases of spring wheat. PLoS ONE 9(9):e108179. https://doi.org/10.1371/journal.pone.0108179
Haudry A, Cenci A, Ravel C, Bataillon T, Brunel D, Poncet C, Hochu I, Poirier S, Santoni S, Glémin S (2007) Grinding up wheat: a massive loss of nucleotide diversity since domestication. Mol Biol Evol 24(7):1506–1517
Haugrud PAR, Zhang Z, Richards JK, Friesen TL, Faris JD (2019) Genetics of variable disease expression conferred by inverse gene-for-gene interactions in the wheat-Parastagonospora nodorum pathosystem. Plant Physiol 180(1):420–434. https://doi.org/10.1104/pp.19.00149
Herrera-Foessel S, Singh R, Huerta-Espino J, William H, Djurle A, Yuen J (2008) Molecular mapping of a leaf rust resistance gene on the short arm of chromosome 6B of durum wheat. Plant Dis 92(12):1650–1654
Hussein S, Spies JJ, Pretorius ZA, Labuschagne MT (2005) Chromosome locations of leaf rust resistance genes in selected tetraploid wheats through substitution lines. Euphytica 141(3):209–216
Ishihara A, Matsukawa T, Nomura T, Sue M, Oikawa A, Okazaki Y, Tebayashi S (2015) Involvement of tryptophan-pathway-derived secondary metabolism in the defence responses of grasses. In: Dmello JPF (ed) Amino acids in higher plants. Cabi Publishing-C a B Int, Wallingford, pp 362–389
Jighly A, Alagu M, Makdis F, Singh M, Singh S, Emebiri LC, Ogbonnaya FC (2016) Genomic regions conferring resistance to multiple fungal pathogens in synthetic hexaploid wheat. Mol Breed 36(9):127. https://doi.org/10.1007/s11032-016-0541-4
Juliana P, Singh RP, Singh PK, Poland JA, Bergstrom GC, Huerta-Espino J, Bhavani S, Crossa J, Sorrells ME (2018) Genome-wide association mapping for resistance to leaf rust, stripe rust and tan spot in wheat reveals potential candidate genes. Theor Appl Genet 131(7):1405–1422. https://doi.org/10.1007/s00122-018-3086-6
Karisto P, Hund A, Yu K, Anderegg J, Walter A, Mascher F, McDonald BA, Mikaberidze A (2017) Ranking quantitative resistance to Septoria tritici blotch in elite wheat cultivars using automated image analysis. Phytopathology 108(5):568–581. https://doi.org/10.1094/PHYTO-04-17-0163-R
Kema GHJ, Annone JG, Rachid S, Van Silfhout CH, Van Ginkel M, De Bree J (1996) Genetic variation for virulence and resistance in the wheat-Mycosphaerella graminicola pathosystem I. Interactions between pathogen isolates and host cultivars. Phytopathology. https://doi.org/10.1094/phyto-86-200
Kertho A, Mamidi S, Bonman JM, McClean PE, Acevedo M (2015) Genome-wide association mapping for resistance to leaf and stripe rust in winter-habit hexaploid wheat landraces. PLoS ONE 10(6):e0129580
Kidane YG, Hailemariam BN, Mengistu DK, Fadda C, Pè ME, Dell’Acqua M (2017) Genome-wide association study of Septoria tritici blotch resistance in ethiopian durum wheat landraces. Front Plant Sci 8:1586. https://doi.org/10.3389/fpls.2017.01586
Korte A, Farlow A (2013) The advantages and limitations of trait analysis with GWAS: a review. Plant Methods 9(1):29. https://doi.org/10.1186/1746-4811-9-29
Krattinger SG, Sucher J, Selter LL, Chauhan H, Zhou B, Tang M, Upadhyaya NM, Mieulet D, Guiderdoni E, Weidenbach D, Schaffrath U, Lagudah ES, Keller B (2016) The wheat durable, multipathogen resistance gene Lr34 confers partial blast resistance in rice. Plant Biotechnol J 14(5):1261–1268. https://doi.org/10.1111/pbi.12491
Kthiri D, Loladze A, MacLachlan PR, N’Diaye A, Walkowiak S, Nilsen K, Dreisigacker S, Ammar K, Pozniak CJ (2018) Characterization and mapping of leaf rust resistance in four durum wheat cultivars. PLoS ONE 13(5):e0197317. https://doi.org/10.1371/journal.pone.0197317
Letta T, Olivera P, Maccaferri M, Jin Y, Ammar K, Badebo A, Salvi S, Noli E, Crossa J, Tuberosa R (2014) Association mapping reveals novel stem rust resistance loci in durum wheat at the seedling stage. Plant Genome. https://doi.org/10.3835/plantgenome2013.08.0026
Ling H-Q, Ma B, Shi X, Liu H, Dong L, Sun H, Cao Y, Gao Q, Zheng S, Li Y, Yu Y, Du H, Qi M, Li Y, Lu H, Yu H, Cui Y, Wang N, Chen C, Wu H, Zhao Y, Zhang J, Li Y, Zhou W, Zhang B, Hu W, van Eijk MJT, Tang J, Witsenboer HMA, Zhao S, Li Z, Zhang A, Wang D, Liang C (2018) Genome sequence of the progenitor of wheat A subgenome Triticum urartu. Nature 557(7705):424–428. https://doi.org/10.1038/s41586-018-0108-0
Liu ZH, Faris JD, Meinhardt SW, Ali S, Rasmussen JB, Friesen TL (2004) Genetic and physical mapping of a gene conditioning sensitivity in wheat to a partially purified host-selective toxin produced by Stagonospora nodorum. Phytopathology 94(10):1056–1060. https://doi.org/10.1094/phyto.2004.94.10.1056
Liu Z, Zhang Z, Faris JD, Oliver RP, Syme R, McDonald MC, McDonald BA, Solomon PS, Lu S, Shelver WL, Xu S, Friesen TL (2012) The cysteine rich necrotrophic effector SnTox1 produced by Stagonospora nodorum triggers susceptibility of wheat lines harboring Snn1. PLoS Pathog 8(1):e1002467. https://doi.org/10.1371/journal.ppat.1002467
Liu W, Maccaferri M, Chen X, Laghetti G, Pignone D, Pumphrey M, Tuberosa R (2017) Genome-wide association mapping reveals a rich genetic architecture of stripe rust resistance loci in emmer wheat (Triticum turgidum ssp. dicoccum). Theor Appl Genet 130(11):2249–2270. https://doi.org/10.1007/s00122-017-2957-6
Loladze A, Kthiri D, Pozniak C, Ammar K (2014) Genetic analysis of leaf rust resistance in six durum wheat genotypes. Phytopathology 104(12):1322–1328
Maccaferri M, Sanguineti MC, Mantovani P, Demontis A, Massi A, Ammar K, Kolmer JA, Czembor JH, Ezrati S, Tuberosa R (2010) Association mapping of leaf rust response in durum wheat. Mol Breed 26(2):189–228. https://doi.org/10.1007/s11032-009-9353-0
Maccaferri M, Harris NS, Twardziok SO, Pasam RK, Gundlach H, Spannagl M, Ormanbekova D, Lux T, Prade VM, Milner SG, Himmelbach A, Mascher M, Bagnaresi P, Faccioli P, Cozzi P, Lauria M, Lazzari B, Stella A, Manconi A, Gnocchi M, Moscatelli M, Avni R, Deek J, Biyiklioglu S, Frascaroli E, Corneti S, Salvi S, Sonnante G, Desiderio F, Marè C, Crosatti C, Mica E, Özkan H, Kilian B, De Vita P, Marone D, Joukhadar R, Mazzucotelli E, Nigro D, Gadaleta A, Chao S, Faris JD, Melo ATO, Pumphrey M, Pecchioni N, Milanesi L, Wiebe K, Ens J, MacLachlan RP, Clarke JM, Sharpe AG, Koh CS, Liang KYH, Taylor GJ, Knox R, Budak H, Mastrangelo AM, Xu SS, Stein N, Hale I, Distelfeld A, Hayden MJ, Tuberosa R, Walkowiak S, Mayer KFX, Ceriotti A, Pozniak CJ, Cattivelli L (2019) Durum wheat genome highlights past domestication signatures and future improvement targets. Nat Genet. https://doi.org/10.1038/s41588-019-0381-3
McDonald BA, Linde C (2002) Pathogen population genetics, evolutionary potential and durable resistance. Annu Rev Phytopathol 40(1):349–379. https://doi.org/10.1146/annurev.phyto.40.120501.101443
Muqaddasi QH, Zhao Y, Rodemann B, Plieske J, Ganal MW, Röder MS (2019) Genome-wide association mapping and prediction of adult stage Septoria tritici blotch infection in European winter wheat via high-density marker arrays. Plant Genome. https://doi.org/10.3835/plantgenome2018.05.0029
Naz R, Bano A, Wilson NL, Guest D, Roberts TH (2014) Pathogenesis-related protein expression in the apoplast of wheat leaves protected against leaf rust following application of plant extracts. Phytopathology 104(9):933–944. https://doi.org/10.1094/PHYTO-11-13-0317-R
Neumann K, Kobiljski B, Denčić S, Varshney RK, Börner A (2011) Genome-wide association mapping: a case study in bread wheat (Triticum aestivum L.). Mol Breed 27(1):37–58. https://doi.org/10.1007/s11032-010-9411-7
Ordonez M, Kolmer JA (2007) Virulence phenotypes of a worldwide collection of Puccinia triticina from durum wheat. Phytopathology 97(3):344–351
Ors ME, Randoux B, Selim S, Siah A, Couleaud G, Maumené C, Sahmer K, Halama P, Reignault P (2018) Cultivar-dependent partial resistance and associated defence mechanisms in wheat against Zymoseptoria tritici. Plant Pathol 67(3):561–572. https://doi.org/10.1111/ppa.12760
Phan HTT, Rybak K, Furuki E, Breen S, Solomon PS, Oliver RP, Tan K-C (2016) Differential effector gene expression underpins epistasis in a plant fungal disease. Plant J 87(4):343–354. https://doi.org/10.1111/tpj.13203
Popova EV, Domnina NS, Kovalenko NM, Sokornova S, Tyuterev SL (2018) Influence of chitosan hybrid derivatives on induced wheat resistance to pathogens with different nutrition strategies. Appl Biochem Microbiol 54(5):535–539. https://doi.org/10.1134/S0003683818050150
Qureshi N, Bariana H, Kolmer JA, Miah H, Bansal U (2017) Genetic and molecular characterization of leaf rust resistance in two durum wheat landraces. Phytopathology 107(11):1381–1387. https://doi.org/10.1094/phyto-01-17-0005-r
Rimbert H, Darrier B, Navarro J, Kitt J, Choulet F, Leveugle M, Duarte J, Rivière N, Eversole K, on behalf of The International Wheat Genome Sequencing C, Le Gouis J, on behalf The BreedWheat C, Davassi A, Balfourier F, Le Paslier M-C, Berard A, Brunel D, Feuillet C, Poncet C, Sourdille P, Paux E (2018) High throughput SNP discovery and genotyping in hexaploid wheat. PLoS ONE 13(1):e0186329. https://doi.org/10.1371/journal.pone.0186329
Royo C, Di Fonzo N (2005) Durum wheat breeding. Current approaches and future strategies, vol 1–2. CRC Press, Boca Raton
Saharan MS, Tiwari R (2011) Durable resistance in wheat. Int J Genet Mol Biol 3:108–114
Saintenac C, Lee W-S, Cambon F, Rudd JJ, King RC, Marande W, Powers SJ, Bergès H, Phillips AL, Uauy C, Hammond-Kosack KE, Langin T, Kanyuka K (2018) Wheat receptor-kinase-like protein Stb6 controls gene-for-gene resistance to fungal pathogen Zymoseptoria tritici. Nat Genet 50(3):368–374. https://doi.org/10.1038/s41588-018-0051-x
Saitoh H, Kanzaki H, Fujisaki K, Takagi H, Yoshida K, Terauchi R (2016) Rice blast resistance-inducing mechanisms < i>via </i > effector recognition by NLR immune receptors. Jpn J Phytopathol 82(4):296–300. https://doi.org/10.3186/jjphytopath.82.296
Shi G, Zhang Z, Friesen TL, Raats D, Fahima T, Brueggeman RS, Lu S, Trick HN, Liu Z, Chao W, Frenkel Z, Xu SS, Rasmussen JB, Faris JD (2016) The hijacking of a receptor kinase-driven pathway by a wheat fungal pathogen leads to disease. Sci Adv 2(10):e1600822. https://doi.org/10.1126/sciadv.1600822
Tagle AG, Chuma I, Tosa Y (2015) Rmg7, a new gene for resistance to triticum isolates of Pyricularia oryzae identified in tetraploid wheat. Phytopathology 105(4):495–499. https://doi.org/10.1094/PHYTO-06-14-0182-R
Tommasini L, Schnurbusch T, Fossati D, Mascher F, Keller B (2007) Association mapping of Stagonospora nodorum blotch resistance in modern European winter wheat varieties. Theor Appl Genet 115(5):697–708. https://doi.org/10.1007/s00122-007-0601-6
Vander P, Vårum KM, Domard A, Eddine El Gueddari N, Moerschbacher BM (1998) Comparison of the ability of partially N-acetylated chitosans and chitooligosaccharides to elicit resistance reactions in wheat leaves. Plant Physiol 118(4):1353–1359. https://doi.org/10.1104/pp.118.4.1353
Velazco JG, Rodríguez-Álvarez M, Boer MP, Jordan D, Eilers PHC, Malosetti M, Eeuwijk F (2017) Modelling spatial trends in sorghum breeding field trials using a two-dimensional P-spline mixed model. Theor Appl Genet 130(7):1375–1392. https://doi.org/10.1007/s00122-017-2894-4
Virdi SK, Liu Z, Overlander ME, Zhang Z, Xu SS, Friesen TL, Faris JD (2016) New Insights into the roles of host gene-necrotrophic effector interactions in governing susceptibility of durum wheat to tan spot and Septoria nodorum blotch. G3 (Bethesda, Md) 6(12):4139–4150. https://doi.org/10.1534/g3.116.036525
Yates S, Mikaberidze A, Krattinger SG, Abrouk M, Hund A, Yu K, Studer B, Fouche S, Meile L, Pereira D, Karisto P, McDonald BA (2019) Precision phenotyping reveals novel loci for quantitative resistance to Septoria tritici blotch. Plant Phenom 2019:11. https://doi.org/10.34133/2019/3285904
Zhan J, Linde CC, Jurgens T, Merz U, Steinebrunner F, McDonald BA (2005) Variation for neutral markers is correlated with variation for quantitative traits in the plant pathogenic fungus Mycosphaerella graminicola. Mol Ecol 14(9):2683–2693. https://doi.org/10.1111/j.1365-294X.2005.02638.x
Zhan SW, Mayama S, Tosa Y (2008) Identification of two genes for resistance to Triticum isolates of Magnaporthe oryzae in wheat. Genome 51(3):216–221. https://doi.org/10.1139/g07-094
Zhong Z, Marcel TC, Hartmann FE, Ma X, Plissonneau C, Zala M, Ducasse A, Confais J, Compain J, Lapalu N, Amselem J, McDonald BA, Croll D, Palma-Guerrero J (2017) A small secreted protein in Zymoseptoria tritici is responsible for avirulence on wheat cultivars carrying the Stb6 resistance gene. New Phytol 214(2):619–631
Zhou X (2017) A unified framework for variance component estimation with summary statistics in genome-wide association studies. Ann Appl Stat 11(4):2027–2051. https://doi.org/10.1214/17-aoas1052
Zhou X, Stephens M (2012) Genome-wide efficient mixed-model analysis for association studies. Nat Genet 44(7):821–824. https://doi.org/10.1038/ng.2310
Zhu T, Wang L, Rodriguez JC, Deal KR, Avni R, Distelfeld A, McGuire PE, Dvorak J, Luo M-C (2019) Improved genome sequence of wild emmer wheat Zavitan with the aid of optical maps. G3 (Bethesda, Md) 9(3):619–624. https://doi.org/10.1534/g3.118.200902
Acknowledgements
Highbred, CASDAR-BURRITOS, FSOV-WEAB and in particular Richard Oliver (Curtin University, Australia) for providing purified Tox1.
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
ESM_1
Manhattan plots from association mapping in the EPO durum wheat panel of 180 lines for SNB and WB traits. The map is based on Zavitan v2.0. A1 Septoria nodorum blotch SNB isolate inoculation chromosome A, B1 chromosome B. A2 Septoria nodorum blotch Tox1 infiltration chromosome A, B2 chromosome B. A3 Wheat blast BR43 isolate inoculation chromosome A, B3 chromosome B. A4 Wheat blast BR32 isolate inoculation chromosome A, B4 chromosome B (TIFF 453 kb)
ESM_2
Manhattan plots from association mapping in the EPO durum wheat panel of 180 lines for STB and LR traits. The map is based on Zavitan v2.0. A1 Septoria tritici blotch STB isolate inoculation, % of surface with necrose, chromosome A, B1 chromosome B. A2 Septoria tritici blotch STB isolate inoculation, % of surface with pycnidia chromosome A, B2 chromosome B. A3 Leaf rust isolate inoculation, mock treated 6 h before inoculation, chromosome A, B3 chromosome B. A4 Leaf rust isolate inoculation, chitin treated 6 h before inoculation, chromosome A, B4 chromosome B (TIFF 363 kb)
ESM_3
Annotation of candidate genes for SNB_Tox7A and SNB_Nec4B. Annotation and genome location are based on Zavitan v2.0 (XLSX 18 kb)
ESM_4
Histogram of STB symptoms in EPO lines. The % of diseased leaf area was recorded for necrosis symptoms (dark grey) and % of surface with pycnidia for pycnidia production (light grey) (TIFF 54 kb)
ESM_5
A Number of EPO lines carrying each allele at the STB_Pyc3A and STB_Nec3A loci. B Number of EPO lines and Elite lines resistant for pycnidia density, depending on their genotype at STB_Pyc3A. The number of resistant lines and the total number of lines carrying the corresponding allele are represented. The frequency of the resistant phenotype is in parenthesis. C Resistance of selected EPO lines and Elite lines based on percentage of necrotic leaf area and depending of their genotype at STB_Nec3A and at Stb6. The number of resistant lines and the total number of lines carrying the corresponding allele are represented. The frequency of the resistant phenotype is shown in parenthesis (JPEG 95 kb)
ESM_6
A Sequence of the kinase at Stb6 locus. The three alleles identified in this study (Stb6-7, Stb6-1, and Stb6-12) were aligned with seven previously published alleles (Saintenac et al. 2018). B Amino acid changes due to the different nucleotide polymorphism are indicated (TIFF 1274 kb)
ESM_7
Sequence alignment of AvrStb6 allele in both isolates P1a and M1a. Two other isolates carrying AvrStb6 allele (1E4, 323) and 1A5 (VirStb6) were sequenced and aligned on three of the published sequences (Hap2, Hap3 and Hap11) found in Brunner and McDonald 2018 (TIFF 53 kb)
ESM_8
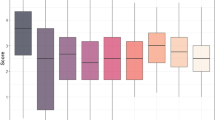
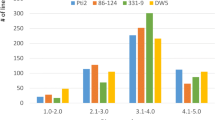
A Distribution of leaf rust severity evaluated as the percentage of leaf surface covered with pustules after prophylactic induction with chitin treatment (dark grey) or following mock treatment (light grey). B Boxplot of leaf rust severity for completely resistant and partially resistant lines. Complete resistance was considered when no pustules were observed on the leaf surface (JPEG 31 kb)
Rights and permissions
About this article
Cite this article
Ballini, E., Tavaud, M., Ducasse, A. et al. Genome wide association mapping for resistance to multiple fungal pathogens in a panel issued from a broad composite cross-population of tetraploid wheat Triticum turgidum. Euphytica 216, 92 (2020). https://doi.org/10.1007/s10681-020-02631-9
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s10681-020-02631-9