Abstract
Structural proteins, including silk fibroins, play an important role in shaping the skeletons and structures of cells, tissues, and organisms. The amino acid sequences of structural proteins often show characteristic features, such as a repeating tandem motif, that are notably different from those of functional proteins such as enzymes and antibodies. In recent years, materials composed of or containing structural proteins have been studied and developed as biomedical, apparel, and structural materials. This review outlines the definition of structural proteins, methods for characterizing structural proteins as polymeric materials, and potential applications.
Similar content being viewed by others
Definition
Proteins are polymers in which the 20 natural amino acids are linked by amide bonds. In addition to the 20 natural amino acids, there are amino acids that are not directly synthesized from ribosomes, such as l-3,4-dihydroxyphenylalanine (DOPA), hydroxyproline (Hyp), dityrosine, and selenomethionine, and these compounds are synthesized via posttranslational modifications. These nonribosomal peptides and amino acids often play an important role in structural and functional proteins. In many cases, structural proteins have a characteristic amino acid sequence that repeats to form a higher-order structure by intermolecular and/or intramolecular hydrogen bonding [1]. One example is the disulfide bond formed between two cysteine residues, which can induce dimerization and hierarchical structures in proteins [2]. However, in general, the amino acid sequence and the structure being formed depend on the type of structural protein, and it is difficult to identify a structural protein based on the sequence or structure alone. In particular, when nonnatural sequences created by gene recombination techniques and chemical synthesis are considered [3], defining a structural protein is even more difficult and complicated. Therefore, in this focus review, I would like to define a structural protein as “a protein that possesses a characteristic amino acid sequence or motif that repeats and forms a skeleton or contributes to the mechanical properties of a living organism, cell, or material” (Fig. 1). Structural proteins can be globular or fibrillar proteins. In most cases, such motifs are expected to form higher-order structures by intermolecular or intramolecular interactions, leading to the expression of physical characteristics. However, considering protein-based amorphous materials, the requirement of a hierarchical structure should not be included in this definition.
Natural structural proteins
To mimic and develop structural protein-like materials, scientists and researchers need to understand the molecular and hierarchical mechanisms influencing natural structural proteins. In this section, several natural structural proteins are reviewed from the perspective of biological polymeric materials.
Spider silk
Many types of spiders are known to produce multiple types of fibers, and some spin up to 7 types (Fig. 2) [4,5,6]. The thread spun from the major ampullate gland is called dragline and is used as a framework for reticular spider webs and as lifelines. As can be inferred from their role in nature, spider draglines exhibit excellent mechanical properties and are known as typical “tough spider silk fibers”. In particular, the toughness of these fibers in terms of resistance to breakage due to stretching is significantly superior to that of synthetic polymers and other natural materials because of their relatively high strength and elongation at break [7, 8]; hence, these fibers are expected to be applicable in lightweight structural materials such as automotive parts. The mechanical and physical properties of these fibers are significantly influenced by the deformation rate and humidity [9,10,11]. Unlike silkworm silk fibers, the physical properties of spider dragline, namely, its supercontraction, are influenced by interactions with water molecules. Recently, proline and diglutamine in the amino acid sequences have been proposed to contribute to supercontraction based on comparative studies of the supercontraction of different spider silks [9]. Spider dragline silk has a tandem sequence of polyalanine-rich and glycine-rich sequences. Polyalanine is a potentially crystalline motif, whereas glycine-rich sequences form amorphous regions. This characteristic repeating motif allows the formation of higher-order structures in which microcrystals comprising beta sheet structures are highly oriented along the fiber axis [12,13,14].
Silkworm silk and other silks
Silk protein is produced not only by spiders but also by animals such as silkworms and scorpions (Fig. 2). The presence of beta sheet structures as crystal components is similar, but the amino acid sequences and physical properties vary markedly among the different species. Silkworm silk fiber is known to be coated with the adhesive protein sericin, and many studies focusing on the material properties of sericin have been reported [15]. In recent years, a consortium of Asian countries has reevaluated silk as a polymer material and reported the variety of physical properties offered by silkworm silk thread (Fig. 3) [16]. Differences in mechanical and thermal properties can occur due to variations in the hierarchical structure resulting from the amino acid sequence of the silk protein. As silk fibers have been used as sutures, silk is attractive as a biomedical material for regenerative medicine, tissue engineering, and drug delivery [17, 18] because of its excellent biocompatibility, biodegradability, and low cytotoxicity [19, 20]. In addition, some silks have an arginine-glycine-aspartic acid (RGD) sequence, which contributes to cell adhesion, making them useful for biomaterials [21,22,23].
Comparison of phylogenetic trees derived from silkworm silks. Japan (Jp), China (Cn), Thailand (Th), and India (In) denote the country where the silk was sampled. Reprinted with permission from ref. [16]
The relationships among the amino acid sequences, secondary structures, and thermal properties of silkworm silk have been investigated and are summarized in Fig. 4. The predominant secondary structure can be polyglycine II, polyglycine I, or beta sheet structures depending on the composition of alanine and glycine in the copolymer, as these are the amino acids that make up the crystal region of the silk, and the pyrolysis behavior is also influenced by the alanine/glycine composition [24]. This correlation has been confirmed not only in model peptides obtained by copolymerization but also in silkworm-derived natural silk fibers [16]. This result suggests that differences in the partial polypeptide sequences in proteins greatly affect the material’s physical properties, especially for silk materials, for which the thermal and mechanical properties are fully determined by the crystalline sequences. This is recognized as an important finding for the molecular-level design of silk materials with the desired thermophysical properties. In this way, the basic information required for controlling the physical and mechanical properties of structural proteins such as silk is gradually being reported.
Collagen
Collagen is one of the most abundant proteins in mammals and is a major structural component of extracellular matrices. It contains many periodically repeating 3-amino-acid sequences containing Gly. The Gly-containing repeating sequences include ~1400 amino acid residues [25]. The most common tripeptide unit of collagen is Gly-Pro-Hyp, and these three residues form one helical turn. As a structural protein, collagen has excellent biocompatibility and cell adhesion and can promote cell proliferation and differentiation.
Elastin
Consisting mainly of glycine, valine, alanine, and proline residues and possessing a molecular weight of ~66 kDa, elastin is present in connective, vascular, and load-bearing tissues and has very elastic mechanical properties. In rat skin, elastin plays a mechanical role at small stress with small deformations [26]. The elastic modulus, strength, extensibility, toughness, and resilience are 0.0011 GPa, 0.002 GPa, 150%, 1.6 MJm−3 and 90%, respectively [27]. Therefore, elastin is considered an important structural protein and material for scaffolds that require elastic physical properties in cell culture. However, elastin is poorly soluble in aqueous solutions, making it difficult to process, and this is a common problem among structural proteins. In addition, since elastin is difficult to obtain, synthesize, and produce in large quantities, reports on biomaterials composed of elastin alone are very limited compared with reports on scaffolds composed of other structural proteins. A scaffold of elastin derived from bovine ligament has been reported, and its mechanical properties have been studied, with a focus on its elastic modulus [28]. The elasticity of elastin makes it an effective additive for improving collagen-based tissue engineering vessels, showing that elastin hybrid composites offer unique mechanical properties such as significant elasticity, indicating that elastin is a useful structural protein for preparing elastic structural materials [29].
Resilin
Resilin, a structural protein, is an entropy elastic material (rubber) found in biological structures that require energy storage and long-range elasticity, and it exhibits 300–400% elongation at break, low solubility, and thermal stability up to ~140 °C [30]. Resilience (the ratio of the energy elastically returned to the added energy) of resilin is ~92% due to covalent crosslinking between tyrosine residues [31]. The mechanical properties of elastin are similar to those of resilin, but resilin shows higher resilience and lower modulus. Resilin can be found in various biological joints, such as the vein joints of dragonfly wings [32]; however, natural resilin is not widely available for biological and physical characterization (Fig. 5). Therefore, by partially cloning and expressing the Drosophila gene that has been identified to encode resilin, a recombinant protein for resilin was successfully designed and obtained, and a rubber-like material could be prepared by rapid photochemical crosslinking [33]. This crosslinking was achieved by peroxidase, which catalyzes dityrosine formation. The recombinant resilin proteins were expressed and evaluated as biomaterials [34,35,36]. Due to the attractive physical characteristics of resilin, combinations of resilin with other proteins have also been investigated [37, 38]. Elastomers composed of biomolecules, including resilin, are often photochemically or chemically crosslinked. These biomaterials behave as rubber-like materials and exhibit high elasticity with small strains, and they are expected to be applicable as new muscle-mimetic biomaterials. On the other hand, examples of research on resilin are limited compared to the number of studies on elastin, as many researchers have studied the use of the latter as a biologically derived elastomer.
Distribution of resilin in the vein joints of the Rhinocypha fenestrella wing. a Resilin present on the dorsal side of the wing. The purple dots indicate resilin. b Fluorescence microscopy (left panels) and scanning electron microscopy images (right panels) of the selected joints. Adopted from ref. [32]
Keratin
Keratin is the main structural protein that forms the hair, wool, feathers, nails, and horns of many types of animals. This protein has a high content of cysteine (7–20% of the total amino acid residues), which is known to form intramolecular and intermolecular disulfide bonds [39, 40]. It has been reported that α-keratin, which has a helical structure, can be reduced to form β-keratin upon stretching treatment, and this structural change affects the mechanical, thermal, and chemical properties of the material [41]. The disulfide bonds in a tissue containing keratin, such as wool, can be disrupted by reductive treatment, affording highly soluble keratin. Keratin can be extracted with an aqueous solution containing urea, mercaptanol, and sodium dodecyl sulfate as a surfactant, and then, insoluble films and biodegradable sponge scaffolds can be prepared [42,43,44].
Reflectin
Reflectin has been found in certain cephalopods (especially squid), including Euprymna scolopes and Doryteuthis opalescens [45]. Squid can change their body color to camouflage themselves, that is, mimic their surroundings, through the function of reflectin, but research on reflectin protein is very limited, as is our understanding of the molecular mechanism. Reflectin is speculated to have random coil and beta barrels as the main motifs [46], but the structure has not been fully elucidated. Protein families that exist in cephalopod pigment cells (rainbow cells, a type of pigment cell that changes the wavelength of reflected light) and white pigment vesicles and contribute to camouflage function have been recently reported. The amino acid sequence of reflectin is rich in aromatic and sulfur-containing amino acids, and these motifs are used to refract incident light in certain environments in certain cephalopods [47, 48]. However, not all the molecular mechanisms have been fully clarified. Aliivibrio fischeri, a marine luminescent bacterium, possesses the reflectin gene of cephalopods due to horizontal gene transmission [49]. Recently, several material scientists have been interested in reflectin as a new functional biopolymer and have reported novel optical biomaterials containing recombinant reflectin [50].
Synthesis and production
The synthesis methods suitable for preparing structural proteins are dependent on many factors, such as their amino acid sequence, solubility in a given solvent, and toxicity to the host cells used for production. In this section, the current synthesis methods are outlined by taking spider silk protein as an example, as it has been studied in a relatively wide range of host organisms. Spiders are carnivorous and cannibalistic, and hence, it is extremely difficult to raise and culture them. Thus, it is enormously challenging to obtain natural spider silk from spiders in bulk. Therefore, in attempts to artificially synthesize spider silk, various organisms, such as plants, goats, Escherichia coli, yeast, and photosynthetic bacteria as well as animals that construct cocoons, have been used as hosts to synthesize silk protein, the main component of spider fibers. Here, the production and synthesis methods are divided into biological synthesis and chemical synthesis methods.
Biological synthesis
Escherichia coli
E. coli is the most convenient and widely used protein synthesis/production host. In research institutions, such as universities, many researchers use this organism in the laboratory. Accordingly, purification methods have been established at the laboratory level (small scale). The synthesis of recombinant proteins such as recombinant silk proteins can be broadly defined in two major steps: (1) design, construction, and cloning of the genes and (2) expression and purification of the protein polymers. E. coli strains such as BL21, JM109, SG13009 pREP4, and RY-3041 have been used as host systems, and protein yields were inversely correlated with the size of the synthetic genes. Synthetic genes encoding dragline silk from Trichonephila clavipes (formally Nephila clavipes) have been successfully constructed, cloned, and expressed [51]. The synthesis of these recombinant genes is based on the repetitive sequences found in native dragline silk genes. The methods for construction of these repeats were previously reported, using smaller oligonucleotide repeats and subsequent multimerization by traditional methods with restriction enzymes and ligase. Thus, the size and sequence of the protein generated can be controlled by the primary sequence synthesized as oligonucleotides. The use of synthetic gene technology to control silk protein size allows for the study of relationships between sequence length and structure-function, along with the study of novel compositions. In general, expression levels obtained from synthetic protein genes are low, with yields of purified protein from 1 to 10 mg/L, usually representing less than 5% of the total protein in the cell, depending upon the size of the protein of interest [52,53,54,55]. By optimizing the sequence and culture conditions, the productivity of recombinant proteins can be drastically improved.
Yeast
A wide variety of yeasts including Saccharomyces cerevisiae have been widely used in the fermentation industry, and many companies prefer to use these microbes in fermentation production. Spider silk production in yeast has a long history; for example, recombinant spider silk protein was successfully expressed in the methylotrophic yeast Pichia pastoris [56, 57]. P. pastoris was reported to exhibit relatively high yield and productivity, with 300–1000 mg/L recombinant spider silk proteins.
Transgenic plants and animals
Tobacco and potato have been used as host plants to produce spider silk proteins [58, 59]. However, the synthesis of the full-length spider silk protein, namely, native‐sized spidroins, from plants remains challenging [60]. In addition to plants, transgenic goats have been designed to synthesize recombinant spider silk proteins in their milk. Furthermore, the silkworm B. mori was genetically modified to synthesize recombinant spider silk proteins [61].
Photosynthetic bacteria
As a “sustainable cell factory”, photosynthetic bacteria are attractive biological hosts to capture and fix CO2 and N2 (Fig. 6) [62]. The author has been interested in purple photosynthetic bacteria because of their biopolymer productivity and their ability to transform technological platforms [63,64,65,66,67,68]. Silk proteins were successfully synthesized by using recombinant nonsulfur photosynthetic bacteria in the marine environment. Importantly, through photosynthesis and nitrogen fixation processes, structural proteins can be synthesized under marine conditions by utilizing low-cost and abundant renewable resources such as light (energy), CO2 (carbon source), and N2 (nitrogen source).
Chemical synthesis
Solid-phase and liquid-phase methods
Solid-phase synthesis is one of the most common techniques used to synthesize relatively short polypeptides (peptides of up to ~50 amino acids) [69, 70]. From a solid surface modified with an amino group, the amino acid chain is extended one residue at a time by dehydration condensation to obtain the target peptide/polypeptide, and then, the solid surface is cleaved to obtain the target peptide/polypeptide. In the condensation reaction, the amino acid side chain must be protected so that it does not react. In general, the method is not scalable, and it is considered unsuitable for synthesizing proteins and polypeptides in bulk. Reactions in a solvent that do not use a solid surface are called liquid-phase reactions, and the basic mechanism is similar to that of solid-phase synthesis [71]. The liquid-phase method is relatively suitable for large-scale synthesis.
Native chemical ligation
In the solid-phase and liquid-phase methods described above, the synthesis efficiency decreases as the molecular weight of the target increases. To compensate for this technical challenge, native chemical ligation is considered a strong complementary reaction [72, 73]. With native chemical ligation, a linked polypeptide can be obtained by mixing a polypeptide with a thioester at the C terminus and a polypeptide with a cysteine at the N terminus, and these precursors can be synthesized by solid-phase or liquid-phase methods. Thus, polypeptides and proteins with more than 200 residues can be chemically synthesized [74]. In recent years, systems other than those based on cysteine have been reported, and their versatility is increasing [75].
α-Amino acid-N-carboxylic anhydride (NCA) ring-opening polymerization
NCA has high electrophilicity and can efficiently afford polypeptides by living anion ring-opening polymerizations using an amine as an initiator [76]. Various polypeptides can be synthesized using an appropriate initiator amine [77]. Currently, the NCA method is not used industrially because a highly toxic reagent such as phosgene or triphosgene is necessary for the preparation of NCA. In recent years, phosgene-free syntheses of NCA have been reported and used to synthesize polypeptides [78, 79]. However, sequence control is still technically difficult, and protein sequences are not easy to mimic.
Chemoenzymatic polymerization
Chemoenzymatic polymerization allows the synthesis of peptides in a relatively short time and with high yield and atom economy [80, 81]. In chemoenzymatic polymerizations for peptide synthesis, amino acid derivatives such as amino acid esters are polymerized via protease-catalyzed aminolysis (Fig. 7) [82, 83]. In addition to natural amino acids [84,85,86], unnatural amino acids, branched structures, and blocky sequences can be easily introduced [87,88,89,90]. By copolymerizing 2-aminoisobutyric acid and nylon units with peptides, the secondary structure and thermal properties of the resulting material can be dramatically altered [90,91,92]. The disadvantage of this method is that it is difficult to control the molecular weight and sequence of the polymer. At present, the application of chemoenzymatic polymerization to the synthesis of polypeptides is still in development.
Schematic of the chemoenzymatic polymerization of alanine ester mediated by papain (a) and a tripeptide (b). c Reaction scheme of chemoenzymatic peptide synthesis mediated by papain. In the first step, acylation, the acyl intermediate is formed upon release of a leaving group from the substrate. The acyl intermediate can undergo either aminolysis or hydrolysis depending on the nucleophile present in the reaction. Adopted from ref. [101]
Synthetic protein material products in the industry
Protein-based materials have been well reviewed by several groups [93,94,95]. Generally, the mechanical/physical properties of structural proteins in nature are expected in their regenerated materials (Table 1). However, it is not easy to reproduce the native physical properties and biological functions of a structural protein using regenerated or recombinant proteins, even though structural proteins can be synthesized by various methods. The reason is that the hierarchical structure and natural biological arrangement are difficult to reproduce by artificial processes. The material processing method needs to be considered and optimized to achieve the desired physical/biological properties of the structural protein material. Here, several examples of structural proteins that have already been commercialized or produced in bulk as a material as well as materials containing a structural protein are listed.
Bolt Threads Microsilk™
Although not disclosed in detail, Bolt Threads (CA, USA) has developed artificial silk fiber, Microsilk™, by a recombinant protein based on the major ampullate spidroin (Fig. 8). The Bolt Threads Microsilk™ Tie is composed of 100% Microsilk™. In 2017, 50 ties were sold at $314 each, and they sold out immediately. Best Made Co. x Bolt Threads Cap of Courage was released in 2017 with 100 colors and 10 patterns at $198 each, and they sold out immediately as well. These hats are a composite of Microsilk™ and American Rambouillet wool. The Stella McCartney x Bolt Threads Gold Microsilk™ Dress is made of 100% Microsilk™ and was designed by Stella McCartney. It was presented and exhibited at “Items: Is Fashion Modern?” at the Museum of Contemporary Art in New York.
Spiber Inc. QMONOS™ and Brewed Protein™
Based on spider silk-derived proteins, Spiber Inc. developed artificial silk fibers with various molecular designs, which can be used for the production of apparel and materials (Fig. 9). Moon Parka™ and T-shirts made of structural protein fibers and textiles were commercialized in 2019. The company also demonstrated silk resin, film, and other silk materials, although these materials have not yet been commercialized as of April 2020 (Fig. 9).
Casein plastic
Casein is a phosphorylated protein present in milk and cheese and is not generally regarded as a structural protein [96]. The colloidal casein in milk solids is composed of not a single type of protein but a combination of α-casein, β-casein, and κ-casein. Focusing on the amino acid composition of casein, most of the serine residues are phosphorylated, and this is regarded as a characteristic structural feature of casein. However, in recent years, biomaterials using casein as structural and/or bulk materials have been developed and produced commercially. Casein plastic was invented in Germany in 1898. Acid is added to milk to solidify casein into a material. The precipitated casein becomes a thermoplastic with an appearance similar to that of ivory. This material is called casein plastic or lactic casein and is used industrially for impressions, clothing buttons, etc. It is not actually a thermoplastic, but it is recognized as a biodegradable, biologically derived material.
Promix
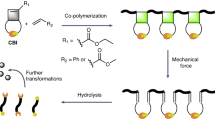
Promix, a fiber made by graft polymerization of acrylonitrile on casein (Fig. 10), was developed by Toyobo Co., Ltd, in the 1970s. The texture is similar to that of silk. Materials that combine Promix and bioplastics such as polylactic acid are also commercially available.
Plant protein fibers
Fibers can be prepared from proteins extracted from plants that have a high protein content, such as soybeans (Fig. 10). Textiles have been developed from peanuts, corn and soybeans. These proteins are difficult to fibrillate as a single substance and are commercialized by compounding with polyester or nylon. For example, a composite fiber was made of 20% soybean protein, 20% nylon, and 60% polyester.
Detection of structural proteins in polymeric materials
There are various natural and synthetic structural proteins and structural protein-like polymers, as introduced above. The content of structural proteins is always important when considering the biomass origin or biodegradability of a material. Furthermore, structural protein quantification is necessary for determining the content of structural proteins in blends and/or composite materials for commercialization. The methods for quantifying structural proteins are the same as those typically used for protein quantification; however, a suitable method must be identified based on the structure, shape, morphology, and size of the material to be measured.
Quantification and qualification
When the whole material is composed of proteins, there is no need to quantify the protein content. However, when in composite materials with other materials, such as polymer resin, the protein content in the material must be determined. If the structural protein is separable from other resins in a composite/blend material, the structural protein can be quantified by the weight ratio (Fig. 11); i.e., the separated structural protein parts are quantified by weight. If the structural protein parts are inseparable, the structural protein needs to be extracted for the content to be quantified.
To extract the protein from a composite/blend material, a pretreatment is necessary prior to extraction. The material is first finely pulverized and subsequently treated with a reducing agent in a denaturing buffer. Typical examples of reducing agents are 10 mM dithiothreitol (DTT), mercaptoethanol, and tris(2-carboxyethyl)phosphine, whereas examples of denaturing buffer are 6 M guanidine hydrochloride, 8 M urea, and formic acid. However, each reducing agent and denaturing buffer combination must be considered with caution because dangerous reactions can occur such as that between urea and chlorine bleach.
To quantify a protein, namely, to determine the concentration of protein in a solution, the UV method, Biuret method, Lowry method, BCA method, Bradford method, etc., can be used. The UV method involves detecting the absorption at ~280 nm, which is the major absorption of tyrosine and tryptophan, but this strategy is not suitable for the quantification of structural proteins because it depends on the tyrosine and tryptophan content of the protein. The Biuret method is based on the detection of Cu+, which is produced when Cu2+ reacts with a protein under alkaline conditions, is an appropriate technique, as are the Lowry method and BCA method, which are advanced variations of the Biuret method. However, the Lowry method is easily affected by a reducing agent such as DTT and is not suitable for this purpose considering the extraction conditions for structural proteins. On the other hand, the BCA method is inhibited in the presence of ~1 mM DTT, ~4 M guanidine hydrochloride, and ~3 M urea but can be used for quantification if the structural protein solution is sufficiently dilute. The Bradford method is conducted under acidic conditions, while the BCA method uses alkaline conditions. Many structural proteins have lower solubility under acidic conditions, making BCA quantification under alkaline conditions the more suitable method. When using the BCA method, both the denaturation buffer and the protein extract must be sufficiently diluted prior to use.
Confirmation of the presence of protein
To confirm that the test material contains some amount of protein, the amino acids, and amide bonds of proteins must be detected by the ninhydrin method [97,98,99]. In the ninhydrin method, the sample is dissolved in a protein denaturing solution, and the resulting solution is used for amino acid analysis. For amino acid composition analysis, a protein sample is hydrolyzed with hydrochloric acid, and each amino acid is quantified by HPLC. Using this strategy, 17 types of amino acids can be confirmed because tryptophan, asparagine, and glutamine cannot be quantified individually. The contents of asparagine and glutamine are combined with those of aspartic acid and glutamic acid, respectively, due to the hydrolysis of asparagine and glutamine during the sample preparation process. Tryptophan is also degraded during preparation, and hence, other protocols for determining tryptophan are needed. When posttranslationally modified peptides and amino acids, such as Hyp, DOPA, and dityrosine, in structural proteins need to be quantified, standard substances corresponding to the target amino acids must be added to the mixture [100].
Confirmation of structural proteins among various proteins
Pure structural protein can be quantified by the general protein quantification method, as described above. However, if structural proteins and other proteins are mixed, they cannot be distinguished by the abovementioned method. To not only quantify a protein but also confirm that it is a structural protein, after treating the extracted protein with a degrading enzyme such as trypsin or separating the protein by sodium dodecyl sulfate–polyacrylamide gel electrophoresis and extracting the protein from the gel, matrix-assisted laser desorption/ionization time-of-flight mass spectrometry should be performed, and databases can be searched for possible sequences. This method requires the protein to have a known sequence, and it is difficult to quantify a novel structural protein prepared by chemical synthesis or genetic recombination technology using this strategy. In addition, shotgun proteomics as well as targeted proteomics using selected reaction monitoring and data-independent acquisition methods are alternative methods. However, modification with synthetic polymers will significantly prevent those characterizations. Thus, in protein quantification, it is generally difficult to specifically quantify proteins with unknown structures. Furthermore, impurity (contaminant) proteins contained in the protein materials synthesized by microbial fermentation must be considered in these analyses.
Summary and future perspective
In this focus review, we focused on structural proteins and broadly outlined them in terms of their definitions, properties, and syntheses as well as their potential as materials and quantification techniques. As mentioned at the beginning of this review, although structural proteins such as silk and elastin are used as structural materials in nature, they have not been put into practical use as bulk-scale structural materials for human use. The challenges of using proteins and polypeptides as structural materials are multifaceted and include their material design, synthesis, purification, and processing. In the future, the author hopes that more scientists will enter this field and establish innovative techniques for molecular design, synthesis, material design, and processing to achieve structural proteins with optimal physical and biological properties.
References
Askarieh G, Hedhammar M, Nordling K, Saenz A, Casals C, Rising A. et al. Self-assembly of spider silk proteins is controlled by a pH-sensitive relay. Nature. 2010;465:236–8.
Hagn F, Eisoldt L, Hardy JG, Vendrely C, Coles M, Scheibel T. et al. A conserved spider silk domain acts as a molecular switch that controls fibre assembly. Nature. 2010;465:239–42.
Tsuchiya K, Numata K. Chemical synthesis of multiblock copolypeptides inspired by spider dragline silk proteins. ACS Macro Lett. 2017;6:103–6.
Babb PL, Lahens NF, Correa-Garhwal SM, Nicholson DN, Kim EJ, Hogenesch JB. et al. The Nephila clavipes genome highlights the diversity of spider silk genes and their complex expression. Nat Genet. 2017;49:895–903.
Vollrath F, Porter D, Holland C. There are many more lessons still to be learned from spider silks. Soft Matter. 2011;7:9595–600.
Katashima T, Malay AD, Numata K. Chemical modification and biosynthesis of silk-like polymers. Curr Opin Chem Eng. 2019;24:61–8.
Gosline JM, Denny MW, DeMont ME. Spider silk as rubber. Nature. 1984;309:551–2.
Gosline JM, Guerette PA, Ortlepp CS, Savage KN. The mechanical design of spider silks: from fibroin sequence to mechanical function. J Exp Biol. 1999;202:3295–303.
Malay AD, Arakawa K, Numata K. Analysis of repetitive amino acid motifs reveals the essential features of spider dragline silk proteins. Plos ONE. 2017;12:e0183397.
Yazawa K, Ishida K, Masunaga H, Hikima T, Numata K. Influence of water content on the beta-sheet formation, thermal stability, water removal, and mechanical properties of silk materials. Biomacromolecules. 2016;17:1057–66.
Yazawa K, Malay AD, Masunaga H, Numata K. Comms Mater. 2020;1:10.
Numata K, Masunaga H, Hikima T, Sasaki S, Sekiyama K, Takata M. Use of extension-deformation-based crystallisation of silk fibres to differentiate their functions in nature. Soft Matter. 2015;11:6335–42.
van Beek JD, Hess S, Vollrath F, Meier BH. The molecular structure of spider dragline silk: folding and orientation of the protein backbone. Proc Natl Acad Sci USA. 2002;99:10266–71.
Xu M, Lewis RV. Structure of a protein superfiber: spider dragline silk. Proc Natl Acad Sci USA. 1990;87:7120–4.
Zhang YQ. Applications of natural silk protein sericin in biomaterials. Biotechnol Adv. 2002;20:91–100.
Malay AD, Sato R, Yazawa K, Watanabe H, Ifuku N, Masunaga H. et al. Relationships between physical properties and sequence in silkworm silks. Sci Rep. 2016;6:27573.
Holland C, Numata K, Rnjak-Kovacina J, Seib FP. The biomedical use of silk: past, present, future. Adv Health Mater. 2019;8:e1800465.
Numata K, Kaplan DL. Silk-based delivery systems of bioactive molecules. Adv Drug Deliv Rev. 2010;62:1497–508.
Numata K, Cebe P, Kaplan DL. Mechanism of enzymatic degradation of beta-sheet crystals. Biomaterials. 2010;31:2926–33.
Numata K, Katashima T, Sakai T. State of water, molecular structure, and cytotoxicity of silk hydrogels. Biomacromolecules. 2011;12:2137–44.
Gupta P, Kumar M, Bhardwaj N, Kumar JP, Krishnamurthy CS, Nandi SK. et al. Mimicking form and function of native small diameter vascular conduits using mulberry and non-mulberry patterned silk films. ACS Appl Mater Inter. 2016;8:15874–88.
Numata K, Sato R, Yazawa K, Hikima T, Masunaga H. Crystal structure and physical properties of Antheraea yamamai silk fibers: Long poly(alanine) sequences are partially in the crystalline region. Polymer. 2015;77:87–94.
Mandal BB, Das S, Choudhury K, Kundu SC. Implication of silk film RGD availability and surface roughness on cytoskeletal organization and proliferation of primary rat bone marrow cells. Tissue Eng A. 2010;16:2391–403.
Ageitos JM, Yazawa K, Tateishi A, Tsuchiya K, Numata K. The benzyl ester group of amino acid monomers enhances substrate affinity and broadens the substrate specificity of the enzyme catalyst in chemoenzymatic copolymerization. Biomacromolecules. 2016;17:314–23.
Lee CH, Singla A, Lee Y. Biomedical applications of collagen. Int J Pharm. 2001;221:1–22.
Oxlund H, Manschot J, Viidik A. The role of elastin in the mechanical-properties of skin. J Biomech. 1988;21:213–8.
Aaron BB, Gosline JM. Elastin as a random-network elastomer—a mechanical and optical analysis of single elastin fibers. Biopolymers. 1981;20:1247–60.
Kirkpatrick SJ, Hinds MT, Duncan DD. Acousto-optical characterization of the viscoelastic nature of a nuchal elastin tissue scaffold. Tissue Eng. 2003;9:645–56.
Berglund JD, Nerem RM, Sambanis A. Incorporation of intact elastin scaffolds in tissue-engineered collagen-based vascular grafts. Tissue Eng. 2004;10:1526–35.
Tatham AS, Shewry PR. Comparative structures and properties of elastic proteins. Philos Trans R Soc Lond B. 2002;357:229–34.
Gosline J, Lillie M, Carrington E, Guerette P, Ortlepp C, Savage K. Elastic proteins: biological roles and mechanical properties. Philos Trans R Soc Lond B. 2002;357:121–32.
Mamat N, Yazawa K, Numata K, Norma-Rashid Y. Morphological and mechanical properties of flexible resilin joints on damselfly wings (Rhinocypha spp.). PLoS ONE. 2018;13:e0193147.
Elvin CM, Carr AG, Huson MG, Maxwell JM, Pearson RD, Vuocolo T. et al. Synthesis and properties of crosslinked recombinant pro-resilin. Nature. 2005;437:999–1002.
Qin GK, Lapidot S, Numata K, Hu X, Meirovitch S, Dekel M. et al. Expression, cross-linking, and characterization of recombinant chitin binding resilin. Biomacromolecules. 2009;10:3227–34.
Qin GK, Rivkin A, Lapidot S, Hu X, Preis I, Arinus SB. et al. Recombinant exon-encoded resilins for elastomeric biomaterials. Biomaterials. 2011;32:9231–43.
Qin GK, Hu X, Cebe P, Kaplan DL. Mechanism of resilin elasticity. Nat Commun. 2012;3:1003.
Cao Y, Li H. Polyprotein of GB1 is an ideal artificial elastomeric protein. Nat Mater. 2007;6:109–14.
Lv S, Dudek DM, Cao Y, Balamurali MM, Gosline J, Li H. Designed biomaterials to mimic the mechanical properties of muscles. Nature. 2010;465:69–73.
Dowling LM, Crewther WG, Parry DA. Secondary structure of component 8c-1 of alpha-keratin. Anal Amino Acid Sequence Biochem J. 1986;236:705–12.
Dowling LM, Crewther WG, Inglis AS. The primary structure of component 8c-1, a subunit protein of intermediate filaments in wool keratin. Relationships with proteins from other intermediate filaments. Biochem J. 1986;236:695–703.
Pauling L, Corey RB. Compound helical configurations of polypeptide chains: structure of proteins of the alpha-keratin type. Nature. 1953;171:59–61.
Yamauchi K, Yamauchi A, Kusunoki T, Kohda A, Konishi Y. Preparation of stable aqueous solution of keratins, and physiochemical and biodegradational properties of films. J Biomed Mater Res. 1996;31:439–44.
Katoh K, Shibayama M, Tanabe T, Yamauchi K. Preparation and physicochemical properties of compression-molded keratin films. Biomaterials. 2004;25:2265–72.
Katoh K, Tanabe T, Yamauchi K. Novel approach to fabricate keratin sponge scaffolds with controlled pore size and porosity. Biomaterials. 2004;25:4255–62.
Crookes WJ, Ding LL, Huang QL, Kimbell JR, Horwitz J, McFall-Ngai MJ. Reflectins: the unusual proteins of squid reflective tissues. Science. 2004;303:235–8.
Kramer RM, Crookes-Goodson WJ, Naik RR. The self-organizing properties of squid reflectin protein. Nat Mater. 2007;6:533–8.
DeMartini DG, Izumi M, Weaver AT, Pandolfi E, Morse DE. Structures, organization, and function of reflectin proteins in dynamically tunable reflective cells. J Biol Chem. 2015;290:15238–49.
Naughton KL, Phan L, Leung EM, Kautz R, Lin QY, Van Dyke Y. et al. Self-assembly of the cephalopod protein reflectin. Adv Mater. 2016;28:8405–12.
Guan Z, Cai TT, Liu ZM, Dou YF, Hu XS, Zhang P. et al. Origin of the reflectin gene and hierarchical assembly of its protein. Curr Biol. 2017;27:2833–42.
Qin GK, Dennis PB, Zhang YJ, Hu X, Bressner JE, Sun ZY. et al. Recombinant reflectin-based optical materials. J Polym Sci Pol Phys. 2013;51:254–64.
Prince JT, McGrath KP, DiGirolamo CM, Kaplan DL. Construction, cloning, and expression of synthetic genes encoding spider dragline silk. Biochemistry. 1995;34:10879–85.
Numata K, Hamasaki J, Subramanian B, Kaplan DL. Gene delivery mediated by recombinant silk proteins containing cationic and cell binding motifs. J Control Release. 2010;146:136–43.
Numata K, Kaplan DL. Silk-based gene carriers with cell membrane destabilizing peptides. Biomacromolecules. 2010;11:3189–95.
Numata K, Reagan MR, Goldstein RH, Rosenblatt M, Kaplan DL. Spider silk-based gene carriers for tumor cell-specific delivery. Bioconjug Chem. 2011;22:1605–10.
Numata K, Subramanian B, Currie HA, Kaplan DL. Bioengineered silk protein-based gene delivery systems. Biomaterials. 2009;30:5775–84.
Fahnestock SR, Bedzyk LA. Production of synthetic spider dragline silk protein in Pichia pastoris. Appl Microbiol Biotechnol. 1997;47:33–9.
Jansson R, Lau CH, Ishida T, Ramstrom M, Sandgren M, Hedhammar M. Functionalized silk assembled from a recombinant spider silk fusion protein (Z-4RepCT) produced in the methylotrophic yeast Pichia pastoris. Biotechnol J. 2016;11:687–99.
Yang JJ, Barr LA, Fahnestock SR, Liu ZB. High yield recombinant silk-like protein production in transgenic plants through protein targeting. Transgenic Res. 2005;14:313–24.
Scheller J, Guhrs KH, Grosse F, Conrad U. Production of spider silk proteins in tobacco and potato. Nat Biotechnol. 2001;19:573–7.
Hauptmann V, Weichert N, Rakhimova M, Conrad U. Spider silks from plants—a challenge to create native-sized spidroins. Biotechnol J. 2013;8:1183–92.
Xu J, Dong QL, Yu Y, Niu BL, Ji DF, Li MW. et al. Mass spider silk production through targeted gene replacement in Bombyx mori. Proc Natl Acad Sci USA. 2018;115:8757–62.
Higuchi-Takeuchi M, Numata K. Marine purple photosynthetic bacteria as sustainable microbial production hosts. Front Bioeng Biotech. 2019;7:258.
Higuchi-Takeuchi M, Morisaki K, Numata K. Method for the facile transformation of marine purple photosynthetic bacteria using chemically competent cells. Microbiologyopen. 2020;9:e953.
Higuchi-Takeuchi M, Numata K. Acetate-inducing metabolic states enhance polyhydroxyalkanoate production in marine purple non-sulfur bacteria under aerobic conditions. Front Bioeng Biotech. 2019;7:118.
Foong CP, Higuchi-Takeuchi M, Numata K. Optimal iron concentrations for growth-associated polyhydroxyalkanoate biosynthesis in the marine photosynthetic purple bacterium Rhodovulum sulfidophilum under photoheterotrophic condition. Plos ONE. 2019;14:e0212654.
Higuchi-Takeuchi M, Motoda Y, Kigawa T, Numata K. Class I polyhydroxyalkanoate synthase from the purple photosynthetic bacterium Rhodovulum sulfidophilum predominantly exists as a functional dimer in the absence of a substrate. ACS Omega. 2017;2:5071–8.
Higuchi-Takeuchi M, Morisaki K, Toyooka K, Numata K. Synthesis of high-molecular-weight polyhydroxyalkanoates by marine photosynthetic purple bacteria. Plos ONE. 2016;11:e0160981.
Higuchi-Takeuchi M, Morisaki K, Numata K. A screening method for the isolation of polyhydroxyalkanoate-producing purple non-sulfur photosynthetic bacteria from natural seawater. Front Microbiol. 2016;7:1509.
Fields GB, Noble RL. Solid phase peptide synthesis utilizing 9-fluorenylmethoxycarbonyl amino acids. Int J Pept Protein Res. 1990;35:161–214.
Merrifield RB. Solid-phase peptide synthesis. Adv Enzymol Ramb. 1969;32:221–96.
Bayer E, Mutter M. Liquid-phase synthesis of peptides. Nature. 1972;237:512–3.
Dawson PE, Muir TW, Clarklewis I, Kent SBH. Synthesis of proteins by native chemical ligation. Science. 1994;266:776–9.
Johnson ECB, Kent SBH. Insights into the mechanism and catalysis of the native chemical ligation reaction. J Am Chem Soc. 2006;128:6640–6.
Johnson ECB, Durek T, Kent SBN. Total chemical synthesis, folding, and assay of a small protein on a water-compatible solid support. Angew Chem Int Ed. 2006;45:3283–7.
Agouridas V, El Mahdi O, Diemer V, Cargoet M, Monbaliu JM, Melnyk O. Native chemical ligation and extended methods: mechanisms, catalysis, scope, and limitations. Chem Rev. 2019;119:7328–443.
Goodman M, Hutchison J. Mechanisms of polymerization of N-unsubstituted N-carboxyanhdrides. J Am Chem Soc. 1966;88:3627
Deming TJ. Synthetic polypeptides for biomedical applications. Prog Polym Sci. 2007;32:858–75.
Koga K, Sudo A, Endo T. Revolutionary phosgene-free synthesis of alpha-amino acid N-carboxyanhydrides using diphenyl carbonate based on activation of alpha-amino acids by converting into imidazolium salts. J Polym Sci Poly Chem. 2010;48:4351–5.
Yamada S, Koga K, Sudo A, Goto M, Endo T. Phosgene-free synthesis of polypeptides: useful synthesis for hydrophobic polypeptides through polycondensation of activated urethane derivatives of amino acids. J Polym Sci Poly Chem. 2013;51:3726–31.
Kobayashi S, Makino A. Enzymatic polymer synthesis: an opportunity for green polymer chemistry. Chem Rev. 2009;109:5288–353.
Kobayashi S, Uyama H, Kimura S. Enzymatic polymerization. Chem Rev. 2001;101:3793–818.
Numata K. Poly(amino acid)s/polypeptides as potential functional and structural materials. Polym J. 2015;47:537–45.
Tsuchiya K, Numata K. Chemoenzymatic synthesis of polypeptides for use as functional and structural materials. Macromol Biosci. 2017;17:1700177.
Ma YN, Li ZB, Numata K. Synthetic short peptides for rapid fabrication of monolayer cell sheets. ACS Biomater Sci Eng. 2016;2:697–706.
Ma YA, Sato R, Li ZB, Numata K. Chemoenzymatic synthesis of oligo(L-cysteine) for use as a thermostable bio-based material. Macromol Biosci. 2016;16:151–9.
Baker PJ, Numata K. Chemoenzymatic synthesis of poly(L-alanine) in aqueous environment. Biomacromolecules. 2012;13:947–51.
Tsuchiya K, Kurokawa N, Gimenez-Dejoz J, Gudeangadi PG, Masunaga H, Numata K. Periodic introduction of aromatic units in polypeptides via chemoenzymatic polymerization to yield specific secondary structures with high thermal stability. Polym J. 2019;51:1287–98.
Ageitos JM, Baker PJ, Sugahara M, Numata K. Proteinase K-catalyzed synthesis of linear and star oligo(L-phenylalanine) conjugates. Biomacromolecules. 2013;14:3635–42.
Fagerland J, Finne-Wistrand A, Numata K. Short one-pot chemo-enzymatic synthesis of L-lysine and L-alanine diblock co-oligopeptides. Biomacromolecules. 2014;15:735–43.
Tsuchiya K, Numata K. Chemoenzymatic synthesis of polypeptides containing the unnatural amino acid 2-aminoisobutyric acid. Chem Commun. 2017;53:7318–21.
Yazawa K, Gimenez-Dejoz J, Masunaga H, Hikima T, Numata K. Chemoenzymatic synthesis of a peptide containing nylon monomer units for thermally processable peptide material application. Polym Chem. 2017;8:4172–6.
Yazawa K, Numata K. Papain-catalyzed synthesis of polyglutamate containing a nylon monomer unit. Polymers. 2016;8:194.
Hu X, Cebe P, Weiss AS, Omenetto F, Kaplan DL. Protein-based composite materials. Mater Today. 2012;15:208–15.
Abascal NC, Regan L. The past, present and future of protein-based materials. Open Biol. 2018;8:180113.
van Hest JC, Tirrell DA. Protein-based materials, toward a new level of structural control. Chem Commun. 2001;19:1897–904.
Farrell HM, Malin EL, Brown EM, Qi PX. Casein micelle structure: What can be learned from milk synthesis and structural biology? Curr Opin Colloid. Interface. 2006;11:135–47.
Starcher B. A ninhydrin-based assay to quantitate the total protein content of tissue samples. Anal Biochem. 2001;292:125–9.
Brewer JM, Roberts CW, Stimson WH, Alexander J. Accurate determination of adjuvant-associated protein or peptide by ninhydrin assay. Vaccine. 1995;13:1441–4.
Doi E, Shibata D, Matoba T. Modified colorimetric ninhydrin methods for peptidase assay. Anal Biochem. 1981;118:173–84.
Numata K, Baker PJ. Synthesis of adhesive peptides similar to those found in blue mussel (Mytilus edulis) using papain and tyrosinase. Biomacromolecules. 2014;15:3206–12.
Gimenez-Dejoz J, Tsuchiya K, Numata K. Insights into the stereospecificity in papain-mediated chemoenzymatic polymerization from quantum mechanics/molecular mechanics simulations. ACS Chem Biol. 2019;14:1280–92.
Acknowledgements
The author thanks Dr Lindsay Wray, Bolt Threads, for providing the photographs in shown Fig. 7 and appreciates Hiroyuki Nakamura, Spiber Inc., for providing the photographs in presented Fig. 8. This work was financially supported by the RIKEN Engineering Network Project, Impulsing Paradigm Change through the Disruptive Technologies Program (ImPACT) and JST ERATO (grant number JPMJER1602).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The author declares that he has no conflict of interest.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Numata, K. How to define and study structural proteins as biopolymer materials. Polym J 52, 1043–1056 (2020). https://doi.org/10.1038/s41428-020-0362-5
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41428-020-0362-5
This article is cited by
-
Cross-linking polybutadiene rubber via a thiol-ene reaction with polycysteine as a degradable cross-linker
Polymer Journal (2024)
-
Comparative proteomics of the shell matrix proteins of Nautilus pompilius and the conchiferans provide insights into mollusk shell evolution at the molecular level
Marine Biology (2023)
-
Development of chiral functional materials based on natural chiral compounds
Polymer Journal (2022)
-
Phosphorylase-catalyzed synthesis and self-assembled structures of cellulose oligomers in the presence of protein denaturants
Polymer Journal (2022)
-
Identification of the fibroin of Stigmaeopsis nanjingensis by a nanocarrier-based transdermal dsRNA delivery system
Experimental and Applied Acarology (2022)