Abstract
Background
Fertility is the most important economic trait in sows, as it is critical for profitability. Considerable phenotypic variation in litter size exists in Large White sows. However, relatively little is known about the underlying molecular and genetic bases.
Objective
An experiment was conducted to screen key genes that affect the fecundity of pigs during the luteal (L) and follicular phases (F) of the estrous cycle.
Methods
Eight sows (n = 4 for high fertility sows and n = 4 for low fertility sows) were sacrificed on day 14 (day 1 = first day of estrus) after estrus in the L phase. Another eight sows were slaughtered on day 20 of the estrous cycle in the F phase. Sixteen ovarian tissue samples were collected at the different sacrifice time points. Total RNA extracted was used to construct the library and then sequence on an Illumina HiSeq X10 system. Differentially expressed genes (DEGs) between high and low fertility in Large White sows were identified, and their potential biological functions were analyzed using bioinformatics analysis.
Results
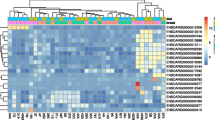
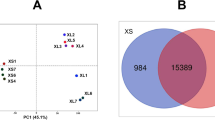
In total, 457 DEGs (161 up-regulated and 296 down-regulated genes) were detected in the ovarian tissues of the high and low fertility groups in the L phase of the estrous cycle. Furthermore, 475 DEGs (253 up-regulated and 222 down-regulated genes) were identified in the F phase. Twenty-nine DEGs were common to both comparisons. Kyoto Encyclopedia of Genes and Genomes enrichment analysis showed that the DEGs were mainly associated with steroid biosynthesis, the Hippo signaling pathway, and lysosomes. Others, such as MSMO1, CYP27B1, and CTSB, were related to reproduction.
Conclusion
These results will contribute to a better understanding of the individual differences in fertility at the transcriptome level, which may provide useful information to explore new ways to improve fertility in pigs.




Similar content being viewed by others
References
Aits S, Jäättelä M (2013) Lysosomal cell death at a glance. J Cell Sci 126:1905–1912. https://doi.org/10.1242/jcs.091181
Audic S, Claverie JM (1997) The significance of digital gene expression profiles. Genome Res 7:986–995. https://doi.org/10.1101/gr.7.10.986
Benjamini Y, Yekutieli D (2001) The control of the false discovery rate in multiple testing under dependency. Ann Stat 29:1165–1188. https://doi.org/10.1214/aos/1013699998
Buaas FW, Lee K, Edelhoff S, Disteche C, Braun RE (1999) Cloning and characterization of the mouse interleukin enhancer binding factor 3 (Ilf3) homolog in a screen for RNA binding proteins. Mamm Genome 10:451–456. https://doi.org/10.1007/s003359901022
Dicken CL, Israel DD, Davis JB, Sun Y, Shu J, Hardin J, Neal-Perry G (2012) Peripubertal vitamin D(3) deficiency delays puberty and disrupts the estrous cycle in adult female mice. Biol Reprod 87:51. https://doi.org/10.1095/biolreprod.111.096511
Dong J, Albertini DF, Kumar Nishimori K, Rajendra Kumar T, Lu N, Matzuk MM (1996) Growth differentiation factor-9 is required during early ovarian folliculogenesis. Nature 383:531–535. https://doi.org/10.1038/383531a0
Feng X, Li F, Wang F, Zhang G, Pang J, Ren C, Zhang T, Yang H, Wang Z, Zhang Y (2018) Genome-wide differential expression profiling of mRNAs and lncRNAs associated with prolificacy in Hu sheep. Biosci Rep. https://doi.org/10.1042/BSR20171350
Groh KJ, Nesatyy VJ, Segner H, Eggen RI, Suter MJ (2011) Global proteomics analysis of testis and ovary in adult zebrafish (Danio rerio). Fish Physiol Biochem 37:619–647. https://doi.org/10.1007/s10695-010-9464-x
Groh KJ, Schönenberger R, Eggen RI, Segner H, Suter MJ (2013) Analysis of protein expression in zebrafish during gonad differentiation by targeted proteomics. Gen Comp Endocrinol 193:210–220. https://doi.org/10.1016/j.ygcen.2013.07.020
Halloran BP, DeLuca HF (1980) Effect of vitamin D deficiency on fertility and reproductive capacity in the female rat. J Nutr 110:1573–1580. https://doi.org/10.1093/jn/110.8.1573
Hsueh AJ, Kawamura K, Cheng Y, Fauser BC (2015) Intraovarian control of early folliculogenesis. Endocr Rev 36:1–24. https://doi.org/10.1210/er.2014-1020
Jang H, Choi Y, Yoo I, Han J, Hong JS, Kim YY, Ka H (2017) Vitamin D-metabolic enzymes and related molecules: expression at the maternal-conceptus interface and the role of vitamin D in endometrial gene expression in pigs. PLoS One 12:e0187221. https://doi.org/10.1371/journal.pone.0187221
Kanehisa M, Goto S, Kawashima S, Nakaya A et al (2000) KEGG: Kyoto Encyclopaedia of Genes and Genomes. Nucleic Acids Res 28:27–30. https://doi.org/10.1093/nar/28.1.27
Kanehisa M, Araki M, Goto S, Hattori M, Hirakawa M, Itoh M, Katayama T, Kawashima S, Okuda S, Tokimatsu T, Yamanishi Y (2008) KEGG for linking genomes to life and the environment. Nucleic Acids Res 36:D480–D484. https://doi.org/10.1093/nar/gkm882
Knapczyk-Stwora K, Grzesiak M, Witek P, Duda M, Koziorowski M, Slomczynska M (2019) Neonatal exposure to agonists and antagonists of sex steroid receptors induces changes in the expression of oocyte-derived growth factors and their receptors in ovarian follicles in gilts. Theriogenology 134:42–52. https://doi.org/10.1016/j.theriogenology.2019.05.018
Langmead B, Trapnell C, Pop M, Salzberg SL (2009) Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome Biol 10:R25. https://doi.org/10.1186/gb-2009-10-3-r25
Li EL, Xie XH, Xu YF, Xie Z, Chen L, Liu HL, Li QF (2010) Relationship between the mRNA expression level of TGF-β receptor genes in tissues and ovulation rate in Hu sheep. Agric Sci China 9:1659–1666. https://doi.org/10.1016/S1671-2927(09)60263-7
Li J, Luo W, Huang T, Gong Y (2019) Growth differentiation factor 9 promotes follicle-stimulating hormone-induced progesterone production in chicken follicular granulosa cells. Gen Comp Endocrinol 276:69–76. https://doi.org/10.1016/j.ygcen.2019.03.005
Ling YH, Xiang H, Li YS, Liu Y, Zhang YH, Zhang ZJ, Ding JP, Zhang XR (2014) Exploring differentially expressed genes in the ovaries of uniparous and multiparous goats using the RNA-Seq (quantification) method. Gene 550:148–153. https://doi.org/10.1016/j.gene.2014.08.008
Ling YH, Guo XF, Chen T, Ding JP, Ma YH, Chu MX, Di R, Zhang YH, Zhang XR (2016) Characterization and analysis of differentially expressed microRNAs in hircine ovaries during the follicular and luteal phases. Anim Reprod Sci 166:47–57. https://doi.org/10.1016/j.anireprosci.2016.01.003
Linville RC, Pomp D, Johnson RK, Rothschild MF (2001) Candidate gene analysis for loci affecting litter size and ovulation rate in swine. J Anim Sci 79:60–67. https://doi.org/10.2527/2001.79160x
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2(−Delta Delta C (T)) Method. Methods 25:402–408. https://doi.org/10.1006/meth.2001.1262
Lou Q, Li T, Wu P, Qiu C, Zhang G, Wang J (2019) Polymorphism identification in GDF9 gene and its association analysis with reproduction traits in Jinghai Yellow chicken. Anim Biotechnol 30:332–341. https://doi.org/10.1080/10495398.2018.1516222
McBride D, Carré W, Sontakke SD, Hogg CO, Law A, Donadeu FX, Clinton M (2012) Identification of miRNAs associated with the follicular–luteal transition in the ruminant ovary. Reproduction 144:221–233. https://doi.org/10.1530/REP-12-0025
Miao XY, Luo QM, Qin XY (2016) Genome-wide transcriptome analysis in the ovaries of two goats identifies differentially expressed genes related to fecundity. Gene 582:69–76. https://doi.org/10.1016/j.gene.2016.01.047
Mortazavi A, Williams BA, McCue K, Schaeffer L, Wold B (2008) Mapping and quantifying mammalian transcriptomes by RNA-Seq. Nat Methods 5:621–628. https://doi.org/10.1038/nmeth.1226
Ollivier L, Messer LA, Rothschild MF, Legault C (1997) The use of selection experiments for detecting quantitative trait loci. Genet Res 69:227–232. https://doi.org/10.1017/s0016672397002802
Panda DK, Miao D, Tremblay ML, Sirois J, Farookhi R, Hendy GN, Goltzman D (2001) Targeted ablation of the 25-hydroxyvitamin D 1α-hydroxylase enzyme: evidence for skeletal, reproductive, and immune dysfunction. Proc Natl Acad Sci USA 98:7498–7503. https://doi.org/10.1073/pnas.131029498
Roehe R, Kennedy BW (1995) Estimation of genetic parameters for litter size in Canadian Yorkshire and Landrace swine with each parity of farrowing treated as a different trait. J Anim Sci 73:2959–2970. https://doi.org/10.2527/1995.73102959x
Rothschild MF, Messer L, Day A, Wales R, Short T, Southwood O, Plastow G (2000) Investigation of the retinol binding protein (RBP4) gene as a candidate gene for litter size in the pig. Mamm Genome 11:75–77. https://doi.org/10.1007/s003350010015
Rydhmer L (2000) Genetics of sow reproduction, including puberty, oestrus, pregnancy, farrowing and lactation. Livest Prod Sci 66:1–12. https://doi.org/10.1016/s0301-6226(99)00170-0
Short TH, Rothschild MF, Southwood OI, McLaren DG, de Vries A, van der Steen H, Eckardt GR, Tuggle CK, Helm J, Vaske DA, Mileham AJ, Plastow GS (1997) Effect of the estrogen receptor locus on reproduction and production traits in four commercial pig lines. J Anim Sci 75:3138–3142. https://doi.org/10.2527/1997.75123138x
Soede NM, Langendijk P, Kemp B (2011) Reproductive cycles in pigs. Anim Reprod Sci 124:251–258. https://doi.org/10.1016/j.anireprosci.2011.02.025
Spötter A, Distl O (2006) Genetic approaches to the improvement of fertility traits in the pig. Vet J 172:234–247. https://doi.org/10.1016/j.tvjl.2005.11.013
Steinheuer R, Drögemüller C, Hamann H, Distl O (2003) Possible uses of genetic markers for improving fertility and health in swine production. Dtsch Tierarztl Wochenschr 110:255–265
Uimari P, Sironen A, Sevón-Aimonen ML (2011) Whole-genome SNP association analysis of reproduction traits in the Finnish Landrace pig breed. Genet Sel Evol 43:1–8. https://doi.org/10.1186/1297-9686-43-42
Zhang X, Huang L, Wu T, Feng Y, Ding Y, Ye P, Yin Z (2015) Transcriptomic analysis of ovaries from pigs with high and low litter size. PLoS One 10:e0139514. https://doi.org/10.1371/journal.pone.0139514
Zhang FP, Niu X, Tang LT, Ran XQ, Li S, Wang JF, Huang SH (2018) Identification of candidate genes associated with Xiang pig estrus by genome-wide transcriptome analysis. Int Clin Exp Med 11:12079–12087
Zhao M, Antunes F, Eaton JW, Brunk UT (2003) Lysosomal enzymes promote mitochondrial oxidant production, cytochrome c release and apoptosis. Eur J Biochem 270:3778–3786. https://doi.org/10.1046/j.1432-1033.2003.03765.x
Acknowledgements
We thank Hebei Shunde-Tianzhao Livestock Technology Co., Ltd (Wanquan, Hebei, China.) for their assistance in sample collection. We thank Charles worth Author Services (http://www.cwauthors.com) for its linguistic assistance during the preparation of this manuscript. This study was supported by the Modern Agriculture Industry Technology System Foundation of Hebei Province (No. HBCT2018110201).
Author information
Authors and Affiliations
Contributions
Conceptualization: QJ; methodology: QJ, HH, and BZ; formal analysis and investigation: HH, JZ, ZL, and ZL; writing—original draft preparation: HH; writing—review and editing: QJ; supervision: QJ.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical approval
All applicable international, national, and/or institutional guidelines for the care and use of animals were followed.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Hu, H., Jia, Q., Zhou, B. et al. Comparative analysis of the ovarian transcriptome reveals novel insights into fertility differences in Large White sows. Genes Genom 42, 715–725 (2020). https://doi.org/10.1007/s13258-020-00926-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13258-020-00926-8