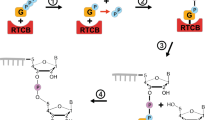
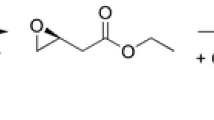
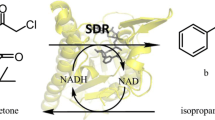
Abstract
Brasiliamides are a class of piperazine-containing alkaloids produced by Penicillium brasilianum with a range of pharmaceutical activities. The mechanism of brasiliamide biosynthesis, including piperazine ring formation and multiple tailoring modifications, still remains unclear. In this study, the biosynthetic gene cluster of brasiliamides, brs, was identified from the marine-derived fungal strain Penicillium brasilianum WZXY-M122-9. Deletion of a histone deacetylase–encoding gene using a CRISPR/Cas9 gene editing system led to the production of a new compound, namely brasiliamide I (1). The brs-encoded single-module nonribosomal peptide synthetase (NRPS) BrsA is involved in the formation of the piperazine skeleton of brasiliamides. Full-length BrsA protein (113.6 kDa) was purified, and reconstitution of enzymatic activity in vitro confirmed that BrsA stereoselectively accepts l-phenylalanine as the substrate. Multiple deletion of tailoring genes and analysis of purified proteins in vitro enabled us to propose a brasiliamide biosynthetic pathway. In the tailoring steps, an α-ketoglutarate (KG)-dependent nonheme iron dioxygenase, BrsJ, was identified to catalyze piperazine ring cleavage during biosynthesis of brasiliamide A (2).
Key Points
-
The gene cluster encoding brasiliamide biosynthesis, brs, is identified.
-
Deletion of a histone deacetylase–encoding gene produces brasiliamide I.
-
BrsA catalyzes brasiliamide piperazine skeleton formation.
-
BrsJ catalyzes piperazine ring cleavage to produce brasiliamide A.

Graphical abstract






Similar content being viewed by others
References
Arai T, Takahashi K, Nakahara S, Kubo A (1980) The structure of a novel antitumor antibiotic, saframycin A. Experientia 36(9):1025–1027. https://doi.org/10.1007/bf01965946
Brito AF, Moreira L, Menegatti R, Costa EA (2019) Piperazine derivatives with central pharmacological activity used as therapeutic tools. Fundam Clin Pharmacol 33(1):13–24. https://doi.org/10.1111/fcp.12408
Chiba T, Asami Y, Suga T, Watanabe Y, Nagai T, Momose F, Nonaka K, Iwatsuki M, Yamada H, Omura S, Shiomi K (2017) Herquline A, produced by Penicillium herquei FKI-7215, exhibits anti-influenza virus properties. Biosci Biotechnol Biochem 81(1):59–62. https://doi.org/10.1080/09168451.2016.1162084
Conti E, Stachelhaus T, Marahiel MA, Brick P (1997) Structural basis for the activation of phenylalanine in the non-ribosomal biosynthesis of gramicidin S. EMBO J 16(14):4174–4183. https://doi.org/10.1093/emboj/16.14.4174
Corey EJ, Gin DY, Kania RS (1996) Enantioselective total synthesis of ecteinascidin 743. J Am Chem Soc 118(38):9202–9203. https://doi.org/10.1021/ja962480t
Cuevas C, Francesch A (2009) Development of Yondelis (trabectedin, ET-743): a semisynthetic process solves the supply problem. Nat Prod Rep 26(3):322–337. https://doi.org/10.1039/b808331m
Enomoto Y, Shiomi K, Hayashi M, Masuma R, Kawakubo T, Tomosawa K, Iwai Y, Omura S (1996) Herquline B, a new platelet aggregation inhibitor produced by Penicillium herquei Fg-372. J Antibiot (Tokyo) 49(1):50–53. https://doi.org/10.7164/antibiotics.49.50
Fill TP, Da SB, Rodrigues-Fo E (2010) Biosynthesis of phenylpropanoid amides by an endophytic Penicillium brasilianum found in root bark of Melia azedarach. J Microbiol Biotechnol 20(3):622–629. https://doi.org/10.4014/jmb.0908.08018
Forseth RR, Amaike S, Schwenk D, Affeldt KJ, Hoffmeister D, Schroeder FC, Keller NP (2013) Homologous NRPS-like gene clusters mediate redundant small-molecule biosynthesis in Aspergillus flavus. Angew Chem Int Ed Engl 52(5):1590–1594. https://doi.org/10.1002/anie.201207456
Fujita T, Hayashi H (2004) New brasiliamide congeners, brasiliamides C, D and E, from Penicillium brasilianum Batista JV-379. Biosci Biotechnol Biochem 68(4):820–826. https://doi.org/10.1271/bbb.68.820
Fujita T, Makishima D, Akiyama K, Hayashi H (2002) New convulsive compounds, brasiliamides A and B, from Penicillium brasilianum Batista JV-379. Biosci Biotechnol Biochem 66(8):1697–1705. https://doi.org/10.1271/bbb.66.1697
Gao X, Haynes SW, Ames BD, Wang P, Vien LP, Walsh CT, Tang Y (2012) Cyclization of fungal nonribosomal peptides by a terminal condensation-like domain. Nat Chem Biol 8(10):823–830. https://doi.org/10.1038/nchembio.1047
Guo XC, Xu LL, Yang RY, Yang MY, Hu LD, Zhu HJ, Cao F (2019) Anti-vibrio indole-diterpenoids and C-25 epimeric steroids from the marine-derived fungus Penicillium janthinellum. Front Chem 7(80):1–7. https://doi.org/10.3389/fchem.2019.00080
Jin S, Gorfajn B, Faircloth G, Scotto KW (2000) Ecteinascidin 743, a transcription-targeted chemotherapeutic that inhibits MDR1 activation. Proc Natl Acad Sci U S A 97(12):6775–6779. https://doi.org/10.1073/pnas.97.12.6775
Li P, Li J, Guo Z, Tang W, Han J, Meng X, Hao T, Zhu Y, Zhang L, Chen Y (2015) An efficient blue-white screening based gene inactivation system for Streptomyces. Appl Microbiol Biotechnol 99(4):1923–1933. https://doi.org/10.1007/s00253-014-6369-0
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2(-ΔΔCt) method. Methods 25(4):402–408. https://doi.org/10.1006/meth.2001.1262
Mao XM, Xu W, Li D, Yin WB, Chooi YH, Li YQ, Tang Y, Hu Y (2015) Epigenetic genome mining of an endophytic fungus leads to the pleiotropic biosynthesis of natural products. Angew Chem Int Ed Engl 54(26):7592–7596. https://doi.org/10.1002/anie.201502452
Omura S, Hirano A, Iwai Y, Masuma R (1979) Herquline, a new alkaloid produced by Penicillium herquei: fermentation, isolation and properties. J Antibiot (Tokyo) 32(8):786–790. https://doi.org/10.7164/antibiotics.32.786
Ono E, Nakai M, Fukui Y, Tomimori N, Fukuchi-Mizutani M, Saito M, Satake H, Tanaka T, Katsuta M, Umezawa T, Tanaka Y (2006) Formation of two methylenedioxy bridges by a Sesamum CYP81Q protein yielding a furofuran lignan, (+)-sesamin. Proc Natl Acad Sci U S A 103(26):10116–10121. https://doi.org/10.1073/pnas.0603865103
Paolinelli-Alfonso M, Galindo-Sanchez CE, Hernandez-Martinez R (2016) Quantitative real-time PCR normalization for gene expression studies in the plant pathogenic fungi Lasiodiplodia theobromae. J Microbiol Methods 127:82–88. https://doi.org/10.1016/j.mimet.2016.05.021
Podobnik B, Stojan J, Lah L, Kraševec N, Seliškar M, Rižner TL, Rozman D, Komel R (2008) CYP53A15 of Cochliobolus lunatus, a target for natural antifungal compounds. J Med Chem 51:3480–3486. https://doi.org/10.1021/jm800030e
Prieto C, Garcia-Estrada C, Lorenzana D, Martin JF (2012) NRPSsp: non-ribosomal peptide synthase substrate predictor. Bioinformatics 28(3):426–427. https://doi.org/10.1093/bioinformatics/btr659
Sahner JH, Empting M, Kamal A, Weidel E, Groh M, Borger C, Hartmann RW (2015) Exploring the chemical space of ureidothiophene-2-carboxylic acids as inhibitors of the quorum sensing enzyme PqsD from Pseudomonas aeruginosa. Eur J Med Chem 96:14–21. https://doi.org/10.1016/j.ejmech.2015.04.007
Shinohara Y, Kawatani M, Futamura Y, Osada H, Koyama Y (2016) An overproduction of astellolides induced by genetic disruption of chromatin-remodeling factors in Aspergillus oryzae. J Antibiot (Tokyo) 69(1):4–8. https://doi.org/10.1038/ja.2015.73
Tamai S, Kaneda M, Nakamura S (1982) Piperazinomycin, a new antifungal antibiotic. I. Fermentation, isolation, characterization and biological properties. J Antibiot (Tokyo) 35(9):1130–1136. https://doi.org/10.7164/antibiotics.35.1130
Tanifuji R, Koketsu K, Takakura M, Asano R, Minami A, Oikawa H, Oguri H (2018) Chemo-enzymatic total syntheses of jorunnamycin a, saframycin A, and N-Fmoc saframycin Y3. J Am Chem Soc 140(34):10705–10709. https://doi.org/10.1021/jacs.8b07161
Weber T, Blin K, Duddela S, Krug D, Kim HU, Bruccoleri R, Lee SY, Fischbach MA, Müller R, Wohlleben W, Breitling R, Takano E, Medema MH (2015) antiSMASH 3.0-a comprehensive resource for the genome mining of biosynthetic gene clusters. Nucleic Acids Res 43:W237–W243. https://doi.org/10.1093/nar/gkv437
Winkler M (2018) Carboxylic acid reductase enzymes (CARs). Curr Opin Chem Biol 43:23–29. https://doi.org/10.1016/j.cbpa.2017.10.006
Wu G, Zhou H, Zhang P, Wang X, Li W, Zhang W, Liu X, Liu HW, Keller NP, An Z, Yin WB (2016) Polyketide production of pestaloficiols and macrodiolide ficiolides revealed by manipulations of epigenetic regulators in an endophytic fungus. Org Lett 18(8):1832–1835. https://doi.org/10.1021/acs.orglett.6b00562
Xu X, Liu L, Zhang F, Wang W, Li J, Guo L, Che Y, Liu G (2014) Identification of the first diphenyl ether gene cluster for pestheic acid biosynthesis in plant endophyte Pestalotiopsis fici. Chembiochem 15(2):284–292. https://doi.org/10.1002/cbic.201300626
Yu X, Liu F, Zou Y, Tang MC, Hang L, Houk KN, Tang Y (2016) Biosynthesis of strained piperazine alkaloids: uncovering the concise pathway of herquline A. J Am Chem Soc 138(41):13529–13532. https://doi.org/10.1021/jacs.6b09464
Zhang X, Liu CJ (2015) Multifaceted regulations of gateway enzyme phenylalanine ammonia-lyase in the biosynthesis of phenylpropanoids. Mol Plant 8(1):17–27. https://doi.org/10.1016/j.molp.2014.11.001
Zhang J, Yuan B, Liu D, Gao S, Proksch P, Lin W (2018) Brasilianoids A-F, new meroterpenoids fom the sponge-associated fungus Penicillium brasilianum. Front Chem 6:314. https://doi.org/10.3389/fchem.2018.00314
Zhang J, Wu Y, Yuan B, Liu D, Zhu K, Huang J, Proksch P, Lin W (2019) DMOA-based meroterpenoids with diverse scaffolds from the sponge-associated fungus Penicillium brasilianum. Tetrahedron 75(14):2193–2205. https://doi.org/10.1016/j.tet.2019.02.037
Acknowledgments
The authors thank Dr. Jinwei Ren for the measurements of NMR spectroscopic data.
Author contribution statement
BY conducted genetic manipulation. DL performed the fungus fermentation. XG and YY analyzed the bioinformatics. JZ helped to record the spectroscopic data. DY and YZ assisted with gene deletion steps. MM and WL conceived and designed the research protocol and wrote the manuscript. All authors read and approved the manuscript.
Funding
This work has been funded by the National Natural Science Foundation of China (81991525, 21861142006, 81872793, 81630089, 81673332, and 81573326), COMRA (DY135-B-05), and MOST (2018ZX09711001-001-008).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
This article does not contain any studies with animals performed by any of the authors.
Conflict of interest
The authors declare that they have no conflict of interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
ESM 1
(PDF 2.78 mb)
Rights and permissions
About this article
Cite this article
Yuan, B., Liu, D., Guan, X. et al. Piperazine ring formation by a single-module NRPS and cleavage by an α-KG-dependent nonheme iron dioxygenase in brasiliamide biosynthesis. Appl Microbiol Biotechnol 104, 6149–6159 (2020). https://doi.org/10.1007/s00253-020-10678-w
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-020-10678-w