Abstract
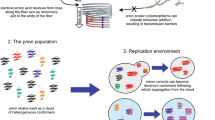
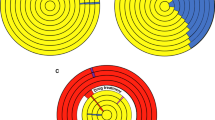
Prions are often considered as anomalous proteins associated primarily with disease rather than as a fundamental source of diversity within biological proteomes. Whereas this longstanding viewpoint has its genesis in the discovery of the original namesake prions as causative agents of several complex diseases, the underlying assumption of a strict disease basis for prions could not be further from the truth. Prions and the spectrum of functions they comprise, likely represent one of the largest paradigm shifts concerning molecular-encoded phenotypic diversity since identification of DNA as the principle molecule of heredity. The ability of prions to recruit similar proteins to alternate conformations may engender a reservoir of diversity supplementing the genetic diversity resulting from stochastic mutations of DNA and subsequent natural selection. Here we present several currently known prions and how many of their functions as well as modes of transmission are intricately linked to adaptation from an evolutionary perspective. Further, the stability of some prion conformations across generations indicates that heritable prion-based adaptation is a reality.
Similar content being viewed by others
References
Aguzzi A (2009) Beyond the prion principle. Nature 459:924–925. https://doi.org/10.1038/459924a
Aguzzi A, Polymenidou M (2004) Mammalian prion biology: one century of evolving concepts. Cell 116:313–327. https://doi.org/10.1016/S0092-8674(03)01031-6
Aguzzi A, Rajendran L (2009) The transcellular spread of cytosolic amyloids, prions, and prionoids. Neuron 64:783–790. https://doi.org/10.1016/j.neuron.2009.12.016
Alberti S, Halfmann R, King O et al (2009) A systematic survey identifies prions and illuminates sequence features of prionogenic proteins. Cell 137:146–158. https://doi.org/10.1016/j.cell.2009.02.044.A
Alper T, Cramp WA, Haig DA, Clarke MC (1967) Does the agent of Scrapie replicate without nucleic acid? Nature 214:764–766. https://doi.org/10.1038/214764a0
An L, Fitzpatrick D, Harrison PM (2016) Emergence and evolution of yeast prion and prion-like proteins. BMC Evol Biol 16:24. https://doi.org/10.1186/s12862-016-0594-3
Brown JCS, Lindquist S (2009) A heritable switch in carbon source utilization driven by an unusual yeast prion. Genes Dev 23:2320–2332. https://doi.org/10.1101/gad.1839109
Brown P, Bradley R (1998) 1755 and all that: a historical primer of transmissible spongiform encephalopathy. BMJ 317:1688–1692. https://doi.org/10.1136/bmj.317.7174.1688
Cassmann ED, Moore SJ, Smith JD, Greenlee JJ (2019) Sheep are susceptible to the bovine adapted transmissible mink encephalopathy agent by intracranial inoculation and have evidence of infectivity in lymphoid tissues. Front Vet Sci. https://doi.org/10.3389/fvets.2019.00430
Chakrabortee S, Kayatekin C, Newby GA et al (2016) Luminidependens (LD) is an Arabidopsis protein with prion behavior. Proc Natl Acad Sci USA 113:6065–6070. https://doi.org/10.1073/pnas.1604478113
Chakravarty AK, Smejkal T, Itakura AK et al (2020) A non-amyloid prion particle that activates a heritable gene expression program. Mol Cell 77(2):251–265. https://doi.org/10.1016/j.molcel.2019.10.028
Chen C-C, Wang Y-H, Wu K-Y (2013) Consumption of bovine spongiform encephalopathy (BSE) contaminated beef and the risk of Variant Creutzfeldt-Jakob Disease. Risk Anal 33:1958–1968. https://doi.org/10.1111/risa.12079
Chernoff YO (2001) Mutation processes at the protein level: is Lamarck back? Mutat Res Mutat Res 488:39–64. https://doi.org/10.1016/S1383-5742(00)00060-0
Chernova TA, Kiktev DA, Romanyuk AV et al (2017) Yeast short-lived actin-associated protein forms a metastable prion in response to thermal stress. Cell Rep 18:751–761. https://doi.org/10.1016/j.celrep.2016.12.082
Collinge J (1999) Variant Creutzfeldt-Jakob disease. Lancet 354:317–323. https://doi.org/10.1016/S0140-6736(99)05128-4
Collinge J (2016) Mammalian prions and their wider relevance in neurodegenerative diseases. Nature 539:217–226. https://doi.org/10.1038/nature20415
Coustou V, Deleu C, Saupe S, Begueret J (1997) The protein product of the het-s heterokaryon incompatibility gene of the fungus Podospora anserina behaves as a prion analog. Proc Natl Acad Sci USA 94:9773–9778. https://doi.org/10.1073/pnas.94.18.9773
Cox BS (1965) Ψ, A cytoplasmic suppressor of super-suppressor in yeast. Heredity (Edinb) 20:505–521. https://doi.org/10.1038/hdy.1965.65
Crow ET, Li L (2011) Newly identified prions in budding yeast, and their possible functions. Semin Cell Dev Biol 22:452–459. https://doi.org/10.1016/j.semcdb.2011.03.003
Cushman M, Johnson BS, King OD et al (2010) Prion-like disorders: blurring the divide between transmissibility and infectivity. J Cell Sci 123:1191–1201. https://doi.org/10.1242/jcs.051672
Dahanukar A, Walker JA, Wharton RP (1999) Smaug, a novel RNA-binding protein that operates a translational switch in Drosophila. Mol Cell 4:209–218. https://doi.org/10.1016/s1097-2765(00)80368-8
Daubin V, Szöllősi GJ (2016) Horizontal gene transfer and the history of life. Cold Spring Harb Perspect Biol 8:a018036. https://doi.org/10.1101/cshperspect.a018036
Derkatch IL, Bradley ME, Hong JY, Liebman SW (2001) Prions affect the appearance of other prions: the story of [PIN(+)]. Cell 106:171–182. https://doi.org/10.1016/S0092-8674(01)00427-5
Du Z, Park KW, Yu H et al (2008) Newly identified prion linked to the chromatin-remodeling factor Swi1 in Saccharomyces cerevisiae. Nat Genet 40:460–465. https://doi.org/10.1038/ng.112
Eraña H (2019) The Prion 2018 round tables (II): Aβ, tau, α-synuclein… are they prions, prion-like proteins, or what? Prion 13:41–45. https://doi.org/10.1080/19336896.2019.1569451
Forloni G, Chiesa R, Bugiani O et al (2019) PrP 106–126—25 years after. Neuropathol Appl Neurobiol. https://doi.org/10.1111/nan.12538
Garcia DM, Jarosz DF (2014) Rebels with a cause: molecular features and physiological consequences of yeast prions. FEMS Yeast Res 14:136–147. https://doi.org/10.1111/1567-1364.12116
Goncharoff DK, Du Z, Li L (2018) A brief overview of the Swi1 prion—[SWI+]. FEMS Yeast Res. https://doi.org/10.1093/femsyr/foy061
Griffith JS (1967) Nature of the scrapie agent: self-replication and scrapie. Nature 215:1043–1044. https://doi.org/10.1038/2151043a0
Halfmann R, Wright JR, Alberti S et al (2012) Prion formation by a yeast GLFG nucleoporin. Prion 6:391–399. https://doi.org/10.4161/pri.20199
Harbi D, Harrison PM (2014) Classifying prion and prion-like phenomena. Prion 8:161–165. https://doi.org/10.4161/pri.27960
Heinrich SU, Lindquist S (2011) Protein-only mechanism induces self-perpetuating changes in the activity of neuronal Aplysia cytoplasmic polyadenylation element binding protein (CPEB). Proc Natl Acad Sci USA 108:2999–3004. https://doi.org/10.1073/pnas.1019368108
Hofmann JP, Denner P, Nussbaum-Krammer C et al (2013) Cell-to-cell propagation of infectious cytosolic protein aggregates. Proc Natl Acad Sci USA 110:5951–5956. https://doi.org/10.1073/pnas.1217321110
Holmes DL, Lancaster AK, Lindquist S, Halfmann R (2013) Heritable remodeling of yeast multicellularity by an environmentally responsive prion. Cell 153:153–165. https://doi.org/10.1016/j.cell.2013.02.026
Itakura AK, Chakravarty AK, Jakobson CM, Jarosz DF (2020) Widespread prion-based control of growth and differentiation strategies in Saccharomyces cerevisiae. Mol Cell 77(2):266–278. https://doi.org/10.1016/j.molcel.2019.10.027
Jarosz DF, Brown JCS, Walker GA et al (2014) Cross-kingdom chemical communication drives a heritable, mutually beneficial prion-based transformation of metabolism. Cell 158:1083–1093. https://doi.org/10.1016/j.cell.2014.07.025
Johnson RT, Gibbs CJ (1998) Creutzfeldt-Jakob Disease and related transmissible spongiform encephalopathies. N Engl J Med 339:1994–2004. https://doi.org/10.1056/NEJM199812313392707
Kabani M, Melki R (2015) Sup35p in its soluble and prion states is packaged inside extracellular vesicles. MBio. https://doi.org/10.1128/mBio.01017-15
Keefer KM, Stein KC, True HL (2017) Heterologous prion-forming proteins interact to cross-seed aggregation in Saccharomyces cerevisiae. Sci Rep 7(1):5853. https://doi.org/10.1038/s41598-017-05829-5
Kelly AC, Shewmaker FP, Kryndushkin D, Wickner RB (2012) Sex, prions, and plasmids in yeast. Proc Natl Acad Sci USA 109:E2683–E2690. https://doi.org/10.1073/pnas.1213449109
Kurt TD, Sigurdson CJ (2016) Cross-species transmission of CWD prions. Prion 10:83–91. https://doi.org/10.1080/19336896.2015.1118603
Lancaster AK, Nutter-Upham A, Lindquist S, King OD (2014) PLAAC: a web and command-line application to identify proteins with prion-like amino acid composition. Bioinformatics 30:2501–2502. https://doi.org/10.1093/bioinformatics/btu310
Liebman SW, Chernoff YO (2012) Prions in yeast. Genetics 191:1041–1072. https://doi.org/10.1534/genetics.111.137760
Light WH, Freaney J, Sood V et al (2013) A conserved role for human Nup98 in altering chromatin structure and promoting epigenetic transcriptional memory. PLoS Biol 11(3):e1001524. https://doi.org/10.1371/journal.pbio.1001524
Liu S, Hossinger A, Hofmann JP et al (2016) Horizontal transmission of cytosolic Sup35 prions by extracellular vesicles. MBio 7:e00915–e00916. https://doi.org/10.1128/mBio.00915-1600
Lupi O, Dadalti P, Cruz E, Sanberg PR (2006) Are prions related to the emergence of early life? Med Hypotheses 67:1027–1033. https://doi.org/10.1016/J.MEHY.2006.04.056
Majumdar A, Cesario WC, White-Grindley E et al (2012) Critical role of amyloid-like oligomers of Drosophila Orb2 in the persistence of memory. Cell 148:515–529. https://doi.org/10.1016/J.CELL.2012.01.004
Manjrekar J, Shah H (2020) Protein-based inheritance. Semin Cell Dev Biol 97:138–155. https://doi.org/10.1016/j.semcdb.2019.07.007
McKinley MP, Bolton DC, Prusiner SB (1983) A protease-resistant protein is a structural component of the scrapie prion. Cell 35:57–62
Nan H, Chen H, Tuite MF, Xu X (2019) A viral expression factor behaves as a prion. Nat Commun 10:359. https://doi.org/10.1038/s41467-018-08180-z
Nishina S, Kohsaka S, Yamaguchi Y et al (1999) PAX6 expression in the developing human eye. Br J Ophthalmol 83:723. https://doi.org/10.1136/bjo.83.6.723
Pallarès I, Iglesias V, Ventura S (2016) The Rho termination factor of Clostridium botulinum contains a prion-like domain with a highly amyloidogenic core. Front Microbiol. https://doi.org/10.3389/fmicb.2015.01516
Patel BK, Gavin-Smyth J, Liebman SW (2009) The yeast global transcriptional co-repressor protein Cyc8 can propagate as a prion. Nat Cell Biol 11:344–349. https://doi.org/10.1038/ncb1843
Prusiner SB (1982) Novel proteinaceous infectious particles cause scrapie. Science 216:136–144. https://doi.org/10.1126/science.6801762
Ross ED, Edskes HK, Terry MJ, Wickner RB (2005) Primary sequence independence for prion formation. Proc Natl Acad Sci USA 102:12825–12830. https://doi.org/10.1073/pnas.0506136102
Sabate R, Rousseau F, Schymkowitz J et al (2015) Amyloids or prions? That is the question. Prion 9:200–206. https://doi.org/10.1080/19336896.2015.1053685
Saupe SJ, Daskalov A (2012) The [Het-s] prion, an amyloid fold as a cell death activation trigger. PLoS Pathog 8:e1002687. https://doi.org/10.1371/journal.ppat.1002687
Schmidt MC, McCartney RR, Zhang X et al (1999) Std1 and Mth1 proteins interact with the glucose sensors to control glucose-regulated gene expression in Saccharomyces cerevisiae. Mol Cell Biol 19:4561
Shorter J, Lindquist S (2005) Prions as adaptive conduits of memory and inheritance. Nat Rev Genet 6:435–450. https://doi.org/10.1038/nrg1616
Si K (2015) Prions: what are they good for? Annu Rev Cell Dev Biol 31:149–169. https://doi.org/10.1146/annurev-cellbio-100913-013409
Si K, Lindquist S, Kandel ER (2003) A neuronal isoform of the Aplysia CPEB has prion-like properties. Cell 115:879–891. https://doi.org/10.1016/S0092-8674(03)01020-1
Sondheimer N, Lindquist S (2000) Rnq1: An epigenetic modifier of protein function in yeast. Mol Cell 5:163–172. https://doi.org/10.1016/S1097-2765(00)80412-8
Steinert JR (2015) Prion protein as a mediator of synaptic transmission. Commun Integr Biol 8:e1063753. https://doi.org/10.1080/19420889.2015.1063753
Suzuki G, Shimazu N, Tanaka M (2012) A yeast prion, Mod5, promotes acquired drug resistance and cell survival under environmental stress. Science 336(6079):355–359. https://doi.org/10.1126/science.1219491
Tan L, Williams MA, Khan MK et al (1999) Risk of transmission of bovine spongiform encephalopathy to humans in the United States: report of the council on scientific affairs. JAMA 281:2330. https://doi.org/10.1001/jama.281.24.2330
Tariq M, Wegrzyn R, Anwar S et al (2013) Drosophila GAGA factor polyglutamine domains exhibit prion-like behavior. BMC Genomics 14:374. https://doi.org/10.1186/1471-2164-14-374
Tuite MF, Serio TR (2010) The prion hypothesis: from biological anomaly to basic regulatory mechanism. Nat Rev Mol Cell Biol 11:823–833. https://doi.org/10.1038/nrm3007
Tuite MF, Staniforth GL, Cox BS (2015) [PSI+] turns 50. Prion 9:318–332. https://doi.org/10.1080/19336896.2015.1111508
Vitrenko YA, Gracheva EO, Richmond JE, Liebman SW (2007) Visualization of aggregation of the Rnq1 prion domain and cross-seeding interactions with Sup35NM. J Biol Chem 282:1779–1787. https://doi.org/10.1074/jbc.M609269200
Wickner RB (1994) [URE3] as an altered URE2 protein: evidence for a prion analog in Saccharomyces cerevisiae. Science 264:566–569. https://doi.org/10.1126/science.7909170
Wickner RB (2016) Yeast and fungal prions. Cold Spring Harb Perspect Biol 8(9):a023531. https://doi.org/10.1101/cshperspect.a023531
Wickner RB, Edskes HK, Bateman DA et al (2013) Amyloids and yeast prion biology. Biochemistry 52:1514–1527. https://doi.org/10.1021/bi301686a
Yuan AH, Hochschild A (2017) A bacterial global regulator forms a prion. Science 355:198–201. https://doi.org/10.1126/science.aai7776
Author information
Authors and Affiliations
Corresponding author
Additional information
Handling editor: David Alvarez-Ponce
Rights and permissions
About this article
Cite this article
Dixson, J.D., Azad, R.K. Prions: Roles in Development and Adaptive Evolution. J Mol Evol 88, 427–434 (2020). https://doi.org/10.1007/s00239-020-09944-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00239-020-09944-2