Abstract
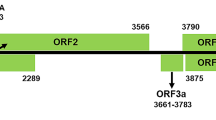
Double-stranded RNA and total RNA purified from sour cherry leaves (Prunus cerasus, cv. Amarelka Chvalkovicka) was analyzed by high-throughput sequencing. BLAST annotation identified contigs with homology to several already known cherry-infecting viruses (prune dwarf virus, prunus necrotic ringspot virus, prunus virus F, little cherry virus 1) as well as contigs with sequences more distantly related to those of members of the family Betaflexiviridae and in particular to prunus virus T of the genus Tepovirus. The full genome sequence of a putative virus (6,847 nucleotides [nt]; GenBank no. MT090966) was assembled and completed at the genome ends. The genome has a typical tepovirus organization, containing three overlapping open reading frames (ORFs), encoding a replication-associated protein, a movement protein and a capsid protein, respectively. Both its genome organization and its phylogenetic relationships show that the virus belongs to the genus Tepovirus, but considering the species demarcation criteria for the family Betaflexiviridae, it appears to represent a novel virus species, and we propose the name “cherry virus T” (ChVT) for this virus.


Similar content being viewed by others
References
Adams MJ, Candresse T, Hammond J, Kreuze JF, Martelli GP, Namba S. Pearson MN, Ryu KH, Saldarelli P, Yoshikawa N (2012) Family Betaflexiviridae. In: King AMQ, Adams MJ, Carstens EB, Lefkowitz EJ (eds.) Virus taxonomy. 9th Report of the ICTV. Elsevier Academic Press, Amsterdam, pp 920–941
Rubio M, Martínez-Gómez P, Marais A, Sánchez-Navarro J, Pallás V, Candresse T (2017) Recent advances and prospects in Prunus virology. Ann Appl Biol 171:125–138
Rubino L, Russo M, De Stradis A, Martelli GP (2012) Tepovirus, a novel genus in the family Betaflexiviridae. Arch Virol 157:1629–1633
Marais A, Faure C, Mustafayev E, Barone M, Alioto D, Candresse T (2015) Characterization by deep sequencing of prunus virus T, a novel Tepovirus infecting Prunus species. Phytopathology 105:135–140
Goh CJ, Park D, Lee JS, Davey PA, Pernice M, Ralph PJ, Hahn Y (2019) Zostera virus T - a novel virus of the genus Tepovirus identified in the eelgrass, Zostera muelleri. Acta Virol 63:366–372
Candresse T, Marais A, Faure C, Gentit P (2013) Association of Little cherry virus 1 with the Shirofugen stunt disease and characterization of the genome of a divergent LChV1 isolate. Phytopathology 103:293–298
Lee DH, Jin SG, Cai S, Pfeifer GP, O’Connor TR (2005) Repair of methylation damage in DNA and RNA by mammalian AlkB homologues. J Biol Chem 280:39448–39459
Acknowledgements
The authors thank the INRAE GenoToul Platform (Toulouse, France) for high-throughput sequencing.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Research involving human and animal participants
This manuscript presents original work that does not involve studies with humans or animals.
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Handling Editor: Ioannis E. Tzanetakis.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
The nucleotide sequence reported in this manuscript has been deposited in the GenBank database under accession number MT090966.
Rights and permissions
About this article
Cite this article
Marais, A., Šafářová, D., Navrátil, M. et al. Complete genome sequence of cherry virus T, a novel cherry-infecting tepovirus. Arch Virol 165, 1711–1714 (2020). https://doi.org/10.1007/s00705-020-04656-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00705-020-04656-w