Abstract
Liometopum apiculatum is a species of ants widely distributed in arid and semi-arid ecosystems where there is a relative food shortage compared with tropical ecosystems. L. apiculatum has established an ecological balance involving symbiotic interactions, which have allowed them to survive through mechanisms that are still unknown. Therefore, the aim of this study was to explore the metabolic potential of isolated bacteria from L. apiculatum using enzymatic activity assay and substrate assimilation. Results revealed a complex bacteria consortium belonging to Proteobacteria, Firmicutes, and Actinobacteria phylum. Most of the isolated bacteria showed activities associated with biopolymers degradation, from them Exiguobacterium and B. simplex showed the highest amylolytic activity (27 U/mg protein), while A. johnsonii and B. pumulis showed the highest cellulolytic and xylanolytic activities (1 and 2.9 U/mg protein, respectively). By other hand, some microorganisms such as S. ficaria, E. asburiae, P. agglomerans, A. johnsonii, S. rubidaea, S. marcescens, S. warneri, and M. hydrocarbonoxydans were able to grow up to 1000 mg/L of phthalates esters. These results not only revealed the important contribution of the symbionts in L apiculatum ants feeding habits, but also have shown a promising source of enzymes with potential biotechnological applications such as lignocellulosic biomass hydrolysis and bioremediation processes.
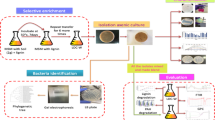
Graphic abstract







Similar content being viewed by others
Data availability
All generated data during this study is included in this published article (and its additional files). The spectral data have been deposited in the PeptideAtlas (https://www.peptideatlas.org/PASS/PASS01383). Request for materials should be addressed to the corresponding author.
References
Adams AS, Aylward FO, Adams SM, Erbilgin N, Aukema BH, Currie CR et al (2013) Mountain pine beetles colonizing historical and naive host trees are associated with a bacterial community highly enriched in genes contributing to terpene metabolism. Appl Environ Microbiol 79:3468–3475. https://doi.org/10.1128/AEM.0006813
Afrisham S, Badoei-Dalfard A, Namaki-shoushtari A, Karami Z (2016) Characterization of a thermostable, CaCl2 -activated and raw-starch hydrolyzing alpha-amylase from Bacillus licheniformis AT70: production under solid state fermentation by utilizing agricultural wastes. J Mol Catal B 132:98–106. https://doi.org/10.1016/j.molcatb.2016.07.002
Amir S, Hafidi M, Merlina G, Hamdi H, Jouraiphy A, El Gharous M et al (2005) Fate of phthalic acid esters during composting of both lagooning and activated sludges. Process Biochem 40:2183–2190. https://doi.org/10.1016/j.procbio.2004.08.012
Ascodeh A, Lagzian M (2012) Purification and characterization of a new glucoamylopullulanase from thermotolerant alkaliphilic Bacillus subtilis DR8806 of a hot mineral spring. Process Biochem 47:806–815. https://doi.org/10.1016/j.procbio.2012.02.018
Avellaneda-Torres LM, Guvera-Pulido CP, Torres-Rojas E (2014) Assessment of cellulolytic microorganisms in soils of Nevados Park, Colombia. Braz J Microbiol 45:1211–1220. https://doi.org/10.1590/s1517-83822014000400011
Benjamin S, Masai E, Kamimura N, Takahashi K, Anderson RC, Faisal PA (2017) Phthalates impact human health: epidemiological evidences and plausible mechanism of action. J Hazard Mater 340:360–383. https://doi.org/10.1016/j.jhazmat.2017.06.036
Castillo-Andrade A, García-Barrientos R, Ruiz-Cabrera MA, Rivera-Bautista C, Pérez-Martínez JD, González-García R et al (2015) Structural and physicochemical changes due to proteolytic deterioration of escamoles (Liometopum apiculatum M) a traditional Mexican food. J Ins Food Feed 1:271–280. https://doi.org/10.3920/JIFF2015.0031
Colman D, Toolson E, Takacs-Vesbach C (2012) Do diet and taxonomy influence insect gut bacterial communities? Mol Ecol 21:5124–5137. https://doi.org/10.1111/j.1365-294X.2012.05752.x
Díaz E (2004) Bacterial degradation of aromatic pollutants: a paradigm of metabolic versatility. Int Microbiol 7:173–180
Engel P, Moran NA (2013) The gut microbiota of insects—diversity in structure and function. FEMS Microbiol Rev 37:699–735. https://doi.org/10.1111/1574-6976.12025
Engel P, Martinson VG, Moran NA (2012) Functional diversity within the simple gut microbiota of the honey bee. Proc Natl Acad Sci 109:11002–11007. https://doi.org/10.1073/pnas.1202970109
Falade AO, Eyisi OAL, Mabinya LV, Nwodo UU, Okoh AI (2017) Peroxidase production and ligninolytic potentials of fresh water bacteria Raoultella ornithinolytica and Ensifer adhaerens. Biotechnol Rep 16:12–17. https://doi.org/10.1016/j.btre.2017.10.001
Fang X, Yano S, Inoue H, Sawayama S (2009) Strain improvement of Acremonium cellulolyticus for cellulase production by mutation. J Biosci Bioeng 107:256–261. https://doi.org/10.1016/j.jbiosc.2008.11.022
García-Herrera E, Méndez-Gallegos S, Talavera-Magaña D (2010) El género agave spp. en México: principales usos de importancia socioeconómica y agroecológica. RESPYN 5:109–129
Gekenidis MT, Studer P, Wüthrich S, Brunisholz R, Drissner D (2014) Beyond the matrix-assisted laser desorption ionization (MALDI) biotyping workflow: In search of microorganism-specific tryptic peptides enabling discrimination of subspecies. Appl Environ Microbiol 80:4234–4241. https://doi.org/10.1128/AEM.00740-14
González-Escobar JL, Grajales-Lagunes A, Adam S, Chagolla-López A, De Léon-Rodríguez A, Barba de la Rosa AP (2018) Microbiota of edible Liometopum apiculatum ant larvae reveals potential functions related to their nutritional value. Food Res Int J 109:497–505. https://doi.org/10.1016/j.foodres.2018.04.049
Gressel J (2018) Microbiome facilitated pest resistance: potential problems and uses. Pest Manage Sci 74:511–515. https://doi.org/10.1002/ps.4777
Gu JD, Li J, Wang Y (2005) Biochemical pathway and degradation of phthalate ester isomers by bacteria. Water Sci Technol 52:241–248
He Z, Xiao H, Lu T, Min H, Lu Z (2013) Biodegradation of di-n-butyl phthalate by a stable bacterial consortium, HD-1, enriched from activated sludge. Bioresour Technol 128:526–532. https://doi.org/10.1016/j.biortech.2012.10.107
Hoey-Chamberlain R, Rust MK, Klotz JH (2013) A Review of the biology, ecology and behaviour of velvety tree ants of North America. Sociobiology 60:1–10. https://doi.org/10.13102/sociobiology.v60i1.1-10
Huerta-Ocampo JA, García-Muñoz MS, Velarde-Salcedo AJ, Hernández-Domínguez EE, González-Escobar JL, Barrera-Pacheco A et al (2018) The proteome map of the escamolera ant (Liometopum apiculatum Mayr) larvae reveals immunogenic proteins and several hexamerin proteoforms. Comp Biochem Physiol D 28:107–121. https://doi.org/10.1016/j.cbd.2018.07.004
Kautz S, Rubin BER, Russell JA, Moreau CS (2013) Surveying the microbiome of ants: comparing 454 pyrosequencing with traditional methods to uncover bacterial diversity. Appl Environ Microbiol 79:525. https://doi.org/10.1128/AEM.03107-12
Kikuchi Y, Hayatsu M, Hosokawa T, Nagayama A, Tago K, Fukatsu T (2012) Symbiont-mediated insecticide resistance. Proc Natl Acad Sci 109:8618–8622. https://doi.org/10.1073/Pnas.1200231109
Kothari R, Tyagi VV, Pathak A (2010) Waste-to-energy: a way from renewable energy sources to sustainable development. Renew Sustain Energy Rev 14:3164–3170. https://doi.org/10.1016/j.rser.2010.05.005
Lama L, Calandrelli V, Gambacorta A, Nicolaus B (2004) Purification and characterization of thermostable xylanase and β-xylosidase by the thermophilic bacterium Bacillus thermantarcticus. Res Microbiol 155:283–239. https://doi.org/10.1016/j.resmic.2004.02.001
Li J, Gu JD, Pan L (2005) Transformation of dimethyl phthalate, dimethyl isophthalate and dimethyl terephthalate by Rhodococcus rubber Sa and modeling the processes using the modified Gompertz model. Int Biodeterior Biodegrad 55:223–232. https://doi.org/10.1016/j.ibiod.2004.12.003
Li F, Liu Y, Wang D, Zhang C, Yang Z, Lu S et al (2018) Biodegradation of di-(2-ethylhexyl) phthalate by a halotolerant consortium LF. PLoS ONE 13(10):e0204324. https://doi.org/10.1371/journal.pone.0204324
Li J, Zhang J, Yadav MP, Li X (2019) Biodegradability and biodegradation pathway of di-(2-ethylhexyl) phthalate by Burkholderia pyrrocinia B1213. Chemosphere 640–641:1121–1131. https://doi.org/10.1016/j.chemosphere.2019.02.194
Liang D, Zhang T, Fang HHP, He J (2008) Phthalates biodegradation in the environment. Appl Microbiol Biotechnol 80:183–198. https://doi.org/10.1007/s00253-008-1548-5
Lilly DG, Dang K, Webb CE, Doggett SL (2016) Evidence for metabolic pyrethorid resistance in the common Bed bug (Hemiptera: Cimicidae). J Econ Entomol Adv Access 109:1364–1368. https://doi.org/10.1093/jee/tow041
Lu Y, Tang F, Wang Y, Zhao J, Zeng X, Luo Q et al (2009) Biodegradation of dimethyl phthalate, diethyl phthalate and di-n-butyl phthalate by Rhodococcus sp. L4 isolated from activated sludge. J Hazard Mater 168:938–943. https://doi.org/10.1016/j.jhazmat.2009.02.126
Miller L (1959) Use of a dinitrosalicylic acid reagent for determination of reducing sugar. Anal Chem 31:426–428. https://doi.org/10.1021/ac60147a030
Pandey J, Chauhan A, Jain RK (2008) Integrative approaches for assessing the ecological sustainability of in situ bioremediation. FEMS Microbiol Rev 33:324–375. https://doi.org/10.1111/j.1574-6976.2008.00133.x
Pilon FM, Visotto LE, Guedes RNV, Oliveira MGA (2013) Proteolytic activity of gut bacteria isolated from the velvet bean caterpillar Anticarsia gemmatalis. J Comp Physiol B 183:735–747. https://doi.org/10.1007/s00360-013-0744-5
Pinheiro GL, Correa RF, Cunha RS, Cardoso AM, Chaia C, Clementino MM et al (2015) Isolation of aerobic cultivable cellulolytic bacteria from different regions of the gastrointestinal tract of giant land snail Achatina fulica. Front Microbiol 6:1–15. https://doi.org/10.3389/fmicb.2015.00860
Pinheiro GL, de Azevedo-Martins AC, Albano RM, de Souza W, Frases S (2017) Comprehensive analysis of the cellulolytic system reveals its potential for deconstruction of lignocellulosic biomass in a novel Streptomyces sp. Appl Microbiol Biotechnol 101:301–319. https://doi.org/10.1007/s00253-016-7851-7
Ramos-Elorduy J (2006) Threatened edible insects in Hidalgo, Mexico and some measures to preserve them. J Ethnobiol Ethnomed 2:1–10. https://doi.org/10.1186/1746-4269-2-51
Rastkari N, Jeddi MZ, Yunesian M, Ahmadkhaniha R (2018) Effect of sunlight exposure on phthalates migration from plastic containers to packaged juices. J Environ Health Sci Eng 16:27–33. https://doi.org/10.1007/s40201-018-0292-8
Sethi A, Slack JM, Kovaleva ES, Buchman GW, Scharf ME (2013) Lignin-associated metagene expression in a lignocellulose-digesting termite. Insect Biochem Mol Biol 43:91–101. https://doi.org/10.1016/j.ibmb.2012.10.001
Shi W, Xie S, Chen X, Sun S, Zhou X, Liu L et al (2013) Comparative genomic analysis of the endosymbionts of herbivorous insects reveals eco-environmental adaptations: Biotechnology applications. PLoS Genet. https://doi.org/10.1371/journal.pgen.1003131
Singh J, Batra N, Sobti RC (2004) Purification and characterisation of alkaline cellulase produced by a novel isolate, Bacillus sphaericus JS1. J Ind Microbiol Biotechnol 31:51–56. https://doi.org/10.1007/s10295-004-0114-0
Shinde AA, Shaikh FK, Padul MV, Kachole MS (2012) Bacillus subtillis RTSBA6 6.00, a new strain isolated from gut of Helicoverpa armigera (Lepidoptera: Noctuidae) produces chymotrypsin-like proteases. Saudi J Biol Sci 19:317–323. https://doi.org/10.1016/j.sjbs.2012.03.001
Su LJ, Lin H, Li Y, Zhang HF, Chen M, Gao XH, Wang FQ, Song AD (2014) Cellulolytic activity and structure of symbiotic bacteria in locust guts. Gen Mol Res 13:7926–7936. https://doi.org/10.4238/2014
Velasco C, Corona-Vargas M, Peña-Martinez R (2007) Liometopum apiculatum (Formicidae: Dolichoderinae) y su relacion trofobiotica con Hemiptera Sternorrhyncha en Tlaxco, Tlaxcala. México Acta Zoológica Mex 23:31–42
Visôtto LE, Oliveira MGA, Ribon AOB, Mares-Guia TR, Guedes RNC (2009) Characterization and identification of proteolytic bacteria from the gut of the velvet bean caterpillar (Lepidoptera: Noctuidae). Environ Entomol 38:1078–1085. https://doi.org/10.1603/022.038.0415
Wang Y, Li F, Ruan X, Song J, Lv L, Chai L et al (2017) Biodegradation of di-n-butyl phthalate by bacterial consortium LV-1 enriched from river sludge. PLoS ONE 12(5):e0178213. https://doi.org/10.1371/journal.pone.0178213
Warnecke F, Luginbühl P, Ivanova N, Ghassemian M, Richardson TH, Stege JT et al (2007) Metagenomic and functional analysis of hindgut microbiota of a wood-feeding higher termite. Nature 450:560–565. https://doi.org/10.1038/nature06269
Wu X, Liang R, Dai Q, Jin D, Wang Y, Chao W (2010) Complete degradation of di-n-octyl phthalate by biochemical cooperation between Gordonia sp. strain JDC-2 and Arthrobacter sp. strain JDC-32 isolated from activated sludge. J Hazard Mater 176:262–268. https://doi.org/10.1016/j.jhazmat.2009.11.022
Xia X, Gurr GM, Vasseur L, Zheng D, Zhong H, Qin B et al (2017) Metagenomic sequencing of Diamondback moth gut microbiome unveils key holobiont adaptations for herbivory. Front Microbiol 8:1–12. https://doi.org/10.3389/fmicb.2017.00663
Xu J, Lu Q, de Toledo RA, Shim H (2017) Degradation of di-2-ethylhexyl phthalate (DEHP) by an indigenous isolate Acinetobacter sp. SN13. Int Biodeterior Biodegrad 117:205–214. https://doi.org/10.1016/j.ibiod.2017.01.004
Yun J, Roh W, Whon W, Jung M, Kim M, Park D et al (2014) Insect gut bacterial diversity determined by environmental habitat, diet, developmental stage, and phylogeny of host. Appl Environ Microbiol 80:5254–5264. https://doi.org/10.1128/AEM.01226-14
Zhao HM, Hu RW, Chen XX, Chen XB, Lü HX, Li YW et al (2018) Biodegradation pathway of di-(2-ethylhexyl) phthalate by a novel Rhodococcus pyridinivorans XB and its bioaugmentation for remediation of DEHP contaminated soil. Sci Total Environ 640–641:1121–1131. https://doi.org/10.1016/j.scitotenv.2018.05.334
Zhang J, Zhang C, Zhu Y, Li J, Li X (2018) Biodegradation of seven phthalate esters by Bacillus mojavensis B1811. Int Biodeterior Biodegrad 132:200–207. https://doi.org/10.1016/j.ibiod.2018.04.006
Acknowledgements
JLGE and MAPC thank CONACYT fellowship 386100 and 331852, respectively. Thanks to A Barrera-Pacheco and VE Balderas-Hernández for their technical assistance.
Funding
This work was financially supported by Fondo Sectorial SAGARPA-CONACYT-B-S-3804, FSAG/ST/025/18 and CONACYT Problemas Nacionales 4601.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that there is no conflict of interests regarding with the publication of this paper.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
González-Escobar, J.L., Pereyra-Camacho, M.A., De Léon-Rodríguez, A. et al. Biodegradation of recalcitrant compounds and phthalates by culturable bacteria isolated from Liometopum apiculatum microbiota. World J Microbiol Biotechnol 36, 73 (2020). https://doi.org/10.1007/s11274-020-02850-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11274-020-02850-1