Abstract
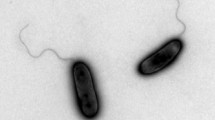
A Gram-negative, aerobic, motile, non-spore-forming and rod-shaped bacterium, designated strain B18-69 T, was isolated from oil-well production liquid in Baolige oilfield, China. The strain was able to grow at pH 6–9.5 (optimum at pH 7), in 0–4% (w/v) NaCl (optimum at 0.5–1%, w/v) and at 35–60 °C (optimum at 55 °C). Major cellular fatty acids were C16:0, C19:0 cyclo ω8c, C17:0 cyclo and C18:1 ω7c. The predominant respiratory quinone was ubiquinone 8. Major polar lipids were phosphatidylethanolamine (PE), phosphatidylglycerol (PG), diphosphatidylglycerol (DPG) and phosphatidylcholine (PC). Phylogenetic analysis based on 16S rRNA gene sequences revealed that strain B18-69 T was most closely related to Tepidiphilus margaritifer DSM 15129 T (98.8% similarity). The draft genome of strain B18-69 T was composed of 2,250,419 bp, and the G+C content was 64.6 mol%. Average nucleotide identity (ANI) and digital DNA–DNA hybridization (dDDH) values between strain B18-69 T and T. margaritifer DSM 15129 T were 90.9% and 68.9%, respectively. Genotypic and phenotypic features indicate that strain B18-69 T represents a novel species of the genus Tepidiphilus, for which the name Tepidiphilus baoligensis sp. nov. is proposed. The type strain is B18-69 T (= CGMCC 1.13573 T = KCTC 62782 T).


Similar content being viewed by others
Data Availability
The GenBank accession numbers for the 16S rRNA gene sequence and the draft genome of strain B18-69T are MH209249 and JAAAUB000000000, respectively.
Abbreviations
- dDDH:
-
Digital DNA–DNA hybridization
- GGDC:
-
Genome-to-genome distance calculation
- ANI:
-
Average nucleotide identity
- ML:
-
Maximum-likelihood
- MP:
-
Maximum-parsimony
- NJ:
-
Neighbor-joining
- DPG:
-
Diphosphatidylglycerol
- PC:
-
Phosphatidylcholine
- PE:
-
Phosphatidylethanolamine
- PG:
-
Phosphatidylglycerol
- HPLC:
-
High-performance liquid chromatography
- TSA:
-
Tryptic soy agar
- CHES:
-
2-(Cyclohexylamino)ethanesulfonic acid
- MES:
-
2-(N-Morpholino)ethanesulfonic acid
- MOPS:
-
3-(N-Morphine)propanesulfonic acid
- TAPS:
-
N-Tris(hydroxymethyl)methyl-3-aminopropanesulfonic acid
References
Manaia CM, Nogales B, Nunes OC (2003) Tepidiphilus margaritifer gen. nov., sp. nov., isolated from a thermophilic aerobic digester. Int J Syst Evol Microbiol 53:1405–1410
Poddar A, Lepcha RT, Das SK (2014) Taxonomic study of the genus Tepidiphilus: transfer of Petrobacter succinatimandens to the genus Tepidiphilus as Tepidiphilus succinatimandens comb. nov., emended description of the genus Tepidiphilus and description of Tepidiphilus thermophilus sp. nov., isolated from a terrestrial hot spring. Int J Syst Evol Microbiol 64:228–235
Salinas MB, Fardeau ML, Cayol JL, Casalot L, Patel BKC, Thomas P, Garcia JL, Ollivier B (2004) Petrobacter succinatimandens gen. nov., sp. nov., a moderately thermophilic, nitrate-reducing bacterium isolated from an Australian oil well. Int J Syst Evol Microbiol 54:645–649
Kieft TL, Fredrickson JK, Onstott TC, Gorby YA, Kostandarithes HM, Bailey TJ, Kennedy DW, Li SW, Plymale AE, Spadoni CM, Gray MS (1999) Dissimilatory reduction of Fe(III) and other electron acceptors by a Thermus isolate. Appl Environ Microbiol 65:1214–1221
Gerhardt P, Murray RGE, Costilow RN, Nester EW, Wood WA, Krieg NR, Phillips GB (1981) Manual of methods for general bacteriology. American Society for Microbiology, Washington, DC
Dong XZ, Cai MY (2001) Determinative manual for routine bacteriology. Scientific, Beijing (in Chinese)
Degryse E, Glansdorff N, Pierard A (1978) A comparative analysis of extreme thermophilic bacteria belonging to the genus Thermus. Arch Microbiol 117:189–196
Collins MD, Pirouz T, Goodfellow M, Minnikin DE (1977) Distribution of menaquinones in actinomycetes and corynebacteria. J Gen Microbiol 100:221–230
Minnikin DE, Collins MD, Goodfellow M (1979) Fatty acid and polar lipid composition in the classification of Cellulomonas, Oerskovia and related taxa. J Appl Bacteriol 47:87–95
Marmur J (1961) A procedure for the isolation of deoxyribonucleic acid from micro-organisms. J Mol Biol 3:208–218
Lane DJ (1991) 16S/23S rRNA sequencing. In: Stackebrandt E, Goodfellow M (eds) Nucleic acid techniques in bacterial systematics. Wiley, Chichester, pp 115–175
Yoon SH, Ha SM, Kwon S, Lim J, Kim Y, Seo H, Chun J (2017) Introducing EzBioCloud: A taxonomically united database of 16S rRNA and whole genome assemblies. Int J Syst Evol Microbiol 67:1613–1617
Kumar S, Stecher G, Tamura K (2016) MEGA7: molecular evolutionary genetics analysis version 7.0 for bigger datasets. Mol Biol Evol 33:1870-1874
Luo R, Liu B, Xie Y, Li Z, Huang W, Yuan J, He G, Chen Y, Pan Q, Liu Y, Tang J, Wu G, Zhang H, Shi Y, Liu Y, Yu C, Wang B, Lu Y, Han C, Cheung DW, Yiu SM, Peng S, Xiaoqian Z, Liu G, Liao X, Li Y, Yang H, Wang J, Lam TW, Wang J (2012) SOAPdenovo2: an empirically improved memory-efficient short-read de novo assembler. Gigascience 1:18
Tatusova T, DiCuccio M, Badretdin A, Chetvernin V, Nawrocki EP, Zaslavsky L, Lomsadze A, Pruitt KD, Borodovsky M, Ostell J (2016) NCBI prokaryotic genome annotation pipeline. Nucleic Acids Res 44:6614–6624
Yoon SH, Ha SM, Lim JM, Kwon SJ, Chun J (2017) A large-scale evaluation of algorithms to calculate average nucleotide identity. Antonie Van Leeuwenhoek 110:1281–1286
Meier-Kolthoff JP, Göker M (2019) TYGS is an automated high-throughput platform for state-of-the-art genome-based taxonomy. Nat Commun 10:2182
Meier-Kolthoff JP, Auch AF, Klenk HP, Göker M (2013) Genome sequence-based species delimitation with confidence intervals and improved distance functions. BMC Bioinform 14:60–73
Chun J, Oren A, Ventosa A, Christensen H, Arahal DR, da Costa MS, Rooney AP, Yi H, Xu XW, Meyer SD, Trujillo ME (2018) Proposed minimal standards for the use of genome data for the taxonomy of prokaryotes. Int J Syst Evol Microbiol 68:461–466
Acknowledgements
This work was supported by the National Key R&D Program of China (2018YFA0902101) and the Huabei Oilfield Company (2017-HB-B19).
Author information
Authors and Affiliations
Contributions
XZ and XM tested the phenotypic and chemotaxonomic characteristics. GW and JLY isolated the strain, and participated in the morphology investigation and the 16S rRNA gene and genome sequence. YX designed the tests, analyzed the data and prepared the manuscript. JY and YM conceived of the study and participated in the data analysis and paper writing. All authors approved the manuscript and its submission.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that no competing interests exist.
Ethical Statement
The authors declare that no ethical issues exist.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Zhang, X., Wang, G., Ma, X. et al. Tepidiphilus baoligensis sp. nov., a Novel Bacterium of the Family Hydrogenophilaceae Isolated from an Oil Reservoir. Curr Microbiol 77, 1939–1944 (2020). https://doi.org/10.1007/s00284-020-01983-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00284-020-01983-8