Abstract
The olive tree is an economically important fruit tree, widely spread in the Mediterranean Basin. The recent availability of two full genomes of this plant offers an opportunity to characterize its disease resistance gene repertoire. The nucleotide-binding site (NBS) gene family accounts for the largest number of known disease resistance genes and is one of the largest gene families in plant genomes. Here, we have identified 270 regular NBS-type genes in the olive genome, roughly representing 0.5% of whole-genome protein-coding genes in this species. A systematic characterization of this gene set was conducted on the bases of conserved protein signatures, gene duplications, phylogenetic relationships, selection pressure, and expression evidence. Several structural features were particularly pronounced in O. europaea, including a very small proportion of TIR-type NBS genes, as well as numerous non-canonical functional domains associated to the NBS. Analyses of duplication and selection pressure strongly suggest that recent duplication, in conjunction with positive selection, played a remarkable role in the evolution of NBS-encoding genes in olive. Based on the phylogenetic tree produced, three pairs of possible orthologs between the olive NBS genes (OeNBS) and those from Arabidopsis thaliana, Oryza sativa Indica, and O. sativa Japonica were identified and could be of interest to characterize the uncharacterized function of the OeNBS genes in olive. Various expression patterns of olive NBS-encoding genes in different tissues were observed using expressed sequence tags (ESTs). Interestingly, we identified a pair of duplicated NLRs integrating an N-terminal RPW8 domain, which are possibly expressed in response to Verticillium dahliae, in addition to a gene encoding C-terminal Armadillo repeat domain, which is possibly induced upon colonization by the endophytic bacterium Pseudomonas fluorescens. This work is the first to characterize NBS family members and report on their architectural, evolutionary, and functional features. Our findings lay an important multifaceted foundation for a deeper understanding of R genes biology in olive tree and provide valuable information for further functional studies.





Similar content being viewed by others
Abbreviations
- OeNBS:
-
Olea europaea NBS genes
- NBS:
-
Nucleotide-binding site
- LRR:
-
Leucine-rich repeat (LRR) domain
- EST:
-
Expressed sequence tag
- Ka:
-
Nonsynonymous substitution rate
- Ks:
-
Synonymous substitution rate
- Mya:
-
Million years ago
- WGD:
-
Whole-genome duplication
- NLRs:
-
NOD-like receptors
- APAF1:
-
The apoptotic protease activating factor 1 of Homo sapiens
References
Allan AC, Hellens RP, Laing WA (2008) MYB transcription factors that colour our fruit. Cell 13:99–102. https://doi.org/10.1016/j.tplants.2007.11.012
Andersen EJ, Ali S, Byamukama E, Yen Y, Nepal MP (2018) Disease resistance mechanisms in plants. Genes. 9:339. https://doi.org/10.3390/genes9070339
Arya P, Kumar G, Acharya V, Singh AK (2014) Genome-wide identification and expression analysis of NBS-encoding genes in Malus x domestica and expansion of NBS genes family in Rosaceae. PLoS One 9:e107987. https://doi.org/10.1371/journal.pone.0107987
Bailey TL, Elkan C (1995) Unsupervised learning of multiple motifs in biopolymers using expectation maximization. Mach Learn 21:51. https://doi.org/10.1007/BF00993379
Baldoni L, Guerrero C, Sossey-Aloui K, Abbott AG, Angiolillo A, Lumaret R (2002) Phylogenetic relationships among Olea species based on nucleotide variation at a non-coding chloroplast DNA region. Plant Biology (Stuttg) 4:346–351. https://doi.org/10.1055/s-2002-32338
Barghini E, Natali L, Giordani T, Cossu RM, Scalabrin S, Cattonaro F, Šimková H, Vrána J, Doležel J, Morgante M, Cavallini A (2015) LTR retrotransposon dynamics in the evolution of the olive (Olea europaea) genome. DNA Res 22(1):91–100. https://doi.org/10.1093/dnares/dsu042
Baroncelli R, Amby DB, Zapparata A, Sarrocco S, Vannacci G, Floch GL, Harrison RJ, Holub E, Sukno SA, Sreenivasaprasad S, Thon MR (2016) Gene family expansions and contractions are associated with host range in plant pathogens of the genus Colletotrichum. BMC Genomics 17:555. https://doi.org/10.1186/s12864-016-2917-6
Besnard G, Rubio de Casas R (2016) Single vs multiple independent olive domestications: the jury is (still) out. New Phytol 209:466–470. https://doi.org/10.1111/nph.13518
Besnard G, Rubio De Casas R, Vargas P (2003) A set of primers for length and nucleotide-substitution polymorphism in chloroplastic DNA of Olea europaea L. (Oleaceae). Mol Ecol Notes 3:651–653. https://doi.org/10.1046/j.1471-8286.2003.00547.x
Besnard G, Terral J-F, Cornille A (2018) On the origins and domestication of the olive: a review and perspectives. Ann Bot 121:385–403. https://doi.org/10.1093/aob/mcx145
Blanc G, Wolfe KH (2004) Widespread paleopolyploidy in model plant species inferred from age distributions of duplicate genes. Plant Cell 16:1667–1678. https://doi.org/10.1105/tpc.021345
Borrelli GM, Mazzucotelli E, Marone D, Crosatti C, Michelotti V, Valè G, Mastrangelo AM (2018) Regulation and evolution of NLR genes: a close interconnection for plant immunity. Int J Mol Sci 19:1662. https://doi.org/10.3390/ijms19061662
Bouktila D, Khalfallah Y, Habachi-Houimli Y, Mezghani-Khemakhem M, Makni M, Makni H (2015) Full-genome identification and characterization of NBS-encoding disease resistance genes in wheat. Mol Gen Genomics 290(1):257–271. https://doi.org/10.1007/s00438-014-0909-2
Bracci T, Busconi M, Fogher C, Sebastiani L (2011) Molecular studies in olive (Olea europaea L.): overview on DNA markers applications and recent advances in genome analysis. Plant Cell Rep 30:449–462. https://doi.org/10.1007/s00299-010-0991-9
Breton C, Pinatel C, Terral J-F, Médial F, Bonhomme F, Bervillé A (2009) The olive domestication in the Mediterranean basin. CR Biologies. https://doi.org/10.1016/j.crvi.2009.08.001
Breton CM, Farinelli D, Shafiq S, Heslop-Harrison JS, Sedgley M, Bervillé AJ (2014) The self-incompatibility mating system of the olive (Olea europaea L.) functions with dominance between S-alleles. Tree Genet Genomes 10:1055–1067. https://doi.org/10.1007/s11295-014-0742-0
Bull PC, Cox DW (1994) Wilson disease and Menkes disease: new handles on heavy-metal transport. Trends Genet 10(7):246–252. https://doi.org/10.1016/0168-9525(94)90172-4
Cabanás G-L, Schiliro E, Valverde-Corredor A, Mercado-Blanco J (2015) Systemic responses in a tolerant olive (Olea europaea L.) cultivar upon root colonization by the vascular pathogen Verticillium dahlia. Front Microbiol 16:928. https://doi.org/10.3389/fmicb.2015.00928
Cheng Y, Li X, Jiang H, Ma W, Miao W, Yamada T, Zhang M (2012a) Systematic analysis and comparison of nucleotide-binding site disease resistance genes in maize. FEBS J 279:2431–2443. https://doi.org/10.1111/j.1742-4658.2012.08621.x
Cheng Y, Zhou Y, Yang Y, Chi YJ, Zhou J, Chen JY, Wang F, Fan B, Shi K, Zhou YH, Yu JQ, Chen Z (2012b) Structural and functional analysis of VQ motif-containing proteins in Arabidopsis as interacting proteins of WRKY transcription factors. Plant Physiol 159:810–825. https://doi.org/10.1104/pp.112.196816
Chisholm ST, Coaker G, Day B, Staskawicz BJ (2006) Host-microbe interactions: shaping the evolution of the plant immune response. Cell 124:803–814. https://doi.org/10.1016/j.cell.2006.02.008
Christopoulou M, Wo SR-C, Kozik A, McHale LK, Truco M-J, Wroblewski T, Michelmore RW (2015) Genome-wide architecture of disease resistance genes in lettuce. G3 (Bethesda) 5(12):2655–2669. https://doi.org/10.1534/g3.115.020818
Cimato A, Attilio C (2011) Worldwide diffusion and relevance of olive culture. In: Schena L, Agosteo GE, Cacciola SO, Sergeeva V (eds) Olive diseases and disorders. Transworld Research Network, Kerala
Cominelli E, Tonelli C (2009) A new role for plant R2R3-MYB transcription factors in cell cycle regulation. Cell Res 19:1231–1232. https://doi.org/10.1038/cr.2009.123
Cruz F, Julca I, Gómez-Garrido J, Loska D, Marcet-Houben M, Cano E, Galán B, Frias L, Ribeca P, Derdak S, Gut M, Sánchez-Fernández M, García JL, Gut IG, Vargas P, Alioto TS, Gabaldón T (2016) Genome sequence of the olive tree, Olea europaea. GigaScience 5:29. https://doi.org/10.1186/s13742-016-0134-5
Dangl JL, Jones JDG (2001) Plant pathogens and integrated defence responses to infection. Nature 411:826–833. https://doi.org/10.1038/35081161
Diez CM, Trujillo I, Martinez-Urdiroz N, Barranco D, Rallo L, Marfil P, Gaut BS (2015) Olive domestication and diversification in the Mediterranean Basin. New Phytol 206:436–447. https://doi.org/10.1111/nph.13181
Dodds PN, Rathjen JP (2010) Plant immunity: towards an integrated view of plant-pathogen interactions. Nat Rev Genet 8:539–548. https://doi.org/10.1038/nrg2812
Domman D, Collingro A, Lagkouvardos I, Gehre L, Weinmaier T, Rattei T, Subtil A, Horn M (2014) Massive expansion of ubiquitination-related gene families within the Chlamydiae. Mol Biol Evol 31(11):2890–2904. https://doi.org/10.1093/molbev/msu227
Dykema P, Sipes P, Marie A, Biermann B, Crowell D, Randall S (1999) A new class of proteins capable of binding transition metals. Plant Mol Biol 41:139–150
Ellis JG (2016) Integrated decoys and effector traps: how to catch a plant pathogen. BMC Biol 14:13. https://doi.org/10.1186/s12915-016-0235-8
FAOSTAT (2019) Food and agriculture organization of the United Nations. Statistics division. http://faostat.fao.org. Accessed 21.05.19
Flor HH (1971) Current status of the gene-for-gene concept. Annu Rev Phytopathol 9:275–296
Garrido-Gala J, Higuera JJ, Muñoz-Blanco J, Amil-Ruiz F, Caballero JL (2019) The VQ motif-containing proteins in the diploid and octoploid strawberry. Sci Rep 9, article number: 4942. https://doi.org/10.1038/s41598-019-41210-4
Giampetruzzi A, Morelli M, Saponari M, Loconsole G, Chiumenti M, Boscia D, Savino VN, Martelli GP, Saldarelli P (2016) Transcriptome profiling of two olive cultivars in response to infection by the CoDiRO strain of Xylella fastidiosa subsp. pauca. BMC Genomics 17:475. https://doi.org/10.1186/s12864-016-2833-9
Grasso F, Coppola M, Carbone F, Baldoni L, Alagna F, Perrotta G, Pérez-Pulido AJ, Garonna A, Facella P, Daddiego L, Lopez L, Vitiello A, Rao R, Corrado G (2017) The transcriptional response to the olive fruit fly (Bactrocera oleae) reveals extended differences between tolerant and susceptible olive (Olea europaea L.) varieties. PLoS one 12(8):e0183050. https://doi.org/10.1371/journal.pone.0183050
Gros-Balthazard M, Besnard G, Sarah G, Holtz Y, Leclercq J, Santoni S, Wegmann D, Glémin S, Khadari B (2019) Evolutionary transcriptomics reveals the origins of olives and the genomic changes associated with their domestication. 100:143–157. https://doi.org/10.1111/tpj.14435
Gu Y, Xing S, He C (2016) Genome-wide analysis indicates lineage-specific gene loss during Papilionoideae evolution. Genome Biol Evol 8:635–648. https://doi.org/10.1093/gbe/evw021
Heywood VH (2012) The role of New World biodiversity in the transformation of Mediterranean landscapes and culture. Bocconea 24:69–93
Hung IH, Casareno RLB, Labesse G, Mathews FS, Gitlin JD (1998) HAH1 is a copper-binding protein with distinct amino acid residues mediating copper homeostasis and antioxidant defense. J Biol Chem 273:1749–1754. https://doi.org/10.1074/jbc.273.3.1749
Jiang SY, Sevugan M, Ramachandran S (2018) Valine-glutamine (VQ) motif coding genes are ancient and non-plant-specific with comprehensive expression regulation by various biotic and abiotic stresses. BMC Genomics 19:342. https://doi.org/10.1186/s12864-018-4733-7
Julca I, Marcet-Houben M, Vargas P, Gabaldón T (2018) Phylogenomics of the olive tree (Olea europaea) disentangles ancient allo- and autopolyploidizations in Lamiales. BMC Biol 16:15. https://doi.org/10.1186/s12915-018-0482-y
Kaniewski D, Van Campo E, Boiy T, Terral JF, Khadari B, Besnard G (2012) Primary domestication and early uses of the emblematic olive tree: palaeobotanical, historical and molecular evidences from the Middle East. Biol Rev Camb Philos Soc 87:885–899. https://doi.org/10.1111/j.1469-185X.2012.00229.x
Kim S, Park M, Yeom S-I, Kim Y-M, Lee JM, Lee H-A et al (2014) Genome sequence of the hot pepper provides insights into the evolution of pungency in Capsicum species. Nat Genet 46:270–278. https://doi.org/10.1038/ng.2877
Kippert F, Gerloff DL (2009) Highly sensitive detection of individual HEAT and ARM repeats with HHpred and COACH. PLoS One 4:e7148. https://doi.org/10.1371/journal.pone.0007148
Kohler A, Rinaldi C, Duplessis S, Baucher M, Geelen D, Duchaussoy F, Meyers BC, Boerjan W, Martin F (2008) Genome-wide identification of NBS resistance genes in Populus trichocarpa. Plant Mol Biol 66:619–636. https://doi.org/10.1007/s11103-008-9293-9
Krattinger SG, Keller B (2016) Trapping the intruder — immune receptor domain fusions provide new molecular leads for improving disease resistance in plants. Genome Biol 17:23. https://doi.org/10.1186/s13059-016-0891-6
Kristensen DM, Wolf YI, Mushegian AR, Koonin EV (2011) Computational methods for gene orthology inference. Brief Bioinform 12(5):379–391. https://doi.org/10.1093/bib/bbr030
Kroj T, Chanclud E, Michel-Romiti C, Grand X, Morel J-B (2016) Integration of decoy domains derived from protein targets of pathogen effectors into plant immune receptors is widespread. New Phytol 210:618–626. https://doi.org/10.1111/nph.13869
Leyva-Pérez MO, Jiménez-Ruiz J, Gómez-Lama Cabanás C, Valverde-Corredor A, Barroso JB, Luque F, Mercado-Blanco J (2018) Tolerance of olive (Olea europaea) cv Frantoio to Verticillium dahliae relies on both basal and pathogen-induced differential transcriptomic responses. 217:671–686. https://doi.org/10.1111/nph.14833
Li W, Godzik A (2006) CD-HIT: a fast program for clustering and comparing large sets of protein or nucleotide sequences. Bioinformatics Applications Note 22:1658–1659. https://doi.org/10.1093/bioinformatics/btl158
Liu J, Liu X, Dai L, Wang G (2007) Recent progress in elucidating the structure, function and evolution of disease resistance genes in plants. J Genet Genomics 34:765–776. https://doi.org/10.1016/S1673-8527(07)60087-3
Lozano R, Ponce O, Ramirez M, Mostajo N, Orjeda G (2012) Genome-wide identification and mapping of NBS-encoding resistance genes in Solanum tuberosum group Phureja. PLoS One 7(4):e34775. https://doi.org/10.1371/journal.pone.0034775
Lukasik E, Takken FLW (2009) STANDing strong, resistance proteins instigators of plant defence. Curr Opin Plant Biol 12:427–436. https://doi.org/10.1016/j.pbi.2009.03.001
Lupas A, Van Dyke M, Stock J (1991) Predicting coiled coils from protein sequences. Science 252:1162–1164. https://doi.org/10.1126/science.252.5009.1162
Lynch M, Conery JS (2000) The evolutionary fate and consequences of duplicate genes. Science 290(5494):1151–1155. https://doi.org/10.1126/science.290.5494.1151
Magadum S, Banerjee U, Murugan P, Gangapur D, Ravikesavan R (2013) Gene duplication as a major force in evolution. J Genet 92:155–161. https://doi.org/10.1007/s12041-013-0212-8
Maqbool A, Saitoh H, Franceschetti M, Stevenson CEM, Uemura A, Kanzaki H, Kamoun S, Terauchi R, Banfield MJ (2015) Structural basis of pathogen recognition by an integrated HMA domain in a plant NLR immune receptor. eLife. 4:e08709. https://doi.org/10.7554/eLife.08709
Mariotti R, Cultrera NGM, Díez CM, Baldoni L, Rubini A (2010) Identification of new polymorphic regions and differentiation of cultivated olives (Olea europaea L.) through plastome sequence comparison. BMC Plant biol 10:211. https://doi.org/10.1186/1471-2229-10-211
Mauricio FN, Soratto TAT, Diogo JA, Boscariol-Camargo RL, De Souza AA, Coletta-Filho HD, Silva JAA, Medeiros AH, Machado MA, Cristofani-Yaly M (2019) Analysis of defense-related gene expression in citrus hybrids infected by Xylella fastidiosa. Phytopathology 109(2):301–306. https://doi.org/10.1094/PHYTO-09-18-0366-FI
McLellan H, Boevink PC, Armstrong MR, Pritchard L, Gomez S, Morales J, Whisson SC, Beynon JL, Birch PRJ (2013) An RxLR effector from Phytophthora infestans prevents relocalisation of two plant NAC transcription factors from the endoplasmic reticulum to the nucleus. PLoS Pathog 9:e1003670. https://doi.org/10.1371/journal.ppat.1003670
Mercado-Blanco J, Rodríguez-Jurado D, Hervás A, Jiménez-Díaz RM (2004) Suppression of Verticillium wilt in olive planting stocks by root-associated fluorescent Pseudomonas spp. Biol Control 30:474–486. https://doi.org/10.1016/j.biocontrol.2004.02.002
Meyers BC, Dickerman AW, Michelmore RW, Sivaramakrishnan S, Sobral BW, Yong ND (1999) Plant disease resistance genes encode members of an ancient and diverse protein family within nucleotide-binding superfamily. Plant J 20:317–332. https://doi.org/10.1046/j.1365-313X.1999.00606.x
Meyers BC, Kozik A, Griego A, Kuang H, Michelmore RW (2003) Genome-wide analysis of NBS-LRR–encoding genes in Arabidopsis. Plant Cell 15(4):809–834. https://doi.org/10.1105/tpc.009308
Meyers BC, Kaushik S, Nandety RS (2005) Evolving disease resistance genes. Curr Opin Plant Biol 8:129–134. https://doi.org/10.1016/j.pbi.2005.01.002
Nei M, Gojobori T (1986) Simple methods for estimating the numbers of synonymous and nonsynonymous nucleotide substitutions. Mol Biol Evol 3:418–426. https://doi.org/10.1093/oxfordjournals.molbev.a040410
Neupane S, Andersen EJ, Neupane A, Nepal MP (2018) Genome-wide identification of NBS-encoding resistance genes in sunflower (Helianthus annuus L.). Genes (Basel) 9(8):384. https://doi.org/10.3390/genes9080384
Nuruzzaman M, Sharoni M, Kikuchi S (2013) Roles of NAC transcription factors in the regulation of biotic and abiotic stress responses in plants. Front Microbiol 4:248. https://doi.org/10.3389/fmicb.2013.00248
Okuyama Y, Kanzaki H, Abe A, Yoshida K, Tamiru M, Saitoh H, Fujibe T, Matsumura H, Shenton M, Clark DG, Undan J, Ito A, Sone T, Terauchi R (2011) A multifaceted genomics approach allows the isolation of the rice Pia-blast resistance gene consisting of two adjacent NBS–LRR protein genes. Plant J 66:467–479. https://doi.org/10.1111/j.1365-313X.2011.04502.x
Pan Q, Wendel J, Fluhr R (2000) Divergent evolution of plant NBS–LRR resistance gene homologues in dicot and cereal genomes. J Mol Evol 50:203–213. https://doi.org/10.1007/s002399910023
Reboredo-Rodríguez P, Varela-López A, Forbes-Hernández TY, Gasparrini M, Afrin S, Cianciosi D, Zhang J, Manna PP, Bompadre S, Quiles JL, Battino M, Giampieri F (2018) Phenolic compounds isolated from olive oil as nutraceutical tools for the prevention and management of cancer and cardiovascular diseases. Int J Mol Sci 19(8). https://doi.org/10.3390/ijms19082305
Roka L, Koudounas K, Daras G, Zoidakis J, Vlahou A, Kalaitzis P, Hatzopoulos P (2018) Proteome of olive non-glandular trichomes reveals protective protein network against (a)biotic challenge. J Plant Physiol 231:210–218. https://doi.org/10.1016/j.jplph.2018.09.016
Sarris PF, Cevik V, Dagdas G, Jones JDG, Krasileva KV (2016) Comparative analysis of plant immune receptor architectures uncovers host proteins likely targeted by pathogens. BMC Biol 14:8. https://doi.org/10.1186/s12915-016-0228-7
Saumitou-Laprade P, Vernet P, Vekemans X, Billiard S, Gallina S, Essalouh L, Mhaïs A, Moukhli A, El Bakkali A, Barcaccia G, Alagna F, Mariotti R, Cultrera NGM, Pandolfi S, Rossi M, Khadari B, Baldoni L (2017) Elucidation of the genetic architecture of self-incompatibility in olive: evolutionary consequences and perspectives for orchard management. Evol Appl 10:867–880. https://doi.org/10.1111/eva.12457
Schiliro E, Ferrara M, Nigro F, Mercado-Blanco J (2012) Genetic responses induced in olive roots upon colonization by the biocontrol endophytic bacterium Pseudomonas fluorescens PICF7. PLoS One 7:e48646. https://doi.org/10.1371/journal.pone.0048646
Schwarz R, Dayhoff M (1979) Matrices for detecting distant relationships. In: Dayhoff M (ed) Atlas of protein sequences. National Biomedical Research Foundation, Silver Spring, pp 353–358
Sharma M, Pandey GK (2015) Expansion and function of repeat domain proteins during stress and development in plants. Front Plant Sci 6:1218. https://doi.org/10.3389/fpls.2015.01218
Song H, Nan Z (2014) Genome-wide analysis of nucleotide-binding site disease resistance genes in Medicago truncatula. Chin Sci Bull 59:1129–1138. https://doi.org/10.1007/s11434-014-0155-3
Suzuki N, Yamaguchi Y, Koizumi N, Sano H (2002) Functional characterization of a heavy metal binding protein CdI19 from Arabidopsis. Plant J 32:165–173. https://doi.org/10.1046/j.1365-313X.2002.01412.x
Tamura K, Peterson D, Peterson N, Steecher G, Nei M, Kumar S (2011) MEGA: molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol Biol Evol 28:2731–2739. https://doi.org/10.1093/molbev/msr121
Tan S, Wu S (2012) Genome wide analysis of nucleotide-binding site disease resistance genes in Brachypodium distachyon. Comp Funct Genomics 418208. https://doi.org/10.1155/2012/418208
Tarr DE, Alexander HM (2009) TIR-NBS-LRR genes are rare in monocots: evidence from diverse monocot orders. BMC Research Notes 2:197. https://doi.org/10.1186/1756-0500-2-197
Tatusov RL, Koonin EV, Lipman DJ (1997) A genomic perspective on protein families. Science 278:631–637. https://doi.org/10.1126/science.278.5338.631
Thornton JW, DeSalle R (2000) Gene family evolution and homology: genomics meets phylogenetics. Annu Rev Genomics Hum Genet 1:41–73. https://doi.org/10.1146/annurev.genom.1.1.41
Timmerman-Vaughan GM, Frew TJ, Weeden NF (2000) Characterization and linkage mapping of R-gene analogous DNA sequencing in pea (Pisum sativum L). Theor Appl Gen 101:241–247
Unver T, Wu Z, Sterck L, Turktas M, Lohaus R, Li Z, Yang M, He L, Deng T, Escalante FJ, Llorens C, Roig FJ, Parmaksiz I, Dundar E, Xie F, Zhang B, Ipek A, Uranbey S, Erayman M, Ilhan E, Badad O, Ghazal H, Lightfoot DA, Kasarla P, Colantonio V, Tombuloglu H, Hernandez P, Mete N, Cetin O, Montagu MV, Yang H, Gao Q, Dorado G, de Peer YV (2017) Genome of wild olive and the evolution of oil biosynthesis. Proc Natl Acad Sci U S A 114:E9413–E9422. https://doi.org/10.1073/pnas.1708621114
Van de Peer Y, Mizrachi E, Marchal K (2017) The evolutionary significance of polyploidy. Nat Rev Genet 18:411–424. https://doi.org/10.1038/nrg.2017.26
Wan H, Yuan W, Ye Q, Wang R, Ruan M, Li Z, Zhou G, Yao Z, Zhao J, Liu S, Yang Y (2012) Analysis of TIR- and non-TIR-NBS-LRR disease resistance gene analogous in pepper: characterization, genetic variation, functional divergence and expression patterns. BMC Genomics 13:502. https://doi.org/10.1186/1471-2164-13-502
Yang S, Zhang X, Yue JX, Tian D, Chen JQ (2008) Recent duplications dominate NBS-encoding gene expansion in two woody species. Mol Gen Genomics 280:187–198. https://doi.org/10.1007/s00438-008-0355-0
Yoshimura S, Yamanouchi U, Katayose Y, Toki S, Wang ZX, Kono I, Kurata N, Yano M, Iwata N, Sasaki T (1998) Expression of Xa1 a bacterial blight-resistance gene in rice is induced by bacterial inoculation. Proc Natl Acad Sci U S A 95:1663–1668. https://doi.org/10.1073/pnas.95.4.1663
Yu J, Tehrim S, Zhang F, Tong C, Huang J, Cheng X, Dong C, Zhou Y, Qin R, Hua W, Liu S (2014) Genome-wide comparative analysis of NBS-encoding genes between Brassica species and Arabidopsis thaliana. 15:3. https://doi.org/10.1186/1471-2164-15-3
Zhang H, Miao H, Wang L, Qu L, Liu H, Wang Q, Yue M (2013) Genome sequencing of the important oilseed crop Sesamum indicum L. Genome Biol vol. 14 pg. 401. https://doi.org/10.1186/gb-2013-14-1-401
Zhao H, Wang X, Jia Y, Minkenberg B, Wheatley M, Fan J, Jia MH, Famoso A, Edwards JD, Wamishe Y, Valent B, Wang GL, Yang Y (2018) The rice blast resistance gene Ptr encodes an atypical protein required for broad-spectrum disease resistance. Nat Commun 9:2039. https://doi.org/10.1038/s41467-018-04369-4
Zhong Y, Cheng ZM (2016) A unique RPW8-encoding class of genes that originated in early land plants and evolved through domain fission, fusion, and duplication. Sci Rep 6:32923. https://doi.org/10.1038/srep32923
Zhong Y, Li Y, Huang K, Cheng ZM (2015) Species-specific duplications of NBS-encoding genes in Chinese chestnut (Castanea mollissima). Sci Rep 5:16638. https://doi.org/10.1038/srep16638
Zipfel C (2008) Pattern-recognition receptors in plant innate immunity. Curr Opin Immunol 20(1):10–16. https://doi.org/10.1016/j.coi.2007.11.003
Zotta Mota AP, Vidigal B, Danchin EGJ, Coiti Togawa R, Leal-Bertioli SCM, Bertioli DJ, Guerra Araujo AC, Miranda Brasileiro AC, Messenberg Guimaraes P (2018) Comparative root transcriptome of wild Arachis reveals NBS-LRR genes related to nematode resistance. BMC Plant Biol 18:159. https://doi.org/10.1186/s12870-018-1373-7
Acknowledgments
This work is part of a doctoral thesis prepared by Inchirah Bettaieb. We gratefully acknowledge Imed Messaoudi, Khaled Chatti, and Nabil Soltani (Higher Institute of Biotechnology of Monastir, Tunisia), for their logistic help.
Funding
This study was financially supported by the Ministry of Higher Education and Scientific Research (Tunisia).
Author information
Authors and Affiliations
Contributions
IB analyzed data using bioinformatics tools, discussed the results, and co-wrote the manuscript, under the guidance of DB. DB designed this study, guided IB in all analyses and discussions, and co-wrote the manuscript. Both authors have read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflicts of interests.
Data archiving statement
The present work is a computational and data-enabled research. All the data used are public and were clearly cited in the manuscript along with the computer codes that generated the findings, for computational reproducibility. The present study does not produce itself any raw data.
Additional information
Editorial Responsibility: G. G. Vendramin
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Highlights
• The first genome-wide identification of NBS family genes was performed in the olive tree (Olea europaea L. subsp. europaea var. europaea), resulting in a ratio of non-TIR-type members to whole-genome genes in the expected lines with most Asteridae sequenced genomes, with a remarkably smaller proportion of TIR-type genes in olive.
• Olive NBS proteins are highly diverse molecules that integrate a wide variety of atypical domains, suggesting a great functional flexibility of this gene family. We first report on Armadillo repeats and Myb-like DNA-binding domain,as inherent components of NBS-type R proteins in olive.
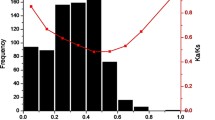
• Recent duplication events contributed to the expansion of NBS-encoding genes in the olive tree, generating novel resistance specificities while compensating for the longer generation time.
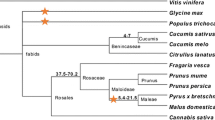
• Three orthologs to O. europaea NBS-encoding genes, from A. thaliana, O. sativa Indica, and O. sativa Japonica, were identified through a comparative phylogenetic approach and could be helpful to understand the function of the OeNBS genes in olive.
• EST-based expression analysis identified a pair of duplicated genes that could be associated with resistance to Verticillium dahliae and a gene possibly induced in olive roots upon colonization by the beneficial bacterium Pseudomonas fluorescens, a biological control agent against V. dahliae.
Rights and permissions
About this article
Cite this article
Bettaieb, I., Bouktila, D. Genome-wide analysis of NBS-encoding resistance genes in the Mediterranean olive tree (Olea europaea subsp. europaea var. europaea): insights into their molecular diversity, evolution and function. Tree Genetics & Genomes 16, 23 (2020). https://doi.org/10.1007/s11295-020-1415-9
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11295-020-1415-9