Abstract
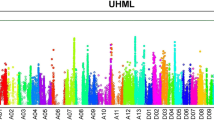
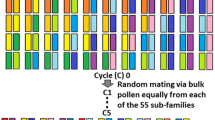
The analysis of association between genotypic markers and phenotypic traits allows identification of quantitative trait loci (QTLs), which can be applied through marker-assisted selection in breeding programs to improve the quality of crops. However, success in these applications depends primarily on the stability and dominance of the QTL. We previously identified a significant fiber length (FL) QTL on chromosome (Chr.) D11 based on the genome-wide association study (GWAS) of a multi-parent advanced generation inter-cross (MAGIC) population in upland cotton. In this report, we conducted mapping studies with two bi-parental populations to confirm the stability of the FL QTL on Chr. D11 and determine the magnitude of its effect on the fiber length phenotype. One of the F2 populations was developed from a cross between the longest fiber and the shortest fiber recombinant inbred lines (RILs) of the MAGIC population originally used for GWAS, whereas the second F2 population was created from a cross between two cotton lines Acala 1517–80 and JJ1145ne, which were not among the eleven MAGIC parental lines. The populations were grown in different environmental conditions. Genetic mapping of these populations confirmed the stability of the FL QTL on Chr. D11. The highest LOD scores of association with fiber length in both populations showed three SNP markers that resided within 360 kb of the QTL region on Chr. D11. Ten genes possessing non synonymous SNPs (nsSNPs) in their protein coding regions were identified in this region. RNAseq analysis detected activity in developing fiber tissue for seven of these candidate genes.




Similar content being viewed by others
References
Applequist WL, Cronn R, Wendel JF (2001) Comparative development of fiber in wild and cultivated cotton. Evol Dev 3(1):3–17
Basra AS, Malik C (1984) Development of the cotton fiber. Int Rev Cytol 89:65–113
Benjamini Y, Yekutieli D (2001) The control of the false discovery rate in multiple testing under dependency. Ann Stat 29:1165–1188
Drenkard E, Richter BG, Rozen S, Stutius LM, Angell NA, Mindrinos M, Cho RJ, Oefner PJ, Davis RW, Ausubel FM (2000) A simple procedure for the analysis of single nucleotide polymorphisms facilitates map-based cloning in Arabidopsis. Plant Physiol 124(4):1483–1492
Du Z, Zhou X, Ling Y, Zhang Z, Su Z (2010) agriGO: a GO analysis toolkit for the agricultural community. Nucleic Acids Res 38(suppl_2):W64–W70. https://doi.org/10.1093/nar/gkq310
Fang DD, Xiao J, Canci PC, Cantrell RG (2010) A new SNP haplotype associated with blue disease resistance gene in cotton (Gossypium hirsutum L.). Theor Appl Genet 120(5):943–953. https://doi.org/10.1007/s00122-009-1223-y
Fang DD, Jenkins JN, Deng DD, McCarty JC, Li P, Wu J (2014) Quantitative trait loci analysis of fiber quality traits using a random-mated recombinant inbred population in Upland cotton (Gossypium hirsutum L.). BMC genomics 15(1):397
Gilbert MK, Turley RB, Kim HJ, Li P, Thyssen G, Tang Y, Delhom CD, Naoumkina M, Fang DD (2013) Transcript profiling by microarray and marker analysis of the short cotton (Gossypium hirsutum L.) fiber mutant Ligon lintless-1 (Li1). BMC Genomics 14:403. https://doi.org/10.1186/1471-2164-14-403
Gilbert MK, Kim HJ, Tang Y, Naoumkina M, Fang DD (2014) Comparative transcriptome analysis of short fiber mutants Ligon-lintless 1 and 2 reveals common mechanisms pertinent to fiber elongation in cotton (Gossypium hirsutum L.). PLoS One 9(4):e95554. https://doi.org/10.1371/journal.pone.0095554
Haigler CH, Betancur L, Stiff MR, Tuttle JR (2012) Cotton fiber: a powerful single-cell model for cell wall and cellulose research. Front Plant Sci 3:104. https://doi.org/10.3389/fpls.2012.00104
Hinchliffe DJ, Meredith WR, Delhom CD, Thibodeaux DP, Fang DD (2011a) Elevated growing degree days influence transition stage timing during cotton fiber development resulting in increased fiber-bundle strength. Crop Sci 51(4):1683–1692
Hinchliffe DJ, Turley RB, Naoumkina M, Kim HJ, Tang Y, Yeater KM, Li P, Fang DD (2011b) A combined functional and structural genomics approach identified an EST-SSR marker with complete linkage to the Ligon lintless-2 genetic locus in cotton (Gossypium hirsutum L.). BMC Genomics 12:445. https://doi.org/10.1186/1471-2164-12-445
Islam MS, Thyssen GN, Jenkins JN, Zeng L, Delhom CD, McCarty JC, Deng DD, Hinchliffe DJ, Jones DC, Fang DD (2016) A MAGIC population-based genome-wide association study reveals functional association of GhRBB1_A07 gene with superior fiber quality in cotton. BMC Genomics 17(1):903. https://doi.org/10.1186/s12864-016-3249-2
Jenkins J, McCarty J, Gutierrez O, Hayes R, Bowman D, Watson C, Jones D (2008) Registration of RMUP-C5, a random mated population of upland cotton germplasm. J Plant Registrations 2(3):239–242
Kim HJ, Triplett BA (2001) Cotton fiber growth in planta and in vitro. Models for plant cell elongation and cell wall biogenesis. Plant Physiol 127(4):1361–1366
Kim D, Langmead B, Salzberg SL (2015) HISAT: a fast spliced aligner with low memory requirements. Nat Methods 12:357. https://doi.org/10.1038/nmeth.3317
Lee JJ, Woodward AW, Chen ZJ (2007) Gene expression changes and early events in cotton fibre development. Ann Bot 100(7):1391–1401
Lin Z-x, He D, X-l Z, Nie Y, Guo X, Feng C, Stewart JM (2005) Linkage map construction and mapping QTL for cotton fibre quality using SRAP, SSR and RAPD. Plant Breed 124(2):180–187
Ma J, Geng Y, Pei W, Wu M, Li X, Liu G, Li D, Ma Q, Zang X, Yu S (2018a) Genetic variation of dynamic fiber elongation and developmental quantitative trait locus mapping of fiber length in upland cotton (Gossypium hirsutum L.). BMC Genomics 19(1):882
Ma Z, He S, Wang X, Sun J, Zhang Y, Zhang G, Wu L, Li Z, Liu Z, Sun G (2018b) Resequencing a core collection of upland cotton identifies genomic variation and loci influencing fiber quality and yield. Nat Genet:1
Naoumkina M, Hinchliffe DJ, Turley RB, Bland JM, Fang DD (2013) Integrated metabolomics and genomics analysis provides new insights into the fiber elongation process in Ligon lintless-2 mutant cotton (Gossypium hirsutumL.). BMC Genomics 14(1):155. https://doi.org/10.1186/1471-2164-14-155
Naoumkina M, Thyssen G, Fang DD, Hinchliffe DJ, Florane C, Yeater KM, Page JT, Udall JA (2014) The Li2 mutation results in reduced subgenome expression bias in elongating fibers of allotetraploid cotton (Gossypium hirsutum L.). PLoS One 9(3):e90830. https://doi.org/10.1371/journal.pone.0090830
Naoumkina M, Thyssen GN, Fang DD (2015) RNA-seq analysis of short fiber mutants Ligon-lintless-1 (Li1) and - 2 (Li2) revealed important role of aquaporins in cotton (Gossypium hirsutum L.) fiber elongation. BMC Plant Biol 15:65. https://doi.org/10.1186/s12870-015-0454-0
Naoumkina M, Thyssen GN, Fang DD, Jenkins JN, McCarty JC, Florane CB (2019) Genetic and transcriptomic dissection of the fiber length trait from a cotton (Gossypium hirsutum L.) MAGIC population. BMC Genomics 20(1):112–114. https://doi.org/10.1186/s12864-019-5427-5
Paterson A, Saranga Y, Menz M, Jiang C-X, Wright R (2003) QTL analysis of genotype× environment interactions affecting cotton fiber quality. Theoret Appl Genetics 106(3):384–396
Pertea M, Pertea GM, Antonescu CM, Chang T-C, Mendell JT, Salzberg SL (2015) StringTie enables improved reconstruction of a transcriptome from RNA-seq reads. Nat Biotechnol 33:290–295. https://doi.org/10.1038/nbt.3122
Said JI, Knapka JA, Song M, Zhang J (2015a) Cotton QTLdb: a cotton QTL database for QTL analysis, visualization, and comparison between Gossypium hirsutum and G. hirsutum× G. barbadense populations. Mol Gen Genomics 290(4):1615–1625
Said JI, Song M, Wang H, Lin Z, Zhang X, Fang DD, Zhang J (2015b) A comparative meta-analysis of QTL between intraspecific Gossypium hirsutum and interspecific G.hirsutum × G.barbadense populations. Mol Gen Genomics 290(3):1003–1025. https://doi.org/10.1007/s00438-014-0963-9
Sun Z, Wang X, Liu Z, Gu Q, Zhang Y, Li Z, Ke H, Yang J, Wu J, Wu L (2017) Genome-wide association study discovered genetic variation and candidate genes of fibre quality traits in Gossypium hirsutum L. Plant Biotechnol J 15(8):982–996
Taliercio EW, Boykin D (2007) Analysis of gene expression in cotton fiber initials. BMC Plant Biol 7:22. https://doi.org/10.1186/1471-2229-7-22
Thaker VS, Saroop S, Vaishnav P, Singh Y (1989) Genotypic variations and influence of diurnal temperature on cotton fibre development. Field Crop Res 22(2):129–141
Thyssen GN, Fang DD, Turley RB, Florane C, Li P, Naoumkina M (2014) Next generation genetic mapping of the Ligon-lintless-2 (Li2) locus in upland cotton (Gossypium hirsutum L.). Theoret Appl Genetics 127(10):2183–2192
Thyssen G, Jenkins J, McCarty J, Zeng L, Campbell T, Delhom C, Islam M, Li P, Jones DC, Condon B, Fang DD (2019) Whole genome sequencing of a MAGIC population identified genomic loci and candidate genes for major fiber quality traits in upland cotton (Gossypium hirsutum L.). Theoret Appl Genetics 132(4):989–999. https://doi.org/10.1007/s00122-018-3254-8
Verkest A, Weinl C, Inzé D, De Veylder L, Schnittger A (2005) Switching the cell cycle. Kip-related proteins in plant cell cycle control. Plant Physiol 139(3):1099–1106. https://doi.org/10.1104/pp.105.069906
Wang R, Liu X, Liang S, Ge Q, Li Y, Shao J, Qi Y, An L, Yu F (2015) A subgroup of MATE transporter genes regulates hypocotyl cell elongation in Arabidopsis. J Exp Bot 66(20):6327–6343
Wang M, Tu L, Yuan D, Zhu D, Shen C, Li J, Liu F, Pei L, Wang P, Zhao G, Ye Z, Huang H, Yan F, Ma Y, Zhang L, Liu M, You J, Yang Y, Liu Z, Huang F, Li B, Qiu P, Zhang Q, Zhu L, Jin S, Yang X, Min L, Li G, Chen L-L, Zheng H, Lindsey K, Lin Z, Udall JA, Zhang X (2019) Reference genome sequences of two cultivated allotetraploid cottons, Gossypium hirsutum and Gossypium barbadense. Nat Genet 51(2):224–229. https://doi.org/10.1038/s41588-018-0282-x
Yan H, You Q, Tian T, Yi X, Liu Y, Du Z, Xu W, Su Z (2017) agriGO v2.0: a GO analysis toolkit for the agricultural community, 2017 update. Nucleic Acids Res 45(W1):W122–W129. https://doi.org/10.1093/nar/gkx382
Zhang T, Hu Y, Jiang W, Fang L, Guan X, Chen J, Zhang J, Saski CA, Scheffler BE, Stelly DM, Hulse-Kemp AM, Wan Q, Liu B, Liu C, Wang S, Pan M, Wang Y, Wang D, Ye W, Chang L, Zhang W, Song Q, Kirkbride RC, Chen X, Dennis E, Llewellyn DJ, Peterson DG, Thaxton P, Jones DC, Wang Q, Xu X, Zhang H, Wu H, Zhou L, Mei G, Chen S, Tian Y, Xiang D, Li X, Ding J, Zuo Q, Tao L, Liu Y, Li J, Lin Y, Hui Y, Cao Z, Cai C, Zhu X, Jiang Z, Zhou B, Guo W, Li R, Chen ZJ (2015) Sequencing of allotetraploid cotton (Gossypium hirsutum L. acc. TM-1) provides a resource for fiber improvement. Nat Biotechnol 33:531–537
Zhang J, Abdelraheem A, Flynn R (2019) Genetic gains of Acala 1517 cotton since 1926. Crop Sci 59(3):1052–1061. https://doi.org/10.2135/cropsci2018.11.0686
Funding
This research was funded by the United States Department of Agriculture-Agricultural Research Service CRIS project 6054–21000-018-00D.
Author information
Authors and Affiliations
Contributions
MN conceive the research and wrote the manuscript; MN, LZ, DDF, CBF, and PL conducted F2 populations studies; MW and GNT performed bioinformatics analysis; DDF conceived the association study and provided genomic sequence data; CD performed fiber testing; all authors read, edited, and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Mention of trade names or commercial products in this article is solely for the purpose of providing specific information and does not imply recommendation or endorsement by the US Department of Agriculture which is an equal opportunity provider and employer.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Naoumkina, M., Zeng, L., Fang, D.D. et al. Mapping and validation of a fiber length QTL on chromosome D11 using two independent F2 populations of upland cotton. Mol Breeding 40, 31 (2020). https://doi.org/10.1007/s11032-020-01111-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11032-020-01111-1