Abstract
Background
LBDs, as the plant-specific gene family, play essential roles in lateral organ development, plant regeneration, as well as abiotic stress and pathogen response. However, the number and characteristic of LBD genes in Pyscomitrella patens were still obscure.
Objective
This study was performed to identify the LBD family gene in moss and to determine the expression profiles of LBDs under the abiotic and pathogen stress.
Methods
Complete genome sequences and transcriptomes of P. patens were downloaded from the Ensembl plant database. The hidden Markov model-based profile of the conserved LOB domain was submitted as a query to identify all potential LOB domain sequences with HMMER software. Expression profiles of PpLBDs were obtained based on the GEO public database and qRT-PCR analysis.
Results
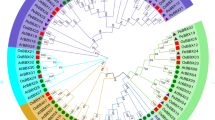
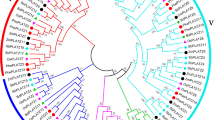
In this study, a total of 31 LBDs were identified in the P. patens genome, divided into two classes based on the presence of the leucine zipper-like coiled-coil motif. A phylogenetic relationship was obtained between 31 proteins from P. patens and 43 proteins from the Arabidopsis thaliana genome, providing insights into their conserved and potential functions. Furthermore, the exon–intron organization of each PpLBD were analyzed. All PpLBD contain the conserved DNA binding motif (CX2CX6CX3C zinc finger-like motif), and were predicted to be located in cell nuclear. The 31 PpLBD genes were unevenly assigned to 18 out of 27 chromosomes based on the physical positions. Among these genes, PpLBD27 was not only remarkably highest expressed in desiccation, but also a susceptible gene to pathogens through jasmonic acid-mediated signaling pathway. Most of PpLBDs were up-regulated with the treatment of mannitol. These results showed they were differentially induced and their potential functions in the environmental stimulus of the early terrestrial colonizers.
Conclusion
Despite significant differences in the life cycle in P. patens and flowering plants, their functions involved in abiotic and biotic stress-regulated by LBDs have been identified and appear to be conserved in the two lineages. These results provided a comprehensive analysis of PpLBDs and paved insights into studies aimed at a better understanding of PpLBDs.





Similar content being viewed by others
References
Agudelo-Romero P, Erban A, Rego C, Carbonell-Bejerano P, Nascimento T, Sousa L, Martinez-Zapater JM, Kopka J, Fortes AM (2015) Transcriptome and metabolome reprogramming in Vitis vinifera cv. Trincadeira berries upon infection with Botrytis cinerea. J Exp Bot 66:1769–1785
Ana C, Stefan GT, Juan Miguel GG, Javier T, Manuel T, Montserrat R (2005) Blast2GO: a universal tool for annotation, visualization and analysis in functional genomics research. Bioinformatics 21:3674–3676
Ariel FD, Diet A, Crespi M, Chan RL (2010) The LOB-like transcription factor Mt LBD1 controls Medicago truncatula root architecture under salt stress. Plant Signal Behav 5:1666–1668
Ba LJ, Kuang JF, Chen JY, Lu WJ (2016) MaJAZ1 attenuates the MaLBD5-mediated transcriptional activation of jasmonate biosynthesis gene MaAOC2 in regulating cold tolerance of banana fruit. J Agric Food Chem 64:738–745
Bell EM, Lin WC, Husbands AY, Yu L, Jaganatha V, Jablonska B, Mangeon A, Neff MM, Girke T, Springer PS (2012) Arabidopsis lateral organ boundaries negatively regulates brassinosteroid accumulation to limit growth in organ boundaries. Proc Natl Acad Sci USA 109:21146–21151
Cao H, Liu CY, Liu CX, Zhao YL, Xu RR (2016) Genomewide analysis of the lateral organ boundaries domain gene family in Vitis vinifera. J Genet 95:515–526
Chen WF, Wei XB, Rety S, Huang LY, Liu NN, Dou SX, Xi XG (2019) Structural analysis reveals a “molecular calipers” mechanism for a LATERAL ORGAN BOUNDARIES DOMAIN transcription factor protein from wheat. J Biol Chem 294:142–156
Ebeed HT, Stevenson SR, Cuming AC, Baker A (2018) Conserved and differential transcriptional responses of peroxisome associated pathways to drought, dehydration and ABA. J Exp Bot 69:4971–4985
Estornell LH, Landberg K, Cierlik I, Sundberg E (2018) SHI/STY genes affect pre- and post-meiotic anther processes in auxin sensing domains in Arabidopsis. Front Plant Sci 9:150–165
Fan M, Xu C, Xu K, Hu Y (2012) LATERAL ORGAN BOUNDARIES DOMAIN transcription factors direct callus formation in Arabidopsis regeneration. Cell Res 22:1169–1180
Gaudet P, Livstone MS, Lewis SE, Thomas PD (2011) Phylogenetic-based propagation of functional annotations within the Gene Ontology consortium. Brief Bioinform 12:449–462
Grimplet J, Pimentel D, Agudelo-Romero P, Martinez-Zapater JM, Fortes AM (2017) The LATERAL ORGAN BOUNDARIES Domain gene family in grapevine: genome-wide characterization and expression analyses during developmental processes and stress responses. Sci Rep 7:15968–15985
Hu Y, Zhang J, Jia H, Sosso D, Li T, Frommer WB, Yang B, White FF, Wang N, Jones JB (2014) Lateral organ boundaries 1 is a disease susceptibility gene for citrus bacterial canker disease. Proc Natl Acad Sci USA 111:E521–E529
Husbands A, Bell EM, Shuai B, Smith HM, Springer PS (2007) LATERAL ORGAN BOUNDARIES defines a new family of DNA-binding transcription factors and can interact with specific bHLH proteins. Nucleic Acids Res 35:6663–6671
Iwakawa H, Ueno Y, Semiarti E, Onouchi H, Kojima S, Tsukaya H, Hasebe M, Soma T, Ikezaki M, Machida C (2002) The ASYMMETRIC LEAVES2 gene of Arabidopsis thaliana, required for formation of a symmetric flat leaf lamina, encodes a member of a novel family of proteins characterized by cysteine repeats and a leucine zipper. Plant Cell Physiol 43:467–478
Kamisugi Y, Schlink K, Rensing SA, Schween G, von Stackelberg M, Cuming AC, Reski R, Cove DJ (2006) The mechanism of gene targeting in Physcomitrella patens: homologous recombination, concatenation and multiple integration. Nucleic Acids Res 34:6205–6214
Kaufman L, Rousseeuw PJ (2009) Finding groups in data: an introduction to cluster analysis. Wiley, Oxford
Kim MJ, Kim M, Lee MR, Park SK, Kim J (2015) LATERAL ORGAN BOUNDARIES DOMAIN (LBD)10 interacts with SIDECAR POLLEN/LBD27 to control pollen development in Arabidopsis. Plant J 81:794–809
Lee HW, Cho C, Kim J (2015) Lateral organ boundaries Domain16 and 18 act downstream of the AUXIN1 and LIKE-AUXIN3 auxin influx carriers to control lateral root development in Arabidopsis. Plant Physiol 168:1792–1806
Lewis MW, Nathalie B, Kayley H, Yadanar H, Angela H, Héctor C, Sarah H (2014) Gene regulatory interactions at lateral organ boundaries in maize. Development 141:4590–4597
Lima JE, Kojima S, Takahashi H, von Wiren N (2010) Ammonium triggers lateral root branching in Arabidopsis in an AMMONIUM TRANSPORTER1;3-dependent manner. Plant Cell 22:3621–3633
Lin WC, Shuai B, Springer PS (2003) The Arabidopsis LATERAL ORGAN BOUNDARIES-domain gene ASYMMETRIC LEAVES2 functions in the repression of KNOX gene expression and in adaxial-abaxial patterning. Plant Cell 15:2241–2252
Lu Q, Shao F, Macmillan C, Wilson IW, van der Merwe K, Hussey SG, Myburg AA, Dong X, Qiu D (2018) Genomewide analysis of the lateral organ boundaries domain gene family in Eucalyptus grandis reveals members that differentially impact secondary growth. Plant Biotechnol J 16:124–136
Luo Y, Ma B, Zeng Q, Xiang Z, He N (2016) Identification and characterization of Lateral Organ Boundaries Domain genes in mulberry, Morus notabilis. Meta Gene 8:44–50
Majer C, Hochholdinger F (2011) Defining the boundaries: structure and function of LOB domain proteins. Trends Plant Sci 16:47–52
McConnell JR, Barton MK (1998) Leaf polarity and meristem formation in Arabidopsis. Development 125:2935–2942
Naito T, Yamashino T, Kiba T, Koizumi N, Kojima M, Sakakibara H, Mizuno T (2007) A link between cytokinin and ASL9 (ASYMMETRIC LEAVES 2 LIKE 9) that belongs to the AS2/LOB (LATERAL ORGAN BOUNDARIES) family genes in Arabidopsis thaliana. Biosci Biotechnol Biochem 71:1269–1278
Nishiyama T, Fujita T, Seki M, Nishide H, Uchiyama I, Kamiya A, Carninci P, Hayashizaki Y, Shinozaki K, Kohara Y (2003) Comparative genomics of Physcomitrella patens gametophytic transcriptome and arabidopsis thaliana: implication for land plant evolution. Proc Natl Acad Sci USA 100:8007–8012
Ren C, Han C, Peng W, Huang Y, Peng Z, Xiong X, Zhu Q, Gao B, Xie D (2009) A leaky mutation in DWARF4 reveals an antagonistic role of brassinosteroid in the inhibition of root growth by jasmonate in Arabidopsis. Plant Physiol 151:1412–1420
Shuai B, Reynaga-Pena CG, Springer PS (2002) The lateral organ boundaries gene defines a novel, plant-specific gene family. Plant Physiol 129:747–761
Stevenson SR, Kamisugi Y, Chi HT, Schmutz J, Jenkins JW, Grimwood J, Muchero W, Tuskan GA, Rensing SA, Lang D (2016) Genetic analysis of Physcomitrella patens identifies ABSCISIC ACID NON-RESPONSIVE, a regulator of ABA responses unique to basal land plants and required for desiccation tolerance. Plant Cell 28:1310–1327
Sugita M, Ichinose M, Ide M, Sugita C (2013) Architecture of the PPR gene family in the moss Physcomitrella patens. RNA Biol 10:1439–1445
Sun SB, Song JP, Meng LS (2012) ASYMMETRIC LEAVES2 gene, a member of LOB/AS2 family of Arabidopsis thaliana, causes an abaxializing leaves in transgenic cockscomb. Mol Biol Rep 39:4927–4935
Thatcher LF, Kazan K, Manners JM (2012a) Lateral organ boundaries domain transcription factors: new roles in plant defense. Plant Signal Behav 7:1702–1704
Thatcher LF, Powell JJ, Aitken EA, Kazan K, Manners JM (2012b) The lateral organ boundaries domain transcription factor LBD20 functions in Fusarium wilt Susceptibility and jasmonate signaling in Arabidopsis. Plant Physiol 160:407–418
Wang Y, Secco D, Poirier Y (2008) Characterization of the PHO1 gene family and the responses to phosphate deficiency of Physcomitrella patens. Plant Physiol 146:646–656
Wang S, Bai Y, Shen C, Wu Y, Zhang S, Jiang D, Guilfoyle TJ, Chen M, Qi Y (2010) Auxin-related gene families in abiotic stress response in Sorghum bicolor. Funct Integr Genom 10:533–546
Xiao L, Yobi A, Koster KL, He Y, Oliver MJ (2018) Desiccation tolerance in Physcomitrella patens: rate of dehydration and the involvement of endogenous abscisic acid (ABA). Plant Cell Environ 41:275–284
Xu C, Luo F, Hochholdinger F (2016) LOB doma in proteins: beyond lateral organ boundaries. Trends Plant Sci 21:159–167
Yang Y, Yu X, Wu P (2006) Comparison and evolution analysis of two rice subspecies LATERAL ORGAN BOUNDARIES domain gene family and their evolutionary characterization from Arabidopsis. Mol Phylogenet Evol 39:248–262
Yang T, Fang GY, He H, Chen J (2016) Genome-wide identification, evolutionary analysis and expression profiles of LATERAL ORGAN BOUNDARIES DOMAIN gene family in Lotus japonicus and Medicago truncatula. PLoS One 11:e0161901
Zhang YM, Zhang SZ, Zheng CC (2014) Genomewide analysis of LATERAL ORGAN BOUNDARIES Domain gene family in Zea mays. J Genet 93:79–91
Zhong CM, Xu H, Ye ST, Wang SY, Li LF, Zhang SC, Wang XJ (2015) Gibberellic acid-stimulated Arabidopsis6 serves as an integrator of gibberellin, abscisic acid, and glucose signaling during seed germination in Arabidopsis. Plant Physiol 169:2288–2303
Acknowledgements
This work was supported by grants from the National Natural Science Foundation of China (Grant nos. 31660554 and 31660046). The talent platform of Qiankehe ([2017] 5726-49 and [2017] 5726-50).
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Huang, X., Yan, H., Liu, Y. et al. Genome-wide analysis of LATERAL ORGAN BOUNDARIES DOMAIN-in Physcomitrella patens and stress responses. Genes Genom 42, 651–662 (2020). https://doi.org/10.1007/s13258-020-00931-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13258-020-00931-x