Abstract
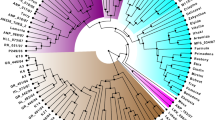
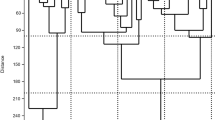
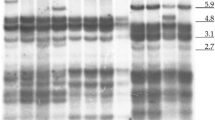
Genetic erosion, mainly caused by the replacement of local landraces with high yielding or exotic varieties is causing loss of agrobiodiversity. Landraces of horticultural species, grown by small producers, represent today an important value in the preservation of agrobiodiversity. Umbria, a region of central Italy, is characterized by agricultural systems linked to tradition and cultivation of local landraces. In this study, the genetic profile of some traditional Umbrian tomato landraces was characterized for the first time, and the landraces uniqueness was evaluated by comparison with commercial varieties. One-hundred and twenty-one plants provided by local farmers and seed companies, represented by local and commercial varieties were analyzed using 19 SSRs markers. A total of 60 alleles were found with moderate levels of diversity. The mean number of alleles per locus was 3.158 and the average polymorphism information content was 0.38. Unweighted UPGMA clustered the accessions into four groups. The gene pool of Umbrian landraces seems to be highly differentiated compared to commercial varieties, with landraces showing a genetic distinctiveness. Furthermore, an identifying fingerprinting code of each tomato landrace was generated and an innovative method for varietal identification based on the ‘QR code’ was proposed. The results obtained in this study will be useful for a better management, conservation and propagation of tomato genetic resources in Umbria region.




Similar content being viewed by others
References
Akkaya MS, Bhagwat AA, Cregan PB (1992) Length polymorphisms of simple sequence repeat DNA in soybean. Genetics 132:1131–1139
Barcaccia G, Lucchin M, Cassandro M (2016) DNA barcoding as a molecular tool to track down mislabeling and food piracy. Diversity 8:2. https://doi.org/10.3390/d8010002
Bergougnoux V (2014) The history of tomato: from domestication to biopharming. Biotechnol Adv 32:170–189. https://doi.org/10.1016/j.biotechadv.2013.11.003
Board on Agriculture of the National Research Council (1993) pp 23–25
Bonneuil C, Goffaux R, Bonnin I, Montalent P, Hamond C, Balfouriere F, Goldringerd I (2012) A new integrative indicator to assess crop genetic diversity. Ecol Ind 23:280–289. https://doi.org/10.1016/j.ecolind.2012.04.002
Caramante M, Rao R, Monti LM, Corrado G (2009) Discrimination of ‘San Marzano’ accessions: a comparison of minisatellite, CAPS and SSR markers in relation to morphological traits. Sci Hortic 120:560–564. https://doi.org/10.1016/j.scienta.2008.12.004
Casañas F, Simó J, Casals J, Prohens J (2017) Toward an evolved concept of landrace. Front Plant Sci 8:145. https://doi.org/10.3389/fpls.2017.00145
Conesa MÀ, Fullana-Pericàs M, Granell A, Galmés J (2020) Mediterranean long shelf-life landraces: an untapped genetic resource for tomato improvement. Front Plant Sci 10:1651. https://doi.org/10.3389/fpls.2019.01651
Corrado G, Piffanelli P, Caramante M, Coppola M, Rao R (2013) SNP genotyping reveals genetic diversity between cultivated landraces and contemporary varieties of tomato. BMC Genomics 14:835. https://doi.org/10.1186/1471-2164-14-835
Corrado G, Caramante M, Piffanelli P, Rao R (2014) Genetic diversity in Italian tomato landraces: Implications for the development of a core collection. Sci Hortic 168:138–144. https://doi.org/10.1016/j.scienta.2014.01.027
Cooke RJ (1995) Varietal identification of crop plants. New diagnostics in crop sciences. CAB International, Wallingford, pp 33–63
Denso Wave Incorporated (1994) QRcode. Aichi (JP)
Earl Dent A, Von Holdt M (2012) STRUCTURE HARVESTER: a website and program for visualizing STRUCTURE output and implementing the Evanno method. Conserv Genet Resour 4:359–361. https://doi.org/10.1007/s12686-011-9548-7
Esquinas-Alcazar J, Nuez F (1999) Situacion taxonomica, domesticacion y divusion del tomate. In: Mundi-Prensa M (ed) El cultivo del tomate. Mexico, Barcelona, pp 13–42
European Commission (2001) Biodiversity in development project. Strategic Approach for Integrating Biodiversity in Development Cooperation. European Commission, Brussels, Belgium, pp 1–82
European Commission (2002) Regulation (EC) No. 178/2002of the European Parliament and of the Council of 28January 2002 laying down the general principles and requirements of food law, establishing the European FoodSafety Authority and laying down procedures in matters of food safety OJ, L31, pp 1–24
European Commission (2009) Plant variety property rights, Regulation EC 874/2009
European Commission (2013) EU regulation 1151/2012 on quality schemes for agricultural products and foodstuffs. https://eur-lex.europa.eu/legal-content/EN/TXT/?uri=CELEX%3A32012R1151. Accessed 14 June 2019
European Commission (2019a) DOOR database. https://ec.europa.eu/agriculture/quality/door/list.html. Accessed 4 June 2019
European Commission (2019b) EU Plant variety database https://ec.europa.eu/food/plant/plant_propagation_material/plant_variety_catalogues_databases/search/public/index.cfm. Accessed 22 June 2019
Evanno G, Regnaut S, Goudet J (2005) Detecting the number of clusters of individuals using the software STRUCTURE: a simulation study. Mol Ecol 14:2611–2620. https://doi.org/10.1111/j.1365-294X.2005.02553.x
Excoffier L, Lischer HEL (2010) Arlequin suite ver 3.5: A new series of programs to perform population genetics analyses under Linux and Windows. Mol Ecol Resour 10:564–567. https://doi.org/10.1111/j.1755-0998.2010.02847.x
FAO (1996) Global plan of action for the conservation and sustainable utilization of plant genetic resources for food and agriculture. Leipzig, Germany.
FAO (1999a) Agricultural biodiversity, multifunctional character of agriculture and land conference, Background paper 1. Maastricht, Netherlands.
FAO (1999b) Women: users, preservers and managers of agrobiodiversity.
FAO (2012) The second report on the state of the world’s plant genetic resources. Food and Agriculture Organization of the United Nations, Rome
FAOSTAT (2017) FAO statistics division. https://faostat.fao.org/2017. Accessed 12 May 2019
Garcıa-Martınez S, Andreani L, Garcıa-Gusano M, Geuna F, Ruiz JJ (2006) Evaluation of amplified fragment length polymorphism and simple sequence repeats for tomato germplasm fingerprinting: utility for grouping closely related traditional cultivars. Genome 49:648–656. https://doi.org/10.1139/g06-016
Gonias ED, Ganopoulos I, Mellidou I (2019) Exploring genetic diversity of tomato (Solanum lycopersicum L.) germplasm of genebank collection employing SSR and SCAR markers. Genet Resour Crop Evol 66:1295–1309. https://doi.org/10.1007/s10722-019-00786-6
Gutiérrez JP, Royo LJ, Álvarez I, Goyache F (2005) MolKin v2.0: a computer program for genetic analysis of populations using molecular coancestry information. J Hered 96:718–721. https://doi.org/10.1093/jhered/esi118
Guichoux E, Lagache L, Wagner S, Chaumeil P, Léger P, Lepais O, Lepoittevin C, Malausa T, Revardel E, Salin F, Petit RJ (2011) Current trends in microsatellite genotyping. Mol Ecol Resour 11:591–611. https://doi.org/10.1111/j.1755-0998.2011.03014.x
Heras J, Domínguez C, Mata E (2015) A survey of tools for analyzing DNA fingerprints. Brief Bioinform 17(6):903–911. https://doi.org/10.1093/bib/bbv016
Kalinowski ST (2005) HP-Rare: a computer program for performing rarefaction on measures of allelic diversity. Mol Ecol Notes 5:187–189. https://doi.org/10.1111/j.1471-8286.2004.00845.x
Karihaloo JL (2015) DNA fingerprinting techniques for plant identification. In: Bahadur B, Venkatrajam M, Sahijram L, Krishnamurthy K (eds) Plant biology and biotechnology. Springer, New Delhi. https://doi.org/10.1007/978-81-322-2283-5_9
Kopelman NM, Mayzel J, Jakobsson M, Rosenberg NA, Mayrose I (2015) CLUMPAK: a program for identifying clustering modes and packaging population structure inferences across K. Mol Ecol Resour 15(5):1179–1191. https://doi.org/10.1111/1755-0998.12387
Liu S, Liu H, Wu A, Hou YL, An Y, Wei C (2017) Construction of fingerprinting for tea plant (Camellia sinensis) accessions using new genomic SSR markers. Mol Breed 37:1–14. https://doi.org/10.1007/s11032-017-0692-y
Mathur PN (2011) Assessing the threat of genetic erosion. In: Guarino L (ed) Collecting plant diversity: technical guidelines. Bioversity International, Rome, pp 1–7
Mazzucato A, Papa R, Bitocchi E (2008) Genetic diversity, structure and marker-trait associations in a collection of Italian tomato (Solanum lycopersicum L.) landraces. Theor Appl Genet 116:657. https://doi.org/10.1007/s00122-007-0699-6
Mazzucato A, Ficcadenti N, Caioni M, Mosconi P, Piccinini E, Rami Reddy Sanampudi V, Sestili S, Ferrari V (2010) Genetic diversity and distinctiveness in tomato (Solanum lycopersicum L.) landraces: the Italian case study of ‘A pera Abruzzese’. Sci Hortic 125:55–62. https://doi.org/10.1016/j.scienta.2010.02.021
Meirmans PG, Van Tienderen PH (2004) GENOTYPE and GENODIVE: two programs for the analysis of genetic diversity of asexual organisms. Mol Ecol Notes 4:792–794. https://doi.org/10.1111/j.1471-8286.2004.00770.x
MIPAAFT (2015) D.M. n.10803, 26/05/2015. Modalità operative inerenti la procedura informatica per l'iscrizione di varietà vegetali. https://www.politicheagricole.it/flex/cm/pages/ServeBLOB.php/L/IT/IDPagina/8771. Accessed 15 June 2019
Negri V, Maxted N, Veteläinen M (2009) European landrace conservation: an introduction. In: European landraces: on farm conservation, management and use: biodiversity technical bulletin no. 15. European Cooperative Programme for Plant Genetic Resources, Rome
Nei M (1973) Analysis of gene diversity in subdivided populations. Proc Nat Acad Sci 70(12):3321–3323. https://doi.org/10.1073/pnas.70.12.3321
Peakall R, Smouse PE (2012) GenAlEx 6.5: genetic analysis in Excel. population genetic software for teaching and research-an update. Bioinformatics 28:2537–2539. https://doi.org/10.1093/bioinformatics/bts460
Petropoulos Spyridon A, Barros L, Ferreira CFR (2019) Rediscovering local landraces: shaping horticulture for the future. Front Plant Sci 10:126. https://doi.org/10.3389/fpls.2019.00126
Porras-Hurtado L, Ruiz Y, Santos C, Phillips C, Carracedo A, Lareu MV (2013) An overview of STRUCTURE: applications, parameter settings, and supporting software. Front Genet 4:98. https://doi.org/10.3389/fgene.2013.00098
Powell W, Machray GC, Provan J (1996) Polymorphism revealed by simple sequence repeats. TrendsPlant Sci 1:215–222. https://doi.org/10.1016/1360-1385(96)86898-1
Pritchard JK, Stephens M, Donnelly P (2000) Inference of population structure using multilocus genotype data. Genetics 155(2):945–959
Rambaut A (2011) FigTree v1.4.4, Institute of Evolutionary Biology, University of Edinburgh.
Rick CM (1976) Tomato family Solanaceae. In: NW Simmonds (ed) Evolution of crop plants. Longman Publications, Harlow, pp 268–273
Rodríguez GR, Muños S, Anderson C, Sim S-C, Michel A, Causse M, Gardener BBM, Francis D, van der Knaap E (2011) Distribution of SUN, OVATE, LC, and FAS in the tomato germplasm and the relationship to fruit shape diversity. Plant Physiol 156:275–285. https://doi.org/10.1104/pp.110.167577
Savo Sardaro MLS, Marmiroli M, Maestri E, Marmiroli N (2013) Genetic characterization of Italian tomato varieties and their traceability in tomato food products. Food Sci Nutr 1(1):54–62. https://doi.org/10.1002/fsn3.8
Scarano D, Rao R, Masi P, Corrado G (2015) SSR fingerprint reveals mislabeling in commercial processed tomato products. Food Control 51:397–401. https://doi.org/10.1016/j.foodcont.2014.12.006
Selkoe KA, Toonen RJ (2006) Microsatellites for ecologists: a practical guide to using and evaluating microsatellite markers. Ecol Lett 9:615–629. https://doi.org/10.1111/j.1461-0248.2006.00889.x
Smulders MJM, Bredemeijer G, Rus-Kortekaas W, Arens P, Vosman B (1997) Use of short microsatellites from database sequences to generate polymorphisms among Lycopersicon esculentum cultivars and accessions of other Lycopersicon species. Theor Appl Genet 97:264–272. https://doi.org/10.1007/s001220050409
Sneath PH, Sokal RR (1973) Numerical taxonomy: the principles and practice of numerical classification, 1st edn. W. H. Freeman, San Francisco
Soressi GP (1969) Il, pomodoro edn. Agricole, Bologna
Takezaki N, Nei M, Tamura K (2010) POPTREE2: Software for constructing population trees from allele frequency data and computing other population statistics with Windows-interface. Mol Biol Evol 27:747–752. https://doi.org/10.1093/molbev/msp312
Valentini P, Galimberti A, Mezzasalma V, De Mattia F, Casiraghi M, Labra M, Pompa PP (2017) DNA barcoding meets nanotechnology: development of a universal colorimetric test for food authentication. Angew Chem Int 56:8094–8098. https://doi.org/10.1002/anie.201702120
Villa TC, Maxted N, Scholten MA, Ford-Lloyd BV (2005) Defining and identifying crop landraces. Plant Genet Res 3:373–384. https://doi.org/10.1079/PGR200591
Weir BS (1990) Genetic data analysis: methods for discrete population genetics data. Sinauer Associates, Sunderland
Weising K, Nybom H, Wolff K, Kahl G (2005) DNA fingerprinting in plants: principles, methods, and applications, 2nd edn. Taylor & Francis Group, Milton Park
Acknowledgements
We wish to acknowledge Dr. Livia Polegri for her advice in the sampling design and all the local farmers, specifically Azienda Melagioni, Antria Maggiore, Azienda Microcosmo and Da Matteo who provided samples and helped in their collection.
Funding
This study was funded by the National Research Program “Sviluppo Rurale per l’Umbria 2014–2020-Misura 16-Sottomisura 16.2”, Project title ‘Pomo d’Umbria’.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Castellana, S., Ranzino, L., Beritognolo, I. et al. Genetic characterization and molecular fingerprint of traditional Umbrian tomato (Solanum lycopersicum L.) landraces through SSR markers and application for varietal identification. Genet Resour Crop Evol 67, 1807–1820 (2020). https://doi.org/10.1007/s10722-020-00942-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10722-020-00942-3