Abstract
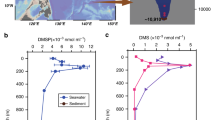
Microbial production and catabolism of dimethylsulfoniopropionate (DMSP), generating the climatically active gases dimethyl sulfide (DMS) and methanethiol (MeSH), have key roles in global carbon and sulfur cycling, chemotaxis, and atmospheric chemistry. Microorganisms in the sea surface microlayer (SML), the interface between seawater and atmosphere, likely play an important role in the generation of DMS and MeSH and their exchange to the atmosphere, but little is known about these SML microorganisms. Here, we investigated the differences between bacterial community structure and the distribution and transcription profiles of the key bacterial DMSP synthesis (dsyB and mmtN) and catabolic (dmdA and dddP) genes in East China Sea SML and subsurface seawater (SSW) samples. Per equivalent volume, bacteria were far more abundant (~ 7.5-fold) in SML than SSW, as were those genera predicted to produce DMSP. Indeed, dsyB (~ 7-fold) and mmtN (~ 4-fold), robust reporters for bacterial DMSP production, were also far more abundant in SML than SSW. In addition, the SML had higher dsyB transcripts (~ 3-fold) than SSW samples, which may contribute to the significantly higher DMSP level observed in SML compared with SSW. Furthermore, the abundance of bacteria with dmdA and their transcription were higher in SML than SSW samples. Bacteria with dddP and transcripts were also prominent, but less than dmdA and presented at similar levels in both layers. These data indicate that the SML might be an important hotspot for bacterial DMSP production as well as generating the climatically active gases DMS and MeSH, a portion of which are likely transferred to the atmosphere.





Similar content being viewed by others
Data Availability
The raw reads of the high throughput sequencing were deposited into the NCBI Sequence Read Archive (SRA) database with accession number SRP174872 under the BioProject PRJNA511511. The partial sequences of the dsyB gene from clone libraries were available in the GenBank database with accession numbers MN232008 to MN232099.
References
Lovelock, J.E, Maggs, R.J, Rasmussen R.A.: Atmospheric dimethyl sulphide and the natural sulphur cycle. Nature. 237, 452–453 (1972). https://doi.org/10.1038/237452a0
Nevitt, G.A, Bonadonna, F.: Sensitivity to dimethyl sulphide suggests a mechanism for olfactory navigation by seabirds. Biol. Lett. 1, 303–305 (2005). https://doi.org/10.1098/rsbl.2005.0350
Steinke, M., Stefels, J., Stamhuis, E.: Dimethyl sulfide triggers search behavior in copepods. Limnol. Oceanogr. 51, 1925–1930 (2006). https://doi.org/10.4319/lo.2006.51.4.1925
Charlson, R.J, Lovelock, J.E, Andreae, M.O, Warren, S.G.: Oceanic phytoplankton, atmospheric sulphur, cloud albedo and climate. Nature 326, 655–661 (1987). https://doi.org/10.1038/326655a0
Vallina, SM., Simo, R.: Strong relationship between DMS and the solar radiation dose over the global surface ocean. Science. 315, 506–508 (2007). https://doi.org/10.1126/science.1133680
Wolfe, G.V., Steinke, M., Kirst, G.O.: Grazing-activated chemical defence in a unicellular marine alga. Nature. 387, 894–897 (1997). https://doi.org/10.1038/43168
Sunda, W., Kieber, D.J., Kiene, R.P., Huntsman, S.: An antioxidant function for DMSP and DMS in marine algae. Nature. 418, 317–320 (2002). https://doi.org/10.1038/nature00851
Seymour J.R., Simo, R., Ahmed, T., Stocker, R.: Chemoattraction to dimethylsulfoniopropionate throughout the marine microbial food web. Science. 329, 342–345 (2010). https://doi.org/10.1126/science.1188418
Kellogg W.W., Cadle R.D., Allen, E.R., Lazrus, A.L., Martell, E.A.: The sulfur cycle. Science. 175, 587–596 (1972). https://doi.org/10.1126/science.175.4022.587
Zhang X.H., Liu, J. Liu, J., Yang, G., Xue, C.X., Curson, A.R.J., Todd, J.D.: Biogenic production of DMSP and its degradation to DMS-their roles in the global sulfur cycle. Sci. China Life Sci. 62, 1296–1319 (2019). https://doi.org/10.1007/s11427-018-9524-y
Curson, A.R., Todd, J.D., Sullivan, M.J., Johnston, A.W.: Catabolism of dimethylsulphoniopropionate: microorganisms, enzymes and genes. Nat. Rev. Microbiol. 9, 849–859 (2011). https://doi.org/10.1038/nrmicro2653
Ackman, R.G., Tocher, C.S., McLachlan, J.: Occurrence of dimethyl-β-propiothetin in marine phytoplankton. J Fish Res Board. 23, 357–364 (1966). https://doi.org/10.1139/f66-030
Otte, M.L., Wilson, G., Morris, J.T., Moran, B.M.: Dimethylsulphoniopropionate (DMSP) and related compounds in higher plants. J. Exp. Bot. 55, 1919–1925 (2004). https://doi.org/10.1093/jxb/erh178
Van Alstyne K.L., Puglisi, M.P.: DMSP in marine macroalgae and macroinvertebrates: distribution, function, and ecological impacts. Aquat. Sci. 69, 394–402 (2007). https://doi.org/10.1007/s00027-007-0888-z
Raina, J.B., Tapiolas, D.M., Foret, S., Lutz, A., Abrego, D., Ceh J., Seneca, F.O., Clode, P.L., Bourne, D.G., Willis, B.L., Motti, C.A.: DMSP biosynthesis by an animal and its role in coral thermal stress response. Nature. 502, 677–680 (2013). https://doi.org/10.1038/nature12677
Dickson, D.M., Jones, R.G., Davenport, J.: Steady state osmotic adaptation in Ulva lactuca. Planta. 150, 158–165 (1980). https://doi.org/10.1007/BF00582360
Curson, A.R.J., Williams, B.T., Pinchbeck, B.J., Sims, L.P., Martinez, A.B., Rivera, P.P.L., Kumaresan, D., Mercade, E., Spurgin, L.G., Carrion, O., Moxon, S., Cattolico, R.A., Kuzhiumparambil, U., Guagliardo P., Clode, P.L., Raina, J.B., Todd, J.D.: DSYB catalyses the key step of dimethylsulfoniopropionate biosynthesis in many phytoplankton. Nat. Microbiol. 3, 430–439 (2018). https://doi.org/10.1038/s41564-018-0119-5
Kageyama, H., Tanaka, Y., Shibata, A., Waditee-Sirisattha, R., Takabe, T.: Dimethylsulfoniopropionate biosynthesis in a diatom Thalassiosira pseudonana: Identification of a gene encoding MTHB-methyltransferase. Arch. Biochem. Biophys. 645, 100–106 (2018). https://doi.org/10.1016/j.abb.2018.03.019
Curson, A.R., Liu, J., Bermejo Martinez, A., Green, R.T., Chan, Y., Carrion, O., Williams, B.T., Zhang, S.H., Yang, G.P., Bulman Page, P.C., Zhang, X.H., Todd, J.D.: Dimethylsulfoniopropionate biosynthesis in marine bacteria and identification of the key gene in this process. Nat. Microbiol. 2, 17009 (2017). https://doi.org/10.1038/nmicrobiol.2017.9
Williams, B.T., Cowles, K., Bermejo Martinez, A., Curson, A.R.J., Zheng, Y., Liu, J., Newton-Payne, S., Hind, A.J., Li, C.Y., Rivera, P.P.L., Carrion, O., Liu, J., Spurgin, L.G., Brearley, C.A., Mackenzie, B.W., Pinchbeck, B.J., Peng, M., Pratscher, J., Zhang, X.H., Zhang, Y.Z., Murrell, J.C., Todd, J.D.: Bacteria are important dimethylsulfoniopropionate producers in coastal sediments. Nat. Microbiol. 4, 1815–1825 (2019). https://doi.org/10.1038/s41564-019-0527-1
Rhodes, D., Gage, D.A., Cooper, A., Hanson, A.D.: S-Methylmethionine conversion to dimethylsulfoniopropionate: Evidence for an unusual transamination reaction. Plant Physiol. 115, 1541–1548 (1997). https://doi.org/10.1104/pp.115.4.1541
Kocsis, M.G., Hanson, A.D.: Biochemical evidence for two novel enzymes in the biosynthesis of 3-dimethylsulfoniopropionate in Spartina alterniflora. Plant Physiol. 123, 1153–1161 (2000). https://doi.org/10.1104/pp.123.3.1153
Tripp H.J., Kitner, J.B., Schwalbach M.S., Dacey J.W., Wilhelm L.J., Giovannoni S.J.: SAR11 marine bacteria require exogenous reduced sulphur for growth. Nature. 452, 741–744 (2008). https://doi.org/10.1038/nature06776
Thume, K., Gebser, B., Chen, L., Meyer, N., Kieber, D.J., Pohnert, G.: The metabolite dimethylsulfoxonium propionate extends the marine organosulfur cycle. Nature. 563, 412–415 (2018). https://doi.org/10.1038/s41586-018-0675-0
Howard, E.C., Sun, S., Biers, E.J., Moran, M.A.: Abundant and diverse bacteria involved in DMSP degradation in marine surface waters. Environ. Microbiol. 10, 2397–2410 (2008). https://doi.org/10.1111/j.1462-2920.2008.01665.x
Varaljay, V.A., Gifford, S.M., Wilson, S.T., Sharma, S., Karl, D.M., Moran, M.A.: Bacterial dimethylsulfoniopropionate degradation genes in the oligotrophic north pacific subtropical gyre. Appl. Environ. Microbiol. 78, 2775–2782 (2012). https://doi.org/10.1128/AEM.07559-11
Cui, Y., Suzuki, S., Omori, Y., Wong, S.K., Ijichi, M., Kaneko, R., Kameyama, S., Tanimoto, H., Hamasaki, K.: Abundance and distribution of dimethylsulfoniopropionate degradation genes and the corresponding bacterial community structure at dimethyl sulfide hot spots in the tropical and subtropical pacific ocean. Appl. Environ. Microbiol. 81, 4184–4194 (2015). https://doi.org/10.1128/AEM.03873-14
Varaljay, V.A., Howard, E.C., Sun, S., Moran, M.A.: Deep sequencing of a dimethylsulfoniopropionate-degrading gene (dmdA) by using PCR primer pairs designed on the basis of marine metagenomic data. Appl. Environ. Microbiol. 76, 609–617 (2010). https://doi.org/10.1128/AEM.01258-09
Zeng, Y.X., Qiao, Z.Y., Yu, Y., Li, H.R., Luo, W.: Diversity of bacterial dimethylsulfoniopropionate degradation genes in surface seawater of Arctic Kongsfjorden. Sci. Rep. 6, 33031 (2016). https://doi.org/10.1038/srep33031
Landa, M., Burns, A.S., Durham, B.P., Esson, K., Nowinski, B., Sharma, S., Vorobev, A., Nielsen, T., Kiene, R.P., Moran, M.A.: Sulfur metabolites that facilitate oceanic phytoplankton-bacteria carbon flux. ISME J. 13, 2536–2550 (2019). https://doi.org/10.1038/s41396-019-0455-3
Alcolombri, U., Ben-Dor, S., Feldmesser, E., Levin, Y., Tawfik, D.S., Vardi, A.: Marine sulfur cycle. Identification of the algal dimethyl sulfide-releasing enzyme: A missing link in the marine sulfur cycle. Science. 348, 1466–1469 (2015). https://doi.org/10.1126/science.aab1586
Johnston, A.W. BIOGEOCHEMISTRY. Who can cleave DMSP? Science. 348, 1430–1431 (2015). https://doi.org/10.1126/science.aac5661
Sun, J., Todd, J.D., Thrash, J.C., Qian, Y., Qian, M.C., Temperton, B., Guo, J., Fowler, E.K., Aldrich, J.T., Nicora, C.D., Lipton, M.S., Smith, R.D., De Leenheer, P., Payne, S.H., Johnston, A.W., Davie-Martin, C.L., Halsey, K.H., Giovannoni, S.J.: The abundant marine bacterium Pelagibacter simultaneously catabolizes dimethylsulfoniopropionate to the gases dimethyl sulfide and methanethiol. Nat. Microbiol. 1, 16065 (2016). https://doi.org/10.1038/nmicrobiol.2016.65
Bullock, H.A., Luo, H., Whitman, W.B.: Evolution of dimethylsulfoniopropionate metabolism in marine phytoplankton and bacteria. Front. Microbiol. 8, 637 (2017). https://doi.org/10.3389/fmicb.2017.00637
Todd, J.D., Curson, A.R., Dupont, C.L., Nicholson, P., Johnston, A.W.: The dddP gene, encoding a novel enzyme that converts dimethylsulfoniopropionate into dimethyl sulfide, is widespread in ocean metagenomes and marine bacteria and also occurs in some Ascomycete fungi. Environ. Microbiol. 11, 1376–1385 (2009). https://doi.org/10.1111/j.1462-2920.2009.01864.x
Kirkwood, M., Todd, J.D., Rypien, K.L., Johnston, A.W.: The opportunistic coral pathogen Aspergillus sydowii contains dddP and makes dimethyl sulfide from dimethylsulfoniopropionate. ISME J. 4, 147–150 (2010). https://doi.org/10.1038/ismej.2009.102
Raina, J.B., Dinsdale, E.A., Willis, B.L., Bourne, D.G.: Do the organic sulfur compounds DMSP and DMS drive coral microbial associations? Trends Microbiol. 18, 101–108 (2010). https://doi.org/10.1016/j.tim.2009.12.002
Liu, J., Liu, J., Zhang, S.H., Liang, J., Lin, H., Song, D., Yang, G.P., Todd, J.D., Zhang, X.H.: Novel insights into bacterial dimethylsulfoniopropionate catabolism in the East China Sea. Front. Microbiol. 9, 3206 (2018). https://doi.org/10.3389/fmicb.2018.03206
Choi, D.H., Park, K.T., An, S.M., Lee. K., Cho, J.C., Lee, J.H., Kim, D., Jeon, D., Noh, J.H.: Pyrosequencing revealed SAR116 clade as dominant dddP-containing bacteria in oligotrophic NW Pacific Ocean. PLoS One. 10, e0116271 (2015). https://doi.org/10.1371/journal.pone.0116271
Howard, E.C., Henriksen, J.R., Buchan, A., Reisch, C.R., Burgmann, H., Welsh, R., Ye, W., Gonzalez, J.M., Mace, K., Joye, S.B., Kiene, R.P., Whitman, W.B., Moran, M.A.: Bacterial taxa that limit sulfur flux from the ocean. Science. 314, 649–652 (2006). https://doi.org/10.1126/science.1130657
Kiene, R.P., Linn, L.J.: Distribution and turnover of dissolved DMSP and its relationship with bacterial production and dimethylsulfide in the Gulf of Mexico. Limnol. Oceanogr. 45, 849–861 (2000). https://doi.org/10.4319/lo.2000.45.4.0849
Liss, P.S., Duce, R.A:. The sea surface and global change. Cambridge University Press, Cambridge (1997)
Parker, B., Barsom, G.: Biological and chemical significance of surface microlayers in aquatic ecosystems. Bioscience. 20, 87–93 (1970). https://doi.org/10.2307/1294827
Hatcher, R.F., Parker, B.C.: Microbiological and chemical enrichment of freshwater-surface microlayers relative to the bulk-subsurface water. Can. J. Microbiol. 20, 1051–1057 (1974). https://doi.org/10.1139/m74-162
Garrett, W.D.: The organic chemical composition of the ocean surface. Deep-Sea Res. Oceanogr. Abstr. 14, 221–227 (1967). https://doi.org/10.1016/0011-7471(67)90007-1
Jarvis, N.L.: Adsorption of surface-active material at the sea-air interface. Limnol. Oceanogr. 12, 213–221 (1967). https://doi.org/10.4319/lo.1967.12.2.0213
Quinn, J.A., Otto, N.C.: Carbon dioxide exchange at the air-sea interface: flux augmentation by chemical reaction. J. Geophys. Res. 76, 1539–1549 (1971). https://doi.org/10.1029/JC076i006p01539
Quickenden, T.I., Barnes, G.T.: Evaporation through monolayers—theoretical treatment of the effect of chain length. J. Colloid Interface Sci. 67, 415–422 (1978). https://doi.org/10.1016/0021-9797(78)90230-8
Wu, M.H., Yang, X.X., Xu, G., Que, C.J., Ma, S.H., Tang, L.: Semivolatile organic compounds in surface microlayer and subsurface water of Dianshan Lake, Shanghai, China: implications for accumulation and interrelationship. Environ. Sci. Pollut. Res. Int. 24, 6572–6580 (2017). https://doi.org/10.1007/s11356-016-8308-3
Franklin, M.P., McDonald, I.R., Bourne, D.G., Owens, N.J., Upstill-Goddard, R.C., Murrell, J.C.: Bacterial diversity in the bacterioneuston (sea surface microlayer): The bacterioneuston through the looking glass. Environ. Microbiol. 7, 723–736 (2005). https://doi.org/10.1111/j.1462-2920.2004.00736.x
Zancker, B., Cunliffe, M., Engel, A.: Bacterial community composition in the sea surface microlayer off the Peruvian coast. Front. Microbiol. 9, 2699 (2018). https://doi.org/10.3389/fmicb.2018.02699
Azevedo JS, Araujo S, Oliveira CS, Correia A, Henriques I (2013) Analysis of antibiotic resistance in bacteria isolated from the surface microlayer and underlying water of an estuarine environment. Microb. Drug Resist. 19, 64–71. https://doi.org/10.1089/mdr.2012.0084
Yang, G.P., Levasseur, M., Michaud, S.: Scarratt, M.: Biogeochemistry of dimethylsulfide (DMS) and dimethylsulfoniopropionate (DMSP) in the surface microlayer and subsurface water of the Western North Atlantic during spring. Mar. Chem. 96, 315–329 (2005). https://doi.org/10.1016/j.marchem.2005.03.003
Yang, G.P., Tsunogai, S., Watanabe, S.: Biogeochemistry of dimethylsulfoniopropionate (DMSP) in the surface microlayer and subsurface seawater of Funka Bay, Japan. J Oceanogr. 61, 69–78 (2005). https://doi.org/10.1007/s10872-005-0020-8
Yang, G.P., Jing, W.W., Kang, Z.Q., Zhang, H.H., Song, G.S.: Spatial variations of dimethylsulfide and dimethylsulfoniopropionate in the surface microlayer and in the subsurface waters of the South China Sea during springtime. Mar. Environ. Res. 65, 85–97 (2008). https://doi.org/10.1016/j.marenvres.2007.09.002
Zhang, H.H., Yang, G.P., Zhu, T.: Distribution and cycling of dimethylsulfide (DMS) and dimethylsulfoniopropionate (DMSP) in the sea-surface microlayer of the Yellow Sea, China, in spring. Cont. Shelf Res. 28, 2417–2427 (2008). https://doi.org/10.1016/j.csr.2008.06.003
Fehon, W.C., Oliver, J.D.: Taxonomy and distribution of surface microlayer bacteria from two estuarine sites. Estuaries. 2, 194–197 (1979). https://doi.org/10.2307/1351735
Zhang, X., Lin, H., Wang, X., Austin, B. Significance of Vibrio species in the marine organic carbon cycle—a review. Science China Earth Sciences. 61, 1357–1368 (2018)
Kanukollu, S., Wemheuer, B., Herber, J., Billerbeck, S., Lucas, J., Daniel, R., Simon, M., Cypionka, H., Engelen, B.: Distinct compositions of free-living, particle-associated and benthic communities of the Roseobacter group in the North Sea. FEMS Microbiol Ecol. 92(1):fiv145 (2016). https://doi.org/10.1093/femsec/fiv145
Liang, J., Liu, J., Wang, X., Lin, H., Liu, J., Zhou, S., Sun, H., Zhang, XH.: Spatiotemporal dynamics of free-living and particle-associated Vibrio communities in the Northern Chinese marginal seas. Appl Environ Microbiol. (2019). https://doi.org/10.1128/AEM.00217-19
Park, B.S., Guo, R., Lim, W.-A., Ki, J.-S.: Importance of free-living and particle-associated bacteria for the growth of the harmful dinoflagellate Prorocentrum minimum: Evidence in culture stages. Mar Freshw Res. 69, 290–299 (2018)
Varaljay, V.A.: Quantitative analysis of bacterial DMSP-degrading gene diversity, abundance, and expression in marine surface water environments. University of Georgia Athens, GA (2012)
Yang, G.-P., Watanabe, S., Tsunogai, S.: Distribution and cycling of dimethylsulfide in surface microlayer and subsurface seawater. Mar. Chem. 76, 137–153 (2001)
Zhang, Y., Zhao, Z., Dai, M., Jiao, N., Herndl, G.J.: Drivers shaping the diversity and biogeography of total and active bacterial communities in the South China Sea. Mol. Ecol. 23, 2260–2274 (2014). https://doi.org/10.1111/mec.12739
Carpenter, J.H.: The accuracy of the Winkler method for dissolved oxygen analysis1. Limnol. Oceanogr. 10, 135–140 (1965). https://doi.org/10.4319/lo.1965.10.1.0135
Liu, J., Fu, B., Yang, H., Zhao, M., He, B., Zhang, X.H.: Phylogenetic shifts of bacterioplankton community composition along the Pearl Estuary: The potential impact of hypoxia and nutrients. Front. Microbiol. 6, 64 (2015). https://doi.org/10.3389/fmicb.2015.00064
Yang, G.-P., Zhang, H,-H., Zhou L.-M., Yang, J.: Temporal and spatial variations of dimethylsulfide (DMS) and dimethylsulfoniopropionate (DMSP) in the East China Sea and the Yellow Sea. Cont. Shelf Res. 31, 1325–1335 (2011). https://doi.org/10.1016/j.csr.2011.05.001
Tan, T.-T., Wu, X., Liu, C.-Y., Yang, G.-P. Distributions of dimethylsulfide and its related compounds in the Yangtze (Changjiang) River Estuary and its adjacent waters in early summer. Cont. Shelf Res. 146, 89–101 (2017). https://doi.org/10.1016/j.csr.2017.08.012
Yin, Q., Fu, B., Li, B., Shi, X., Inagaki, F., Zhang, XH.: Spatial variations in microbial community composition in surface seawater from the ultra-oligotrophic center to rim of the South Pacific Gyre. PLoS One. 8, e55148 (2013). https://doi.org/10.1371/journal.pone.0055148
Liu, J., Zheng, Y., Lin, H., Wang, X., Li, M., Liu, Y., Yu, M., Zhao, M., Pedentchouk, N., Lea-Smith, D.J., Todd, J.D., Magill, C.R., Zhang, W.J., Zhou, S., Song, D., Zhong, H., Xin, Y., Yu, M., Tian, J., Zhang, X.H.: Proliferation of hydrocarbon-degrading microbes at the bottom of the Mariana Trench. Microbiome. 7, 47 (2019). https://doi.org/10.1186/s40168-019-0652-3
Levine, N.M., Varaljay, V.A., Toole, D.A., Dacey, J.W., Doney, S.C., Moran, M.A.: Environmental, biochemical and genetic drivers of DMSP degradation and DMS production in the Sargasso Sea. Environ. Microbiol. 14, 1210–1223 (2012). https://doi.org/10.1111/j.1462-2920.2012.02700.x
Walters, W., Hyde, E.R., Berg-Lyons, D., Ackermann, G., Humphrey, G., Parada, A., Gilbert, J.A., Jansson, J.K., Caporaso, J.G., Fuhrman, J.A., Apprill, A., Knight, R.: Improved bacterial 16S rRNA gene (V4 and V4-5) and fungal internal transcribed spacer marker gene primers for microbial community surveys. mSystems. 1, e00009–e00015 (2016). https://doi.org/10.1128/mSystems.00009-15
Todd, J.D., Curson, A.R., Nikolaidou-Katsaraidou, N., Brearley, C.A., Watmough, N.J., Chan, Y., Page, P.C., Sun, L., Johnston, A.W.: Molecular dissection of bacterial acrylate catabolism--unexpected links with dimethylsulfoniopropionate catabolism and dimethyl sulfide production. Environ. Microbiol. 12, 327–343 (2010). https://doi.org/10.1111/j.1462-2920.2009.02071.x
Curson, A.R., Rogers, R., Todd, J.D., Brearley, C.A., Johnston, A.W.: Molecular genetic analysis of a dimethylsulfoniopropionate lyase that liberates the climate-changing gas dimethylsulfide in several marine alpha-proteobacteria and Rhodobacter sphaeroides. Environ. Microbiol. 10, 757–767 (2008). https://doi.org/10.1111/j.1462-2920.2007.01499.x
Oh HM, Kwon KK, Kang I, Kang SG, Lee JH, Kim SJ, Cho JC (2010) Complete genome sequence of “Candidatus Puniceispirillum marinum” IMCC1322, a representative of the SAR116 clade in the Alphaproteobacteria. J. Bacteriol. 192:3240–3241. https://doi.org/10.1128/JB.00347-10
Kanukollu, S., Voget, S., Pohlner, M., Vandieken, V., Petersen, J., Kyrpides, N.C., Woyke, T., Shapiro, N., Goker, M., Klenk, H.P., Cypionka, H., Engelen, B.: Genome sequence of Shimia str. SK013, a representative of the Roseobacter group isolated from marine sediment. Stand. Genomic Sci. 11, 25 (2016). https://doi.org/10.1186/s40793-016-0143-0
Zeng, Y.: Phylogenetic diversity of dimethylsulfoniopropionatede-pendent demethylase gene dmdA in distantly related bacteria isolated from Arctic and Antarctic marine environments. Acta Oceanol. Sin. 38, 64–71 (2019). https://doi.org/10.1007/s13131-019-1393-7
Engel, A., Bange, H.W., Cunliffe, M., Burrows, S.M., Friedrichs, G., Galgani, L., H.H., N.H., Johnson, M., Liss, P.S., Quinn, P.K., Schartau, M., Soloviev, A., Stolle, C., Upstill-Goddard, R.C., Van Pinxteren, M., Zäncker, B.: The ocean’s vital skin: toward an integrated understanding of the sea surface microlayer. Front. Mar. Sci. 4, 165 (2017). https://doi.org/10.3389/fmars.2017.00165
Hardy, J.T.: The sea surface microlayer: biology, chemistry and anthropogenic enrichment. Prog. Oceanogr. 11, 307–328 (1982). https://doi.org/10.1016/0079-6611(82)90001-5
Williams, P.M., Carlucci, A.F., Henrichs, S.M., Vleet, E.S.V., Horrigan, S.G., Reid, F.M.H., Robertson, K.J. Chemical and microbiological studies of sea-surface films in the Southern Gulf of California and off the West Coast of Baja California. Mar. Chem. 19, 17–98 (1986). https://doi.org/10.1016/0304-4203(86)90033-2
Aller, J.Y., Kuznetsova, M.R., Jahns, C.J., Kemp, P.F.: The sea surface microlayer as a source of viral and bacterial enrichment in marine aerosols. J. Aerosol Sci. 36, 801–812 (2005). https://doi.org/10.1016/j.jaerosci.2004.10.012
Song, Y.K., Hong, SH, Jang, M., Kang, J.H., Kwon, O.Y., Han, GM., Shim, W.J.: Large accumulation of micro-sized synthetic polymer particles in the sea surface microlayer. Environ. Sci. Technol. 48, 9014–9021 (2014). https://doi.org/10.1021/es501757s
Yoch, D.C.: Dimethylsulfoniopropionate: its sources, role in the marine food web, and biological degradation to dimethylsulfide. Appl. Environ. Microbiol. 68, 5804–5815 (2002). https://doi.org/10.1128/aem.68.12.5804-5815.2002
Dahllof, I., Baillie, H., Kjelleberg, S.: rpoB-based microbial community analysis avoids limitations inherent in 16S rRNA gene intraspecies heterogeneity. Appl. Environ. Microbiol. 66, 3376–3380 (2000). https://doi.org/10.1128/aem.66.8.3376-3380.2000
Cunliffe, M., Murrell, J.C.: The sea-surface microlayer is a gelatinous biofilm. ISME J. 3, 1001–1003 (2009). https://doi.org/10.1038/ismej.2009.69
Agogue, H., Casamayor, E.O., Bourrain, M., Obernosterer, I., Joux, F., Herndl, G.J., Lebaron, P.: A survey on bacteria inhabiting the sea surface microlayer of coastal ecosystems. FEMS Microbiol. Ecol. 54, 269–280 (2005). https://doi.org/10.1016/j.femsec.2005.04.002
Cunliffe, M., Harrison, E., Salter, M., Schäfer, H., Upstill-Goddard, R.C., Murrell, J.C.: Comparison and validation of sampling strategies for the molecular microbial analysis of surface microlayers. Aquat. Microb. Ecol. 57, 69–77 (2009). https://doi.org/10.3354/ame01330
Yang, G.-P., Tsunogai, S.: Biogeochemistry of dimethylsulfide (DMS) and dimethylsulfoniopropionate (DMSP) in the surface microlayer of the western North Pacific. Deep-Sea Res. I Oceanogr. Res. Pap. 52, 553–567 (2005). https://doi.org/10.1016/j.dsr.2004.11.013
Keller, M., Bellows, W., Guillard, R.: Biogenic sulfur in the environment. American Chemical Society (Symposium Series 393), pp 167–182 (1989)
Morris, R.M., Rappe, M.S., Connon, S.A., Vergin, K.L., Siebold, W.A., Carlson, C.A., Giovannoni, S.J.: SAR11 clade dominates ocean surface bacterioplankton communities. Nature. 420, 806–810 (2002). https://doi.org/10.1038/nature01240
Acknowledgments
We appreciate all the scientists and crew members on the Dongfang Hong 2 during the expedition for their great efforts and help in sample collection. We thank Yahui Gao of Xiamen University for providing Chl a data, and Yu Xin of Ocean University of China for providing nutrient measurements.
Funding
This work was supported by the National Key Research and Development Program of China (grant 2016YFA0601303) and the National Natural Science Foundation of China (grants 91751202, 41730530, and 41476112) to X-HZ, and the Natural Environmental Research Council, UK (grants NE/N002385, NE/P012671, and NE/S001352) to JDT.
Author information
Authors and Affiliations
Contributions
X-HZ and JDT designed the experiments, analyzed the data, and wrote the manuscript. HS collected samples, performed experiments, analyzed the data, and wrote the manuscript. G-PY, YHZ, and YFZ analyzed the data. SZ performed statistical analysis. ST performed part of the qPCR experiments. Q-YM performed the DMS and DMSP detection. All the authors edited and approved the manuscript.
Corresponding authors
Ethics declarations
Conflict of Interest
The authors declare that they have no conflict of interest.
Electronic Supplementary Material
ESM 1
Five supplementary figures and six supplementary tables are available with this paper. (PDF 1011 kb)
Rights and permissions
About this article
Cite this article
Sun, H., Zhang, Y., Tan, S. et al. DMSP-Producing Bacteria Are More Abundant in the Surface Microlayer than Subsurface Seawater of the East China Sea. Microb Ecol 80, 350–365 (2020). https://doi.org/10.1007/s00248-020-01507-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00248-020-01507-8