Abstract
In cell-based research, the process of visually monitoring cells generates large image datasets that need to be evaluated for quantifiable information in order to track the effectiveness of treatments in vitro. With the traditional, end-point assay-based approach being error-prone, and existing computational approaches being complex, we tested existing machine learning frameworks to find methods that are relatively simple, yet powerful enough to accomplish the goal of analyzing cell microscopy data. This paper details the machine learning pipeline for pixel-based classification and object-based classification. Furthermore, it compares the performances of three classifiers. The classifiers evaluated were the fast-random forest (RF), the sequential minimal optimization (SMO), and the Bayesian network (BN). Images were first preprocessed using smoothing and contrast methods found in FIJI. For pixel-based classification, the preprocessed images were fed into the Trainable Waikato Segmentation (TWS). For object-based classification, training and classification were conducted within the Waikato Environment for Knowledge Analysis (WEKA) interface. All classifiers’ performance was evaluated using the WEKA experimental explorer. In terms of performance, the BN had the lowest classification accuracy for both the pixel-based and object-based model. The object-based SMO classifier had the best performance with the lowest mean absolute error of 0.05. The TWS and WEKA interface allows users to easily create and train classifiers for image analysis. However, for analyzing large image datasets, they are not ideal.

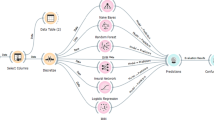
Grapical abstract







Similar content being viewed by others
References
Sommer C, Gerlich DW (2013) Machine learning in cell biology – teaching computers to recognize phenotypes. J Cell Sci 126. https://doi.org/10.1242/jcs.123604
Baatz M, Arini N, Schäpe A, Binnig G, Linssen B (2006) Object-oriented image analysis for high content screening: detailed quantification of cells and sub cellular structures with the cellenger software. Cytometry Part A:69. https://doi.org/10.1002/cyto.a.20289
Tarca AL, Carey VJ, Chen X, Romero R, Drăghici S (2007) Machine learning and its applications to biology. PLoS Comput Biol 3. https://doi.org/10.1371/journal.pcbi.0030116
Arganda-Carreras I, Kaynig V, Rueden C, Eliceiri KW, Schindelin J, Cardona A, Sebastian Seung H (2017) Trainable Weka segmentation: a machine learning tool for microscopy pixel classification. Bioinformatics (Oxford, England) 33. https://doi.org/10.1093/bioinformatics/btx180
Sonka M, Hlavac V, Boyle R (1993) Image pre-processing. In: Image Processing, Analysis and Machine Vision. Springer US, Boston
Chitradevi B, Srimanthi P (2014) An overview on image processing techniques. ISRN Signal Process 2. https://doi.org/10.1155/2013/496701
Maini R, Aggarwal H (2009) Study and Comparison of Various Image Edge Detection Techniques. In: International Journal of Image Processing (IJIP)
Pérez-Benito C, Morillas S, Jordán C, Conejero JA (2017) Smoothing vs. sharpening of color images - together or separated. Appl Math Nonlinear Sci. https://doi.org/10.21042/AMNS.2017.1.00025
Singh A, Yadav S, Singh N (2016) Contrast enhancement and brightness preservation using global-local image enhancement techniques. In: 2016 4th International Conference on Parallel, Distributed and Grid Computing, PDGC 2016
Blaschke T, Burnett C, Pekkarinen A (2004) Image segmentation methods for object-based analysis and classification. In: Jong SMD, Meer FDV (eds) Remote Sensing Image Analysis: Including the Spatial Domain, pp.211-236. Remote Sensing and Digital Image Processing, vol 5. Springer, Dordrecht. https://doi.org/10.1007/978-1-4020-2560-0_12
Yuheng S, Hao Y Image segmentation algorithms overview
Anjna E, Rajandeep Kaur E (2017) Review of image segmsentation technique
Shariff A, Kangas J, Coelho LP, Quinn S, Murphy RF (2010) Automated image analysis for high-content screening and analysis. J Biomol Screen 15. https://doi.org/10.1177/1087057110370894
Kotsiantis SB (2007) Supervised machine learning: a review of classification techniques
Mayo M (2007) Effective classifiers for detecting objects. CIRAS
Bin Othman MF, Yau TMS (2007) Comparison of Different Classification Techniques Using WEKA for Breast Cancer. In: Ibrahim F, Osman NAA, Usman J, Kadri NA (eds) 3rd Kuala Lumpur International Conference on Biomedical Engineering 2006. IFMBE Proceedings, vol 15. Springer, Berlin, Heidelberg. https://doi.org/10.1007/978-3-540-68017-8_131
Kelner R, Lerner B (2012) Learning Bayesian network classifiers by risk minimization. In: International Journal of Approximate Reasoning
Frank E, Hall M, Holmes G, Kirkby R, Pfahringer B, Witten IH, Trigg L Chapter 1 WEKA a machine learning workbench for data mining
Soille P, Vincent LM (1990) Determining watersheds in digital pictures via flooding simulations. In: Visual Communications and Image Processing '90. Society of Photo-Optical Instrumentation Engineers (SPIE). https://doi.org/10.1117/12.24211
Paynter G, Trigg L, Kirkby R (2008) Attribute-relation file format (ARFF). In: The University of Waikato. Accessed 4 Nov 2018
Li W, Mao KZ, Zhang H, Chai T (2010) Selection of gabor filters for improved texture feature extraction. In: Proceedings - International Conference on Image Processing, ICIP
Krishan A (2012) Evaluation of Gabor filter parameters for image enhancement and segmentation. In: International Journal of Advanced Research in Computer Engineering & Technology (IJARCET), Volume 1, Issue 7. ISSN: 2278 – 1323
Bai Y, Guo L, Jin L, Huang Q (2009) A novel feature extraction method using pyramid histogram of orientation gradients for smile recognition. In: Proceedings - International Conference on Image Processing, ICIP
Bouckaert RR, Frank E, Hall M, Kirkby R, Reutemann P, Seewald A, Scuse D (2013) WEKA Manual for Version 3-7-8
Frank E, Hall MA, Witten IH (2016) The WEKA Workbench. Online Appendix for “Data Mining: Practical Machine Learning Tools and Techniques". Morgan Kaufmann, Fourth Edition
Acknowledgments
We thank Timothy Samec for cultivating and imaging the LN-18 cells.
Funding
This research was funded by the Transformative Initiative for Generating Extramural Research (TIGER) grant at Clemson University.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Mbiki, S., McClendon, J., Alexander-Bryant, A. et al. Classifying changes in LN-18 glial cell morphology: a supervised machine learning approach to analyzing cell microscopy data via FIJI and WEKA. Med Biol Eng Comput 58, 1419–1430 (2020). https://doi.org/10.1007/s11517-020-02177-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11517-020-02177-x