Abstract
The smut fungal genus Microbotryum (Microbotryales, Pucciniomycotina) contains species that parasitize plants from many different lineages of euasterids, with host specificity of individual parasite species in general being exceptionally high. Additionally, it has been shown that the location of spore production in some species is related to spore dispersal. In this phylogenetic study based on ITS and LSU rDNA data of 57 Microbotryum spp., host spectra and sorus location are mapped on the phylogeny of Microbotryum species in order to understand the macroevolutionary patterns of these two traits. We find that monophyletic parasite clades correspond well with monophyletic host clades and also that monophyletic parasite groups in general produce their spores in the same plant organ. Ancestral state reconstruction inferred the most probable ancestral trait for sorus location being leaves and the most probable ancestral host family for the genus Microbotryum as being the Polygonaceae. According to molecular analyses, a newly sequenced specimen of Ustilago ducellieri, a seed parasite on Arenaria leptoclados, previously treated as synonym of Microbotryum duriaeanum, belongs to a lineage distinct from specimens of M. duriaeanum. A new combination, Microbotryum ducellieri, is accordingly proposed. Taxonomic implications of the presented analyses for the genera Bauhinus and Haradaea are briefly discussed.
Similar content being viewed by others
Introduction
The fungal genus Microbotryum, with 98 described species (Denchev and Denchev 2011; Vánky 2012; Piątek et al. 2012, 2013; Ziegler et al. 2018; Denchev et al. 2019), is mainly known for containing the agents of anther smut disease in hosts of the Caryophyllaceae. In this group of parasites, it is assumed that the production of teliospores in the anthers is most likely an adaptation to host pollinators that increases parasite dispersal. However, the host range of Microbotryum is much broader and also includes host species in the plant families Asteraceae, Caprifoliaceae, Gentianaceae, Lamiaceae, Lentibulariaceae, Montiaceae Onagraceae, and Polygonaceae (Fig. 1). Additionally, there is one report of a Microbotryum species on a host belonging to the Primulaceae, which is however not confirmed by voucher material. Previous studies have demonstrated that anther smuts on hosts in the Caryophyllaceae form a monophyletic group within Microbotryum (Almaraz et al. 2002; Kemler et al. 2006, 2009). Accordingly, sorus formation in anthers of the Caryophyllaceae species evolved only once. Following this single colonization event, a major radiation led to the formation of Microbotryum species on many members of the Caryophyllaceae (Lutz et al. 2005, 2008; Le Gac et al. 2007; Refrégier et al. 2008; Piątek et al. 2012, 2013; Smith et al. 2017; Denchev et al. 2019; Tang et al. 2019). Anther smut infections are widespread throughout host distributions (Hood et al. 2010). Recurrently, monophyletic host genera within the Caryophyllaceae have been colonized once with subsequent divergence on many host species within the host genus. However, there are exceptions: North American Silene species for instance were colonized independently from European Silene species, whereas others were colonized by host-jumps from non-Silene hosts, such as Silene italica and S. viscosa clustering in a group containing only species on Atocion, Heliosperma, and Viscaria (Lutz et al. 2005, 2008; Piątek et al. 2012, 2013; Kemler et al. 2013; Smith et al. 2017). These studies established that host specificity in the anther smuts on the Caryophyllaceae is higher than previously assumed, and many species were described or re-instated. The pattern of singular colonization, subsequent divergence, and high host specificity thereby seems to be more general in the genus Microbotryum than previously assumed based on morphological characteristics and has been observed to a smaller extent in other host groups (Kemler et al. 2009; Ziegler et al. 2018). However, other observations show that this pattern does not hold true all the time. Anther smuts on Montiaceae for instance are dispersed throughout the clade of caryophyllaceous anther smuts and do not form a monophyletic group (Hood et al. 2010), and parasitism of Polygonaceae can be found throughout the phylogenetic tree of Microbotryum (Kemler et al. 2006, 2009), whereas parasitism on Caprifoliaceae has evolved twice (Kemler et al. 2006, 2009). These examples show that colonization patterns in Microbotryum are complicated, and given that the number of species in the genus examined by molecular methods is still low, further research is needed to understand the patterns of host specificity in Microbotryum.
Examples of Microbotryum species parasitizing different host plant species belonging to diverse families and producing sori in different host organs. aM. salviae in the anthers of Salvia pratensis (Lamiaceae); bM. marginale in the leaves of Persicaria bistorta (Polygonaceae); cM. bistortarum in the flowers of Persicaria vivipara (Polygonaceae); dM. pustulatum in the leaves of Persicaria bistorta (Polygonaceae); eM. alpinum in the anthers of Pinguicula alpina (Lentibulariacae); fM. duriaeanum in the ovules of Cerastium brachypetalum (Caryophyllaceae); gM. tragopogonis-pratensis replacing the florets of Tragopogon pratensis (Asteraceae; a healthy flower head is shown in the forefront). © all pictures M. Kemler except f) M. Piątek
The parasitism of Microbotryum species in their hosts in vivo is visible upon teliospore presentation of the parasite in soral structures. Different Microbotryum species develop sori in different organs of their hosts. The most well-known species, the anthericolous smuts, develop sori in the anthers of Caprifoliaceae, Caryophyllaceae, Lamiaceae, Lentibulariaceae, Montiaceae, and Primulaceae. Spore production of other species occurs in ovaries or seeds (Caprifoliaceae, Caryophyllaceae, Gentianaceae, Montiaceae, Onagraceae, and Polygonaceae), flowers (Asteraceae, Montiaceae, and Polygonaceae), leaves (Polygonaceae), stems (Polygonaceae), or a combination of organs (Polygonaceae). The same sorus location occurs in different host families, but it remains unclear whether or how often the transitions between the different soral sites have occurred within Microbotryum.
In order to gain more insight into these topics, this study addressed the following questions: (1) Are the caryophyllaceous Microbotryum species that form sori in ovules/seeds monophyletic, and how are they related to the caryophyllaceous anther smuts? (2) What is the ancestral state of sorus location in the genus Microbotryum, and how is the location of spore production reflected by Microbotryum phylogeny? (3) Are monophyletic clades of parasites restricted to monophyletic groups of hosts, and what is the most likely ancestral host genus parasitized by members of the genus Microbotryum? To answer these questions, molecular phylogenetic analyses and ancestral state reconstructions were performed based on a broad species sampling that covers 57 Microbotryum species, many of which are considered in phylogenetic analyses here for the first time.
Materials and methods
Specimens and morphological analyses
The specimens newly sequenced for this study are shown in Table 1. The voucher specimens are deposited in the herbaria of the Botanischer Garten und Botanisches Museum Berlin-Dahlem (B), Herbarium Ustilaginales Vánky (HUV, deposited in BRIP), Royal Botanic Gardens, Kew (K(M)), Komarov Botanical Institute, Russian Academy of Sciences, St Petersburg (LE), Botanische Staatssammlung München (M), Royal Botanic Garden, Madrid (MA), the Institute of Biodiversity and Ecosystem Research, Bulgarian Academy of Sciences, Sofia (SOMF), and the herbarium of the University of Tübingen (TUB).
A dried specimen of Ustilago duriaeana, kept at the herbarium of the Royal Botanic Garden, Madrid (MA), was examined by light microscopy (LM) and scanning electron microscopy (SEM). For LM observations and measurements, spores were mounted in lactoglycerol solution (w:la:gl = 1:1:2) on glass slides, gently heated to boiling point to rehydrate the spores, and then cooled. The measurements of spores are given as min–max (extreme values) (mean ± 1 standard deviation). For SEM, spores were attached to specimen holders by double-sided adhesive tape and coated with gold in an ion sputter. The surface structure of spores was observed and photographed at 10 kV accelerating voltage using a ZEISS Sigma VP scanning electron microscope.
DNA isolation, PCR, and sequencing
Genomic DNA was isolated from herbarium material using the DNeasy™ Plant Mini Kit (Qiagen, Hilden, Germany) according to the manufacturer’s instructions. Parts of the ribosomal DNA large subunit (LSU) and the internal transcribed spacer (ITS) were amplified using PCR. The LSU was amplified using the primer pair LR0R/NL4 or NL1/NL4 (White et al. 1990; O’Donnell 1992, 1993). The ITS was either amplified using ITS1f/ITS4 or ITS1f/5.8S and 5.8Srev/ITS4 (Vilgalys and Hester 1990; White et al. 1990, Gardes and Bruns 1993). Samples were sequenced using the BigDye™ Terminator Cycle Sequencing Kit V3.1 on an ABI 3100 Genetic Analyser (Applied Biosystems, Waltham, USA). The sequences obtained in this study were deposited in GenBank under accession numbers MN657185–MN657204 (ITS) and MN657208–MN657227 (LSU; Table 1).
Phylogenetic analyses
To infer the phylogenetic relationships within the genus Microbotryum, a dataset comprising newly generated sequences and representative sequences for Microbotryum species taken from GenBank (Table S1) was assembled. For sequences obtained from GenBank, one specimen per species was chosen. For species retrieved from GenBank, we only used specimens that had both ITS and LSU sequences in GenBank. As we were interested in the host spectrum of species in the Asteraceae, the specimens of parasites on Asteraceae only represented by ITS sequences in GenBank were also included in the analyses. The two DNA regions were aligned separately using MAFFT v7.305b (Katoh et al. 2002; Katoh and Standley 2013) under the “linsi” option. The alignments were concatenated using SequenceMatrix 1.7.8 (Vaidya et al. 2011), and leading and trailing gaps were coded as missing state. A preliminary tree was inferred using RAxML 8.2.11 (Stamatakis 2014) under the GTRGAMMA model of rate heterogeneity and a rapid bootstrap inference of 100 replications. In order to get a better representation of variable regions, the resulting phylogeny was subsequently used as a guide tree for the phylogeny aware aligner PAGAN (Löytynoja et al. 2012). This approach resulted in numerous indels in variable regions, and therefore this additional information was used in a combined approach of DNA and indel information. Indels were coded after Simmons and Ochoterena (2000) in SeqState (Müller 2005). Indel data was then combined with the DNA data, and the resulting partitioned alignment was used as an input for a ML analysis in RAxML using the BINGAMMA model of rate heterogeneity and a rapid bootstrap inference of 1000 replicates. Additionally, a phylogenetic tree from the concatenated alignments of the original PAGAN analysis without indel information was inferred under the same conditions as the original MAFFT alignment.
Ancestral state reconstructions for sorus location and host family were performed in Mesquite 3.51 (Maddison and Maddison 2015) using a ML approach applying the Mk1 model (Maddison and Maddison 2006). The individual states for these traits were taken from Denchev and Denchev (2011), Piątek et al. (2012, 2013), Vánky (2012), and Ziegler et al. (2018). States for sorus location were coded as follows: (a) not applicable (N/A), (b) anthers, (c) anthers and filaments, (d) flower, (e) inflorescences/pedicels/stem/leaves, (f) leaves, and (g) ovaries/ovules/seeds. In categories (e) and (g), not necessarily all species sporulate in all of these organs (for details, see Vánky 2012). States for host family were coded as follows: (a) not applicable (N/A), (b) Juncaceae, (c) Polygonaceae, (d) Caryophyllaceae, (e) Caprifoliaceae, (f) Lamiaceae, (g) Lentibulariaceae, and (h) Asteraceae.
Results
Phylogenetic analyses
The results of the phylogenetic analyses in this study to a large extent agreed with those of previous studies on the intra-generic relationships of Microbotryum. The analyses including indel data resulted both in a statistically well-supported backbone and statistically well-supported terminal clades (Fig. 2), whereas the analyses including DNA data alone showed strong support mainly for terminal clades (Fig. 3). Anther smuts on hosts in the Caryophyllaceae were inferred as monophyletic and formed a sister clade to Microbotryum anomalum on Polygonaceae. A clade containing most anther smuts on hosts in the Dipsacoideae in Caprifoliaceae formed a sister clade to anther smuts on hosts in the Caryophyllaceae and M. anomalum. A group of Microbotryum species that develop teliospores in ovules/seeds of hosts in the Caryophyllaceae were the sister group to the clade containing all the previously mentioned taxa. In our analyses the seed parasites on Cerastium spp. formed a monophyletic group. However, Microbotryum duriaeanum was inferred as paraphyletic as M. afromontanum fell within a clade otherwise only containing M. duriaeanum. Microbotryum intermedium, an anther smut on Scabiosa species (Caprifoliaceae), did not cluster with other anther smuts on hosts in the Caprifoliaceae but formed a monophyletic group with anther smuts on hosts in the Lamiaceae (M. betonicae and M. salviae) and Lentibulariaceae (M. alpinum and M. liroi). Microbotryum species on hosts in the Asteraceae formed a monophyletic clade. Parasite species delimitations were not resolved in two cases in this host group as M. scorzonerae and M. cardui were inferred as paraphyletic. However, paraphyly for M. cardui had low statistic support. Like in previous studies, parasites on hosts in the Polygonaceae formed several groups that were dispersed throughout the phylogenetic tree, as well as formed the earliest diverging lineage of Microbotryum.
Phylogenetic relationships for species of Microbotryum based on likelihood analysis of the dataset using indel coding. Branch thickness is relative to the support values obtained (compare Fig. 3). The phylogeny was rooted with Bauerago abstrusa and Microbotryozyma collariae. K.V.U.E. – K. Vánky Ustilaginales Exsiccata. M. – Microbotryum
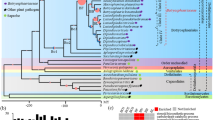
Cladogram of the same phylogenetic tree as in Fig. 2 including host information. Values above branches are bootstrap values of the RAxML analysis for the PAGAN alignment with indel coding, the PAGAN alignment of the original data, and the original MAFFT alignment. Only bootstrap values above 65 are shown. M. – Microbotryum
Ancestral state reconstruction of sorus location indicated that the ancestor of the genus Microbotryum could have either formed its sori in leaves (proportional likelihood: ~0.52) or in inflorescences (combined proportional likelihood: ~0.34; Fig. 4). Anther parasitism seems to have evolved at least two times among Microbotryum spp. There is a medium probability (proportional likelihood: ~0.47) that the ancestor of the clade containing anther smuts on hosts in the Caryophyllaceae and Caprifoliaceae, M. anomalum, and ovules/seeds parasites on hosts in the Caryophyllaceae was an anther smut. Seed/ovary parasitism has evolved twice among the taxa included in the present analysis, once resulting in seed parasites on hosts in the Caryophyllaceae and once in Sphacelotheca. Parasitism in the leaves most likely evolved only once in the taxa included in the present analysis. Interspersed with these “conservative” locations of sorus formation are species that develop sori in more than one organ. These can be found in the anther smuts group (i.e., Microbotryum adenopetalae and M. majus), as well as in the group of leaf smuts (i.e., M. bosniacum and M. emodensis). Additionally, M. parlatorei, which has no close affiliations to other species, can sporulate in stems, petioles, leaves, and flowers, although predominately it produces spores in the stems.
Ancestral state reconstruction of host family (Fig. 2) resulted in a high probability of the Polygonaceae being the first plant family colonized by members of the genus Microbotryum (proportional likelihood: ~0.98). Caryophyllaceae have a medium likelihood of having been colonized only once, irrespective of their parasites developing spores in anthers or in ovules/seed (proportional likelihood: ~0.45). However, there is nearly an equal likelihood that the ancestor of the clade containing all Caryophyllaceae parasites, as well as Caprifoliaceae parasites (except Microbotryum intermedium) and M. anomalum was a parasite on Polygonaceae (proportional likelihood: ~0.42). Caprifoliaceae have been colonized twice independently by Microbotryum species and both times in the anthers. All other host families (i.e., Asteraceae, Lamiaceae, and Lentibulariaceae) except for Polygonaceae, have been colonized only once. However, in the case of Lamiaceae, subsequent jumps to Caprifoliaceae and Lentibulariaceae hosts occurred.
Taxonomy
Two smut fungi have been described on Arenaria spp.: Ustilago ducellieri Maire (on Arenaria serpyllifolia L. from Algeria, Maire 1917) and U. arenariae-bryophyllae Vánky (on Arenaria bryophylla Fernald from India, Vánky 1983). Ustilago ducellieri was reduced to a synonym of U. duriaeana Tul. & C. Tul. by Zundel (1953), which is based on a type specimen on Cerastium glomeratum Thuill., and later transferred to Microbotryum (as M. duriaeanum (Tul. & C. Tul.) Vánky, Vánky 1998). Ustilago arenariae-bryophyllae is considered a distinct species, namely, M. arenariae-bryophyllae (Vánky) Vánky (Vánky 1998, 2012). Although spore ornamentation shows similarity to M. duriaeanum (Vánky 2012: Fig. on page 367), based on the phylogenetic position (Figs. 2 and 3), it is evident that the sequenced specimen on Arenaria leptoclados (Rchb.) Guss. from Spain is not Microbotryum duriaeanum. Moreover, this specimen has reticulate spore ornamentation (Fig. 5) that differs significantly from the verruculosely reticulate ornamentation of Microbotryum arenariae-bryophyllae (Vánky 1983: Fig. 15, 2012: Fig. on page 351). In addition, Arenaria bryophylla has recently been transferred to Eremogone as E. bryophylla (Fernald) Sadeghian & Zarre (Sadeghian et al. 2015). Arenaria leptoclados and A. serpyllifolia on the other hand are sister species within Arenaria sect. Arenaria (Sadeghian et al. 2015). Host specificity within the ovules/seeds smuts on hosts in the Caryophyllaceae might not be that high, as exemplified by Microbotryum duriaeanum (Fig. 2), and therefore, we consider the smut fungi on Arenaria leptoclados and Arenaria serpyllifolia as belonging to the same species. A new combination of Ustilago ducellieri in Microbotryum is proposed here, along with a description and illustrations.
Microbotryum ducellieri (Maire) Kemler, T. Denchev, Denchev & M. Lutz, comb. nov. (Fig. 5)
Index Fungorum number: IF 556468
Basionym: Ustilago ducellieri Maire, Bull. Soc. Hist. Nat. Afrique N. 8: 140, 1917.—Holotype on Arenaria serpyllifolia L., Algeria, Algiers, dunes near El Harrach (as “Dunes de Maison-Carrée”), 10 March 1912, L. Ducellier, Herb. Maire no. 560 (MPU, n.v.).
Infection systemic. Sori destroying the seeds, filling the capsules with a reddish brown, initially semi-agglutinated, later powdery spore mass. Spores globose, subglobose, broadly ellipsoidal or ovoid, sometimes slightly irregular, (11–)12–15(−16.5) × (10–)11–13.5(−14.5) (13.3 ± 0.8 × 12.2 ± 0.7) μm (n = 200), medium yellowish brown to medium reddish brown; wall reticulate, 1.3–1.8(−2.2) μm thick (including reticulum), meshes 5–7(−8) per spore diameter, polyhedral or irregular, 0.7–2.5(−3.0) μm long; muri (17–)18–21(−23) on equatorial circumference, in optical median view subacute or acute, 0.5–1.0(−1.3) μm high. Immature hyaline spores may be present. As seen by SEM meshes rugulose on the bottom, often with a hemispherical protuberance.
Specimen examined: on Arenaria leptoclados, Spain, Andalusia, Jaén, Quesada, at the foot of Cabañas Peak, 13 June 1996, T. Almaraz et al., no. TAL400 (MA-Fungi 37,800 as “Ustilago duriaeana”).
Discussion
This study provides the most comprehensive phylogeny of the genus Microbotryum conducted to date. Compared with previous phylogenies (Kemler et al. 2006, 2009), the current dataset was expanded to include all described species of anther smuts on hosts in the Caprifoliaceae, all Microbotryum species known on hosts in the Asteraceae, and most described ovule/seed parasites of Caryophyllaceae, except for M. alsines, M. arenariae-bryophyllae, and M. nivale. Consequently, this study provides a more complete understanding of the macroevolution within the genus Microbotryum than previous studies.
Microbotryum species that form sori in ovules/seeds of Caryophyllaceae
Phylogenetic analyses revealed a monophyletic origin of ovule/seed parasite species on hosts in the Caryophyllaceae and show that they neither originated from caryophyllaceous anther smuts nor do they form their sister clade. Thus, either an early split of Microbotryum species parasitizing hosts in the Caryophyllaceae with subsequent colonization of hosts in other plant families occurred or independent colonization on the one side led to caryophyllaceous anther smuts and on the other side to caryophyllaceous ovule/seed parasites.
Research on caryophyllaceous anther smuts has shown that there exists a high number of often host-species-specific parasite species, many of which have been described only in recent years based on molecular data, host information, and sometimes also on morphological differences. Currently, 24 species are recognized (Lutz et al. 2005, 2008; Denchev 2007a, b; Denchev et al. 2009, 2019; Denchev and Denchev 2011; Vánky 2012; Piątek et al. 2012, 2013). Based on morphological species concepts prevalent in the existing literature, species diversity and host specificity seem lower in ovule/seed parasites (ten species), but our phylogenies indicate that host specific lineages exist in this group, e.g., the smut fungi infecting different Cerastium spp. and different Moenchia spp. form monophyletic lineages, and it is most likely that cryptic species in this clade will be discovered in future studies. However, species delimitations within the ovule/seed parasites are not always satisfying, as for instance Microbotryum afromontanum (on Cerastium afromontanum) clusters within M. duriaeanum (on C. brachypetalum and C. gracilis). Further studies in this group including many more specimens are needed to understand the patterns of host specificity and also whether the ovule/seed parasites are host specialists, as seen in anther smuts of Silene spp. (Lutz et al. 2005, 2008; Piątek et al. 2012, 2013; Kemler et al. 2013; Denchev et al. 2019), or if they constitute generalists, as the anther parasites of Dianthus spp. (Kemler et al. 2013) and Saponaria spp. (Lutz et al. 2005; Fortuna et al. 2018).
Ancestral state reconstruction of sorus location
Most species in Microbotryum produce their teliospores in the inflorescences of their hosts. However, contrary to previous assumptions (Kemler et al. 2006), ancestral state reconstruction indicates that parasitism in any part of the inflorescence might be a derived trait. Based on the current dataset, there is a high probability that the ancestor of the genus Microbotryum produced its teliospores in leaves. Species that produce spores in leaves (i.e., M. marginale, M. pustulatum, and M. tuberculiforme) and stems (M. nepalense; see Kemler et al. 2006) appear in the earliest diverging lineage and only one other, partially leaf sporulating species (i.e., M. parlatorei) diverged later from species sporulating in flowers. Future work should consider other species (e.g., M. ocrearum and M. piperi) sporulating in leaves and/or stems of their hosts to determine the evolution of sorus location.
The location of spore production often co-correlates well with phylogenetic lineages of Microbotryum and with host lineages (Fig. 4). For instance, all species infecting hosts in the Asteraceae form a monophyletic group and result in similar host morphology by modifying the entire capitula (Fig. 1g). Despite this, sorus location between lineages of Microbotryum is a variable trait in the macroevolution of this genus. It is highly variable between sister groups (e.g., flower sporulating species in hosts of the Asteraceae form a sister clade to anther parasites in hosts of the Caprifoliaceae, Lamiaceae, and Lentibulariaceae). Even within clades, there are species that do not conform to the general trend of the group (e.g., Microbotryum adenopetalae and M. majus among caryophyllaceous anther smuts). Similar variability of sorus location has also been observed within a species during accidental or artificial non-host infection. Studies of incipient host jumps in nature have shown that parasite occurrence of the newly infected hosts results in a distorted host morphology not seen in the original host (Antonovics et al. 2002; Kummer 2010). Such non-host infections can also be accompanied by spore production in tissues that are not used by the same fungal species on the original host. For instance, infection experiments of different caryophyllaceous anther smuts on four different host species conducted to better understand host specificity, frequently resulted in the spores being produced in the ovules instead of the anthers (Sloan et al. 2008).
Monophyletic clades of parasites on monophyletic clades of hosts and ancestral host family
The genus Microbotryum has an enormous host spectrum, and species within the genus occur on hosts of several unrelated plant families, a situation only rivaled among smut fungi by some Ustilaginomycotina smut genera (e.g., Entyloma and Urocystis). Host plants of Microbotryum species belong to the Asterales (Asteraceae), Caryophyllales (i.e., Caryophyllaceae, Montiaceae, and Polygonaceae), Dipsacales (Caprifoliacae), Ericales (Primulaceae), Gentianales (Gentianaceae), Lamiales (Lamiaceae and Lentibulariaceae), and Myrtales (Onagraceae). Supporting results of previous studies (Kemler et al. 2006, 2009), our data on many Microbotryum species from different host species show that monophyletic groups of parasites mainly occur on monophyletic clades of host species. However, some host groups have been colonized independently more than once (i.e., hosts in the Caprifoliaceae and Caryophyllaceae). The colonization of a new host clade initiated a radiation within the clade with adaptation and speciation of Microbotryum spp. on different species in such host clades, a pattern well known from other parasite groups (e.g., McTaggart et al. 2015; Kruse et al. 2018). Additionally, inter-family host jumps sometimes occurred, for example, in the clade of anther smuts on the Caryophyllaceae (jump to Montiaceae; Hood et al. 2010) or Lamiaceae (jump to Lentibulariaceae; Ziegler et al. 2018).
Taxonomic implications for the genera Bauhinus and Haradaea
Our study additionally has some implications on the taxonomy of genera associated with Microbotryum. Moore (1992) proposed the genus Bauhinus for all “dicot” smuts of Ustilago and assigned six species to the genus. Vánky (1993) treated Bauhinus as a nomenclaturally superfluous name and reduced Bauhinus to a synonym of Microbotryum because the species of Microbotryum Léveillé were parasites also on dicotyledonous plants. However, the generic name Bauhinus is validly published and legitimate with the type species Uredo tragopogonis-pratensis Pers. Further, Ustilago species on non-caryophyllaceous hosts were transferred to Bauhinus by Denchev (1997) and Denchev et al. (2006). Based on our phylogeny, following the Bauhinus concept would result in this genus being paraphyletic and in order to define monophyletic genera, many new genera would need to be erected.
A similar problem occurs with the genus Haradaea, which accommodates a group of former Ustilago species destroying the ovules/seeds of several host species in the Caryophyllaceae by filling the capsules with a purplish spore mass (Denchev et al. 2006). Although our analyses confirmed the monophyletic origin of this clade of ovule/seed parasites, it clearly clusters within Microbotryum in its current circumscription. Contrary to this, a previous analysis of Microbotryum (Almaraz et al. 2002) supported Haradaea as a separate genus outside the Microbotryum lineage. Most likely, the previous topology by Almaraz et al. (2002) arose due to the sequencing of contaminating Holtermaniella festucosa (based on BLASTn hits of the ITS sequences AF287152 of Ustilago duriaeana). Like in the case of Bauhinus, following the Haradaea concept in this latter form would result in splitting Microbotryum in a bulk of genera. At this point, we refrain from drawing premature taxonomic conclusions and prefer to keep Microbotryum in its current broad circumscription. Up to date, systematic studies of Microbotryum lack most of the species from different hosts in the Polygonaceae, which is the most species-rich host family and most likely also the ancestral host family of this group of smut fungi.
Conclusions
Our study provides an updated evolutionary framework for the genus Microbotryum that helps to understand trait evolution and host specificity in plant parasitic fungi. We found a correlation between the monophyletic groups within Microbotryum with monophyletic host lineages, as well as with sorus location, but the pattern is not straightforward. In general, monophyletic parasite groups occur on monophyletic host groups with all species in this clade expressing a similar sorus location on their host plants. On the other side, parasite sister clades on different host clades can express a very different location of spore production. Further research using additional tools (e.g., genomics and transcriptomics) is certainly needed to understand the interplay between mechanisms of host specificity and sorus location. Additionally, the study increases the number of species accepted within Microbotryum to 99, emphasizing that species numbers in this genus will continue to rise by the application of molecular methods.
References
Almaraz T, Roux C, Maumont S, Durrieu G (2002) Phylogenetic relationships among smut fungi parasitizing dicotyledons based on ITS sequence analysis. Mycol Res 106:541–548. https://doi.org/10.1017/S0953756202006019
Antonovics J, Hood M, Partain J (2002) The ecology and genetics of a host shift: Microbotryum as a model system. Am Nat 160(S4):S40–S53. https://doi.org/10.1086/342143
Denchev CM (1997) New combinations in Bauhinus (Microbotryaceae). Mycotaxon 65:419–426
Denchev CM (2007a) Microbotryum lagerheimii sp. nov. (Microbotryaceae). Mycol Balc 4:61–67. https://doi.org/10.5281/zenodo.2547791
Denchev CM (2007b) Microbotryum savilei sp. nov. (Microbotryaceae). Mycol Balc 4:69–73. https://doi.org/10.5281/zenodo.2547803
Denchev CM, Denchev TT (2011) New records of smut fungi. 4. Microbotryum coronariae comb. nov. Mycotaxon 118:53–56. https://doi.org/10.5248/118.53
Denchev CM, Moore RT, Shin HD (2006) A reappraisal of the genus Bauhinus (Microbotryaceae). Mycol Balc 3:71–75. https://doi.org/10.5281/zenodo.2547355
Denchev CM, Giraud T, Hood ME (2009) Three new species of anthericolous smut fungi on Caryophyllaceae. Mycol Balc 6:79–84. https://doi.org/10.5281/zenodo.2548933
Denchev TT, Kemler M, Begerow D, Denchev CM (2019) Molecular and morphological evidences reveal a new smut fungus, Microbotryum arcticum (Microbotryaceae), on Silene uralensis (Caryophyllaceae) from Greenland and Canada. Willdenowia 49(2):241–255. https://doi.org/10.3372/wi.49.49212
Fortuna TM, Namias A, Snirc A, Branca A, Hood ME, Raquin C et al (2018) Multiple infections, relatedness and virulence in the anther-smut fungus castrating Saponaria plants. Mol Ecol 27:4947–4959. https://doi.org/10.1111/mec.14911
Gardes M, Bruns TD (1993) ITS primers with enhanced specificity for basidiomycetes - application to the identification of mycorrhizae and rusts. Mol Ecol 2(2):113–118. https://doi.org/10.1111/j.1365-294x.1993.tb00005.x
Hood ME, Mena-Alí JI, Gibson AK, Oxelman B, Giraud T, Yockteng R et al (2010) Distribution of the anther-smut pathogen Microbotryum on species of the Caryophyllaceae. New Phytol 187:217–229. https://doi.org/10.1111/j.1469-8137.2010.03268.x
Katoh K, Standley DM (2013) MAFFT multiple sequence alignment software version 7: improvements in performance and usability. Mol Biol Evol 30:772–780. https://doi.org/10.1093/molbev/mst010
Katoh K, Misawa K, Kuma K, Miyata T (2002) MAFFT: a novel method for rapid multiple sequence alignment based on fast Fourier transform. Nucleic Acids Res 30:3059–3066. https://doi.org/10.1093/nar/gkf436
Kemler M, Göker M, Oberwinkler F, Begerow D (2006) Implications of molecular characters for the phylogeny of the Microbotryaceae (Basidiomycota: Urediniomycetes). BMC Evol Biol 6:35. https://doi.org/10.1186/1471-2148-6-35
Kemler M, Lutz M, Göker M, Oberwinkler F, Begerow D (2009) Hidden diversity in the non-caryophyllaceous plant-parasitic members of Microbotryum (Pucciniomycotina: Microbotryales). Syst Biodivers 7:297–306. https://doi.org/10.1017/S1477200009990028
Kemler M, Martín MP, Telleria MT, Schäfer AM, Yurkov A, Begerow D (2013) Contrasting phylogenetic patterns of anther smuts (Pucciniomycotina: Microbotryum) reflect phylogenetic patterns of their caryophyllaceous hosts. Org Divers Evol 13:111–126. https://doi.org/10.1007/s13127-012-0115-1
Kruse J, Piątek M, Lutz M, Thines M (2018) Broad host range species in specialised pathogens should be treated with suspicion—a case study in Entyloma reveals high, hostcorrelated diversity in pathogens infecting Ranunculus. Persoonia 41:175–201. https://doi.org/10.3767/persoonia.2018.41.09
Kummer V (2010) Bemerkenswerte Pilzfunde auf der 38. Brandenburgischen Botanikertagung in Groß Pinnow/Uckermark 2007. Verh Bot Vereins Berlin Brandenburg 142:223–245
Le Gac M, Hood ME, Fournier E, Giraud T (2007) Phylogenetic evidence of host-specific cryptic species in the anther smut fungus. Evolution 61:15–26. https://doi.org/10.1111/j.1558-5646.2007.00002.x
Löytynoja A, Vilella AJ, Goldman N (2012) Accurate extension of multiple sequence alignments using a phylogeny-aware graph algorithm. Bioinformatics 28:1684–1691. https://doi.org/10.1093/bioinformatics/bts198
Lutz M, Göker M, Piątek M, Kemler M, Begerow D, Oberwinkler F (2005) Anther smuts of Caryophyllaceae: molecular characters indicate host-dependent species delimitation. Mycol Prog 4:225–238. https://doi.org/10.1007/s11557-006-0126-4
Lutz M, Piątek M, Kemler M, Chlebicki A, Oberwinkler F (2008) Anther smuts of Caryophyllaceae: molecular analyses reveal further new species. Mycol Res 112:1280–1296. https://doi.org/10.1016/j.mycres.2008.04.010
Maddison WP, Maddison DR (2006) StochChar: a package of Mesquite modules for stochastic models of character evolution. Version 1.1. http://mesquiteproject.org
Maddison WP, Maddison DR (2015) Mesquite: a modular system for evolutionary analysis. Version 3.03. http://mesquiteproject.org
Maire R (1917) Champignons Nord-Africains nouveaux ou peu connus. Bull Soc Hist Nat Afrique N 8:134–200
McTaggart AR, Doungsa-Ard C, Geering AD, Aime MC, Shivas RG (2015) A co-evolutionary relationship exists between Endoraecium (Pucciniales) and its Acacia hosts in Australia. Persoonia 35:50–62. https://doi.org/10.3767/003158515X687588
Moore RT (1992) The genus Bauhinus gen. nov.: for species of Ustilago on dicot hosts. Mycotaxon 45:97–100
Müller K (2005) SeqState: primer design and sequence statistics for phylogenetic DNA datasets. Appl Bioinforma 4:65–69. https://doi.org/10.2165/00822942-200504010-00008
O’Donnell K (1992) Ribosomal DNA internal transcribed spacers are highly divergent in the phytopathogenic ascomycete Fusarium sambucinum (Gibberella pulicaris). Curr Genet 22:213–220. https://doi.org/10.1007/BF00351728
O’Donnell KL (1993) Fusarium and its relatives. In: Taylor JW, Reynolds DR (eds) The fungal holomorph: mitotic, meiotic and pleomorphic speciation in fungal systematics. CAB International, Wallingford, pp 225–233
Piątek M, Lutz M, Ronikier A, Kemler M, Swiderska-Burek U (2012) Microbotryum heliospermae, a new anther smut fungus parasitic on Heliosperma pusillum in the mountains of the European Alpine system. Fungal Biol 116:185–195. https://doi.org/10.1016/j.funbio.2011.10.011
Piątek M, Lutz M, Kemler M (2013) Microbotryum silenes-saxifragae sp. nov. sporulating in the anthers of Silene saxifraga in southern European mountains. IMA Fungus 4:29–40. https://doi.org/10.5598/imafungus.2013.04.01.04
Refrégier G, Le Gac M, Jabbour F, Widmer A, Shykoff JA, Yockteng R, Hood ME, Giraud T (2008) Cophylogeny of the anther smut fungi and their caryophyllaceous hosts: prevalence of host shifts and importance of delimiting parasite species for inferring cospeciation. BMC Evol Biol 8(1):100. https://doi.org/10.1186/1471-2148-8-100
Sadeghian S, Zarre S, Rabeler RK, Heubl G (2015) Molecular phylogenetic analysis of Arenaria (Caryophyllaceae: tribe Arenarieae) and its allies inferred from nuclear DNA internal transcribed spacer and plastid DNA rps16 sequences. Bot J Linn Soc 178:648–669. https://doi.org/10.1111/boj.12293
Simmons MP, Ochoterena H (2000) Gaps as characters in sequence-based phylogenetic analyses. Syst Biol 49:369–381. https://doi.org/10.1093/sysbio/49.2.369
Sloan DB, Giraud T, Hood ME (2008) Maximized virulence in a sterilizing pathogen: the anther-smut fungus and its co-evolved hosts. J Evol Biol 21:1544–1554. https://doi.org/10.1111/j.1420-9101.2008.01604.x
Smith PA, Lutz M, Ziegler R, Piątek M (2017) Anther smuts of Silene acaulis and S. uniflora in the Outer Hebrides, including an assessment of ITS genotypes of Microbotryum silenes-acaulis. IMA Fungus 8:107–116. https://doi.org/10.5598/imafungus.2017.08.01.08
Stamatakis A (2014) RAxML version 8: a tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinformatics 30:1312–1313. https://doi.org/10.1093/bioinformatics/btu033
Tang H, Hood ME, Ren ZX, Li HD, Zhao YH, Wolfe LM, Li DZ, Wang H (2019) Specificity and seasonal prevalence of anther smut disease Microbotryum on sympatric Himalayan Silene species. J Evol Biol 32:451–462. https://doi.org/10.1111/jeb.13427
Vaidya G, Lohman DJ, Meier R (2011) SequenceMatrix: concatenation software for the fast assembly of multi-gene datasets with character set and codon information. Cladistics 27:171–180. https://doi.org/10.1111/j.1096-0031.2010.00329.x
Vánky K (1983) Ten new species of Ustilaginales. Mycotaxon 18:319–336
Vánky K (1993) Taxonomical studies on Ustilaginales. X Mycotaxon 48:27–44
Vánky K (1998) The genus Microbotryum (smut fungi). Mycotaxon 67:33–60
Vánky K (2012) Smut fungi of the world. APS Press, St. Paul
Vilgalys R, Hester M (1990) Rapid genetic identification and mapping of enzymatically amplified ribosomal DNA from several Cryptococcus species. J Bacteriol 172:4238–4246. https://doi.org/10.1128/jb.172.8.4238-4246.1990
White TJ, Bruns T, Lee S, Taylor J (1990) Amplification and direct sequencing of fungal ribosomal RNA genes for phylogenetics. In: Innis M, Gelfand D, Shinsky J, White T (eds) PCR protocols: a guide to methods and applications. Academic Press, San Diego, pp 315–322. https://doi.org/10.1016/B978-0-12-372180-8.50042-1
Ziegler R, Lutz M, Piątek J, Piątek M (2018) Dismantling a complex of anther smuts (Microbotryum) on carnivorous plants in the genus Pinguicula. Mycologia 110:361–374. https://doi.org/10.1080/00275514.2018.1451697
Zundel GL (1953) The Ustilaginales of the world. Contr Dept Bot School Agric Pennsylvania State Coll 176:1–410
Acknowledgment
We thank Tanja Rollnik for her help with the SEM pictures.
Funding
Open Access funding provided by Projekt DEAL. Parts of this research were supported by the German Science Foundation (BE 2201/7-2).
Author information
Authors and Affiliations
Contributions
All authors contributed to the conception and design of the study. Material preparation and data collection and analysis were performed by Teodor T. Denchev, Martin Kemler, and Matthias Lutz. The first draft of the manuscript was written by Martin Kemler, and all authors commented on previous versions of the manuscript. All authors read and approved the final manuscript.
Corresponding author
Additional information
Section Editor: Meike Piepenbring
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Topical collection on Basidiomycote Mycology in honor of Franz Oberwinkler who passed away in March 2018.
Electronic supplementary material
ESM 1
(XLSX 11 kb)
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Kemler, M., Denchev, T.T., Denchev, C.M. et al. Host preference and sorus location correlate with parasite phylogeny in the smut fungal genus Microbotryum (Basidiomycota, Microbotryales). Mycol Progress 19, 481–493 (2020). https://doi.org/10.1007/s11557-020-01571-x
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11557-020-01571-x